- NEWS AND VIEWS
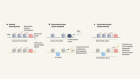
Modification of histone proteins by serotonin in the nucleus
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 567, 464-465 (2019)
doi: https://doi.org/10.1038/d41586-019-00532-z
References
Borrelli, E., Nestler, E. J., Allis, C. D. & Sassone‑Corsi, P. Neuron 60, 961–974 (2008).
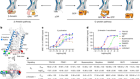
Farrelly, L. A. et al. Nature https://doi.org/10.1038/s41586-019-1024-7 (2019).
Daubert, E. A. & Condron, B. G. Trends Neurosci. 33, 424–434 (2010).
Holloway, T. & Gonzalez-Maeso, J. ACS Chem. Neurosci. 6, 1099–1109 (2015).
Watts, S. W., Priestley, J. R. & Thompson, J. M. PLoS ONE 4, e5682 (2009).
Paulmann, N. et al. PLoS Biol. 7, e1000229 (2009).
Walther, D. J. et al. Cell 115, 851–862 (2003).
Sileno, S. et al. J. Proteomics 96, 314–327 (2014).
Csaba, G. & Kovacs, P. Cell Biol. Int. 30, 861–865 (2006).
Tessarz, P. & Kouzarides, T. Nature Rev. Mol. Cell Biol. 15, 703–708 (2014).
Greer, E. L. & Shi, Y. Nature Rev. Genet. 13, 343–357 (2012).
Karmodiya, K. et al. BMC Genom. 13, 424 (2012).

 Read the paper: Histone serotonylation is a permissive modification that enhances TFIID binding to H3K4me3
Read the paper: Histone serotonylation is a permissive modification that enhances TFIID binding to H3K4me3
 Chromosomes come together to help mice distinguish odours
Chromosomes come together to help mice distinguish odours
 Protein modification fine-tunes the cell’s force producers
Protein modification fine-tunes the cell’s force producers