- NEWS AND VIEWS
Unstable genomes promote inflammation
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 567, 41-42 (2019)
doi: https://doi.org/10.1038/d41586-019-00510-5
References
McNairn, A. J., Chuang, C.-H., Bloom, J. C., Wallace, M. D. & Schimenti, J. C. Nature 567, 105–108 (2019).
Fragkos, M., Ganier, O., Coulombe, P. & Méchali, M. Nature Rev. Mol. Cell Biol. 16, 360–374 (2015).
Técher, H., Koundrioukoff, S., Nicolas, A. & Debatisse, M. Nature Rev. Genet. 18, 535–550 (2017).
Chuang, C.-H., Wallace, M. D., Abratte, C., Southard, T. & Schimenti, J. C. PLoS Genet. 6, e1001110 (2010).
Shima, N. et al. Nature Genet. 39, 93–98 (2007).
Rodier, F. et al. Nature Cell Biol. 11, 973–979 (2009).
Härtlova, A. et al. Immunity 42, 332–343 (2015).
Constanzo, V., Bardelli, A., Siena, S. & Abrignani, S. Open Biol. 8, 180081 (2018).
Zeman, M. K. & Cimprich, K. A. Nature Cell Biol. 16, 2–9 (2014).
Luke-Glaser, S., Luke, B., Grossi, S. & Constantinou, A. EMBO J. 29, 795–805 (2010).
Mackenzie, K. J. et al. Nature 548, 461–465 (2017).
Harding, S. M. et al. Nature 548, 466–470 (2017).
Luo, Y. et al. PLoS Genet. 10, e1004471 (2014).
Coquel, F. et al. Nature 557, 57–61 (2018).
Wang, R.-F. & Wang, H. Y. Cell Res. 27, 11–37 (2017).
Zhang, C.-Z. et al. Nature 522, 179–184 (2015).

 Read the paper: Female-biased embryonic death from inflammation induced by genomic instability
Read the paper: Female-biased embryonic death from inflammation induced by genomic instability
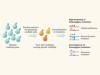 One rogue agent suffices for genomic chaos
One rogue agent suffices for genomic chaos
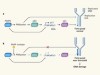 Superfast DNA replication causes damage in cancer cells
Superfast DNA replication causes damage in cancer cells