- NEWS AND VIEWS
Sperm CatSper ion channel swims into sharper focus
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 595, 654-655 (2021)
doi: https://doi.org/10.1038/d41586-021-01945-5
References
Lin, S., Ke, M., Zhang, Y., Yan, Z. & Wu, J. Nature 595, 746–750 (2021).
Qi, H. et al. Proc. Natl Acad. Sci. USA 104, 1219–1223 (2007).
Ren, D. et al. Nature 413, 603–609 (2001).
Quill, T. A., Ren, D., Clapham, D. E. & Garbers, D. L. Proc. Natl Acad. Sci. USA 98, 12527–12531 (2001).
Cai, X., Wang, X. & Clapham, D. E. Mol. Biol. Evol. 31, 2735–2740 (2014).
Wang, H., McGoldrick, L. L. & Chung, J.-J. Nature Rev. Urol. 18, 46–66 (2021).
Kirichok, Y., Navarro, B. & Clapham, D. E. Nature 439, 737–740 (2006).
Hwang, J. Y. et al. Cell 177, 1480–1494 (2019).
Zhao, Y. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.06.19.448910 (2021).
Chung, J.-J. et al. Cell 157, 808–822 (2014).
Competing Interests
The authors declare no competing interests

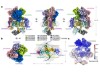 Read the paper: Structure of a mammalian sperm cation channel complex
Read the paper: Structure of a mammalian sperm cation channel complex
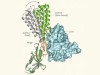 When sperm meets egg
When sperm meets egg
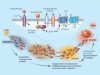 Swimming with sperm
Swimming with sperm