- OUTLOOK
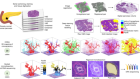
The unexpected role of histones in childhood brain cancer
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 561, S56-S58 (2018)
doi: https://doi.org/10.1038/d41586-018-06712-7
This article is part of Nature Outlook: Brain cancer, an editorially independent supplement produced with the financial support of third parties. About this content.
References
Faury, D. et al. J. Clin. Oncol. 25, 1196–1208 (2007).
Paugh, B. S. et al. J. Clin. Oncol. 28, 3061–3068 (2010).
Wu, G. et al. Nature Genet. 44, 251–253 (2012).
Schwartzentruber, J. et al. Nature 482, 226–231 (2012).
Mackay, A. et al. Cancer Cell 32, 520–537 (2017).
Grasso, C. S. et al. Nature Med. 21, 555–559 (2015).

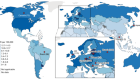
 Breaking down the epidemiology of brain cancer
Breaking down the epidemiology of brain cancer
 Immunotherapy offers a promising bet against brain cancer
Immunotherapy offers a promising bet against brain cancer
 See brain cancer as more than just the sum of biology
See brain cancer as more than just the sum of biology
 Getting cancer drugs into the brain
Getting cancer drugs into the brain
 How cerebral organoids are guiding brain-cancer research and therapies
How cerebral organoids are guiding brain-cancer research and therapies
 Searching for the roots of brain cancer
Searching for the roots of brain cancer
 Improving brain-cancer therapies through mathematical modelling
Improving brain-cancer therapies through mathematical modelling
 The genomics of brain cancer
The genomics of brain cancer
 The innovative therapies that could break the brain-cancer stalemate
The innovative therapies that could break the brain-cancer stalemate