- NEWS AND VIEWS
The origins of memory T cells
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 552, 337-339 (2017)
doi: https://doi.org/10.1038/d41586-017-08280-8
References
Akondy, R. S. et al. Nature 552, 362–367 (2017).
Youngblood, B. et al. Nature 552, 404–409 (2017).
Kaech, S. M. & Cui, W. Nature Rev. Immunol. 12, 749–761 (2012).
Winter, D. R., Jung, S. & Amit, I. Nature Rev. Immunol. 15, 585–594 (2015).
Gray, S. M., Kaech, S. M. & Staron, M. M. Immunol. Rev. 261, 157–168 (2014).
Chang, J. T., Wherry, E. J. & Goldrath, A. W. Nature Immunol. 15, 1104–1115 (2014).

 Read the paper: Origin and differentiation of human memory CD8 T cells after vaccination
Read the paper: Origin and differentiation of human memory CD8 T cells after vaccination
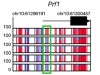 Read the paper: Effector CD8 T cells dedifferentiate into long-lived memory cells
Read the paper: Effector CD8 T cells dedifferentiate into long-lived memory cells
 The patterns of T-cell target recognition
The patterns of T-cell target recognition
 Memory beyond immunity
Memory beyond immunity