- NEWS
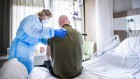
Coronavirus variants are spreading in India — what scientists know so far
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 593, 321-322 (2021)
doi: https://doi.org/10.1038/d41586-021-01274-7
References
Cherian, S. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.04.22.440932 (2021).
Hoffmann, M. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.05.04.442663 (2021).
Yadav, P. D. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.05.05.442760 (2021).
Ferreira, I. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.05.08.443253 (2021).
Yadav, P. D. et al. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciab411 (2021).

 India’s massive COVID surge puzzles scientists
India’s massive COVID surge puzzles scientists
 India’s COVID-vaccine woes — by the numbers
India’s COVID-vaccine woes — by the numbers
 Why India must tackle a mutating virus head-on
Why India must tackle a mutating virus head-on