- NEWS AND VIEWS
The telomerase enzyme and liver renewal
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 556, 181-182 (2018)
doi: https://doi.org/10.1038/d41586-018-02684-w
References
Palm, W. & de Lange, T. Annu. Rev. Genet. 41, 301–334 (2008).
Blasco, M. A. Nature Chem. Biol. 3, 640–649 (2007).
Günes, C. & Rudolph, K. L. Cell 152, 390–393 (2013).
Rudolph, K. L., Chang, S., Millard, M., Schreiber-Agus, N. & DePinho, R. A. Science 287, 1253–1258 (2000).
Calado, R. T. et al. PLoS ONE 4, e7926 (2009).
Lin, S. et al. Nature 556, 244–248 (2018).
Stanger, B. Z. Annu. Rev. Physiol. 77, 179–200 (2015).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Nature 524, 180–185 (2015).
Yanger, K. et al. Cell Stem Cell 15, 340–349 (2014).
Tarlow, B. D. et al. Cell Stem Cell. 15, 605–618 (2014).
Font-Burgada, J. et al. Cell 162, 766–779 (2015).
Raven, A. et al. Nature 547, 350–354 (2017).

 Read the paper: Distributed hepatocytes expressing telomerase repopulate the liver in homeostasis and injury
Read the paper: Distributed hepatocytes expressing telomerase repopulate the liver in homeostasis and injury
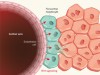 Maintaining liver mass
Maintaining liver mass
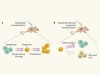 The versatile and plastic liver
The versatile and plastic liver