- NEWS AND VIEWS
A new era of rationally designed antipsychotics
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 555, 170-172 (2018)
doi: https://doi.org/10.1038/d41586-018-02328-z
References
Perala, J. et al. Arch. Gen. Psychiatry 64, 19−28 (2007).
Wang, S. et al. Nature 555, 269–273 (2018).
Kebabian, J. W. & Calne, D. B. Nature 277, 93–96 (1979).
Creese, I., Burt, D. R. & Snyder, S. H. Science 192, 481–483 (1976).
Seeman, P., Lee, T., Chau-Wong, M. & Wong, K. Nature 262, 717–719 (1976).
Sibley, D. R. & Monsma, F. J. Jr Trends Pharmacol. Sci. 13, 61–69 (1992).
Kapur, S. & Remington, G. Biol. Psychiatry 50, 873–883 (2001).
Keck, T. M., Burzynski, C., Shi, L. & Newman, A. H. Adv. Pharmacol. 69, 267–300 (2014).
Wang, S. et al. Science 358, 381–386 (2017).
Moritz, A. E., Free, R. B. & Sibley, D. R. Cell. Signal. 41, 75–81 (2018).
Chien, E. Y. et al. Science 330, 1091–1095 (2010).
Newman, A. H. et al. J. Med. Chem. 55, 6689–6699 (2012).
Löber, S., Hübner, H., Tschammer, N. & Gmeiner, P. Trends Pharmacol. Sci. 32, 148–157 (2011).
Carlsson, J. et al. Nature Chem. Biol. 7, 769–778 (2011).
Kapur, S. & Seeman, P. Am. J. Psychiatry 158, 360–369 (2001).
Sykes, D. A. et al. Nature Commun. 8, 763 (2017).
Seeman, P. Clin. Neurosci. Res. 1, 53–60 (2001).

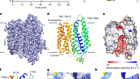
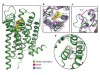 Read the paper: Structure of the D2 dopamine receptor bound to the atypical antipsychotic drug risperidone
Read the paper: Structure of the D2 dopamine receptor bound to the atypical antipsychotic drug risperidone
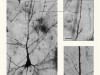 Pruning hypothesis comes of age
Pruning hypothesis comes of age
 Making risk-takers settle
Making risk-takers settle