- NEWS AND VIEWS
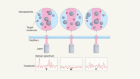
Snapshots of vibrating molecules
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 568, 36-37 (2019)
doi: https://doi.org/10.1038/d41586-019-00987-0
References
Steidtner, J. & Pettinger, B. Phys. Rev. Lett. 100, 236101 (2008).
Zhang, R. et al. Nature 498, 82–86 (2013).
Lee, J., Crampton, K. T., Tallarida, N. & Apkarian, V. A. Nature 568, 78–82 (2019).
Le Ru, E. C. & Etchegoin, P. G. Surface-Enhanced Raman Spectroscopy and Related Plasmonic Effects (Elsevier, 2009).
Sonntag, M. D. et al. J. Phys. Chem. C 116, 478–483 (2012).
Le Ru, E. C. & Etchegoin, P. G. Annu. Rev. Phys. Chem. 63, 65–87 (2012).
Stöckle, R. M., Suh, Y. D., Deckert, V. & Zenobi, R. Chem. Phys. Lett. 318, 131–136 (2000).
Pettinger, B., Ren, B., Picardi, G., Schuster, R. & Ertl, G. Phys. Rev. Lett. 92, 096101 (2004).
Duan, S. et al. J. Am. Chem. Soc. 137, 9515–9518 (2015).
Pozzi, E. A. et al. J. Phys. Chem. Lett. 5, 2657–2661 (2014).

 Read the paper: Visualizing vibrational normal modes of a single molecule with atomically confined light
Read the paper: Visualizing vibrational normal modes of a single molecule with atomically confined light
 A larger palette for biological imaging
A larger palette for biological imaging
 Optical spectroscopy goes intramolecular
Optical spectroscopy goes intramolecular