- NEWS AND VIEWS
How DNA and RNA subunits might have formed to make the first genetic alphabet
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 582, 33-34 (2020)
doi: https://doi.org/10.1038/d41586-020-01566-4
References
Joyce, G. F. & Szostak, J. W. Cold Spring Harb. Perspect. Biol. 10, a034801 (2018).
Gavette, J. V., Stoop, M., Hud, N. V. & Krishnamurthy, R. Angew. Chem. Int. Edn 55, 13204–13209 (2016).
Xu, J. et al. Nature 582, 60–66 (2020).
Powner, M. W., Gerland, B. & Sutherland, J. D. Nature 459, 239–242 (2009).
Patel, B. H., Percivalle, C., Ritson, D. J., Duffy, C. D. & Sutherland, J. D. Nature Chem. 7, 301–307 (2015).
Teichert, J. S., Kruse, F. M. & Trapp, O. Angew. Chem. Int. Edn 58, 9944–9947 (2019).
Becker, S. et al. Science 366, 76–82 (2019).
Yadav, M., Kumar, R. & Krishnamurthy, R. Chem. Rev. https://doi.org/10.1021/acs.chemrev.9b00546 (2020).
Hein, J. E., Tse, E. & Blackmond, D. G. Nature Chem. 3, 704–706 (2011).

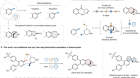
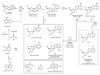 Read the paper: Selective prebiotic formation of RNA pyrimidine and DNA purine nucleosides
Read the paper: Selective prebiotic formation of RNA pyrimidine and DNA purine nucleosides
 The rocky road to biomolecules
The rocky road to biomolecules
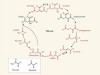 A possible non-biological reaction framework for metabolic processes on early Earth
A possible non-biological reaction framework for metabolic processes on early Earth