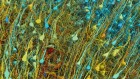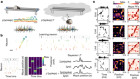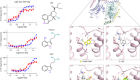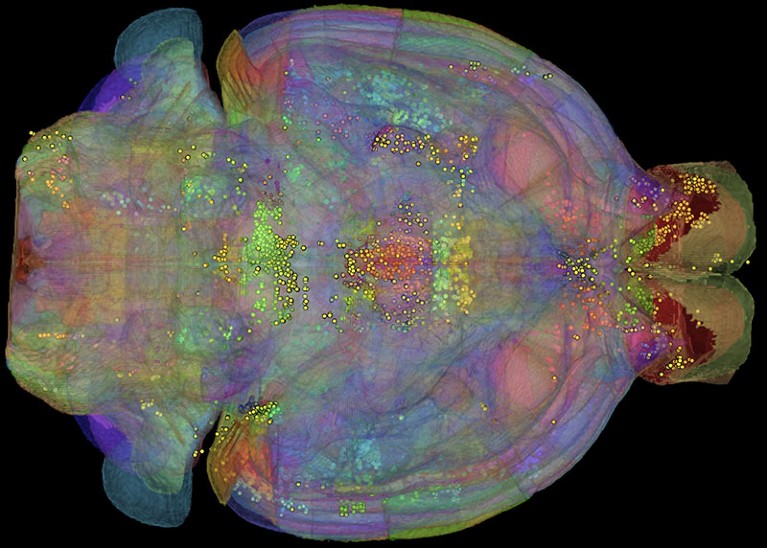
Neuroimaging data sets contain vast amounts of information.Credit: J. T. Vogelstein et al.
Randal Burns recalls that the brain-science community was “abuzz” in 2011. Burns, a computer scientist at Johns Hopkins University in Baltimore, Maryland, was focusing on astrophysics and fluid dynamics data management at the time. But he was intrigued when Joshua Vogelstein, a neuroscientist and colleague at Johns Hopkins, told him that the first large-scale neural-connectivity data sets had just been collected and asked for his help to present them online.
“It was the first time that you had data of that quality, at that resolution and scale, where you had the sense that you could build a neural map of an interesting portion of the brain,” says Burns.
Vogelstein worked with Burns to build a system that would make those data — 20 trillion voxels’ worth — available to the larger neuroscience community. The team has now generalized the software to support different classes of imaging data and describes the system this week (J. T. Vogelstein et al. Nature Meth. 15, 846-847; 2018).
NeuroData is a free, cloud-based collection of web services that supports large-scale neuroimaging data, from electron microscopy to magnetic resonance imaging and fluorescence photomicrographs.
Key to its functionality, Vogelstein says, is the spatial database bossDB, which allows researchers to retrieve images of any section of the brain, at any resolution, and in several standard formats. Users can then explore those data using a tool known as Neuroglancer. As they navigate the images, the URL changes to reflect their specific view, allowing them to share particular visualizations with their colleagues. “These links become a core part of the way in which we communicate and pass data back and forth to one another,” says Forrest Collman, a neuroscientist at the Allen Institute for Brain Science in Seattle, Washington, and a co-author of the paper.
But data browsing “is not where the science happens”, Burns says. “The science happens by doing a statistical analysis of the images.” To that end, NeuroData also includes tools for machine learning, image analysis and 3D volume rendering.
More than 100 public and private data sets have been deposited, Vogelstein says. Two of those came from Nelson Spruston, a neuroscientist at the Howard Hughes Medical Institute Janelia Research Campus in Ashburn, Virginia, who assembled 2D images into 3D brain reconstructions that are several terabytes in size. Because it can take weeks to upload that much data, Spruston’s team actually gave the information to NeuroData on a physical hard drive — an anachronistic, if practical solution.
Erik Bloss, a neuroscientist in Spruston’s laboratory, compiled those data sets to study neural synapses in the hippocampus. But researchers could mine the same data to, for instance, quantify the branching of neural arbors or the density of neural spines, Spruston says. “There’s all kinds of information in those data sets that in principle other people might be interested in,” he says.

 The Human Connectome Project’s neuroimaging approach
The Human Connectome Project’s neuroimaging approach
 Human brain mapped in unprecedented detail
Human brain mapped in unprecedented detail
 Connectomes make the map
Connectomes make the map