- NEWS FEATURE
The explosion of new coronavirus tests that could help to end the pandemic
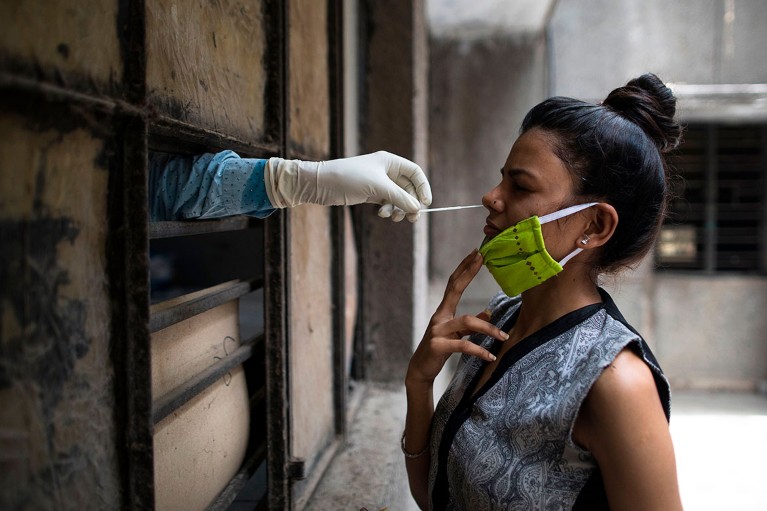

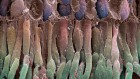
Viral diagnostics often rely on a nasal-swab sample, and researchers are developing faster, simpler and cheaper methods of testing. Credit: Xavier Galiana/AFP via Getty
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 583, 506-509 (2020)
doi: https://doi.org/10.1038/d41586-020-02140-8
References
Yu, L. et al. Preprint at medRxiv https://doi.org/10.1101/2020.02.20.20025874 (2020).
Yu, L. et al. Clin. Chem. https://doi.org/10.1093/clinchem/hvaa102 (2020).
Schmid-Burgk, J. L. et al. Preprint at bioRxiv https://doi.org/10.1101/2020.04.06.025635 (2020)
Joung, J. et al. Preprint at medRxiv https://doi.org/10.1101/2020.05.04.20091231 (2020).
Gootenberg, J. S. et al. Science 356, 438–442 (2017).
Broughton, J. P. et al. Nature Biotechnol. 38, 870–874 (2020).
Chen, J. S. et al. Science 360, 436–439 (2018).
Ackerman, C. M. et al. Nature 582, 277–282 (2020).

 Why more coronavirus testing won’t automatically help the hardest hit
Why more coronavirus testing won’t automatically help the hardest hit
 Scientists baffled by decision to stop a pioneering coronavirus testing project
Scientists baffled by decision to stop a pioneering coronavirus testing project
 The sprint to solve coronavirus protein structures — and disarm them with drugs
The sprint to solve coronavirus protein structures — and disarm them with drugs