- NEWS AND VIEWS
AI tracks a beating heart’s function over time
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 580, 192-194 (2020)
doi: https://doi.org/10.1038/d41586-020-00819-6
References
Martino, T. A. & Young, M. E. J. Biol. Rhythms 30, 183–205 (2015).
Ouyang, D. et al. Nature 580, 252–256 (2020).
Pellikka, P. A. et al. JAMA Netw. Open 1, e181456 (2018).
Lang, R. M. et al. Eur. Heart J. Cardiovasc. Imaging 16, 233–270 (2015).
Morillo, C. A., Banerjee, A., Perel, P., Wood, D. & Jouven, X. J. Geriatr. Cardiol. 14, 195–203 (2017).
Santhanakrishnan, R. et al. Circulation 133, 484–492 (2016).
Zhang, J. et al. Circulation 138, 1623–1635 (2018).
Ghorbani, A. et al. NPJ Digit. Med. 3, 10 (2020).
Ji, S., Xu, W., Yang, M. & Kai, Y. IEEE Trans. Pattern Anal. Mach. Intell. 35, 221–231 (2013).
Tran, D. et al. Proc. IEEE Comput. Soc. Conf. Comput. Vision Pattern Recog. 6450–6459 (2018).
Rahman, S. A. & Adjeroh, D. A. Proc. IEEE Int. Conf. Bioinform. Biomed. 1100–1103 (2019); https://doi.org/10.1109/BIBM47256.2019.8983251
Dolz, J., Desrosiers, C. & Ayed, I. B. NeuroImage 170, 456–470 (2018).
Voigt, J.-U. et al. J. Am. Soc. Echocardiogr. 28, 183–193 (2015).
Suinesiaputra, A., McCulloch, A. D., Nash, M. P., Pontre, B. & Young, A. A. Med. Image Anal. 33, 38–43 (2016).
Varano, V. et al. Med. Image Anal. 46, 35–56 (2018).
Bernardis, E. et al. IEEE Trans. Biomed. Eng. 65, 733–744 (2018).
Omar, A. M. S. et al. JACC Cardiovasc. Imaging 10, 1291–1303 (2017).
Cho, J. S. et al. JACC Cardiovasc Imaging https://doi.org/10.1016/j.jcmg.2020.02.008 (2020).
Competing Interests
P.P.S. is an adviser for HeartSciences, Ultromics and Kencor Health, and has received research grants and support from HeartSciences, Hitachi Aloka, EchoSense and TomTec.

 Read the paper: Video-based AI for beat-to-beat assessment of cardiac function
Read the paper: Video-based AI for beat-to-beat assessment of cardiac function
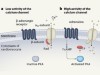 Suspect that modulates the heartbeat is ensnared
Suspect that modulates the heartbeat is ensnared
 Deep learning detects impending organ injury in the clinic
Deep learning detects impending organ injury in the clinic