Abstract
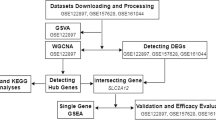
The rupture of intracranial aneurysm (IA) is the leading cause for devastating subarachnoid hemorrhage. This study aimed to investigate genes related to IA and potential diagnosis targets. Two data sets (GSE15629 and GSE54083) were downloaded from Gene Expression Omnibus database. GSE15629 contained eight RI (ruptured IA), six UI (unruptured IA) and five control IA samples. GSE54083 included 8 RI, 5 UI and 10 superficial temporal artery samples. In total, 452 differentially expressed genes (DEGs) between RI and control, and 570 DEGs between UI and control, were identified. Protein–protein interaction networks for two kinds of DEGs related to RI and UI were constructed, respectively. Module networks were searched for DEGs related to RI or UI based on WGCNA (weighted gene coexpression network analysis). In the significant modules, FOS, CCL2, COL4A2 and CXCL5 were screened as crucial nodes with high degrees. Among them, FOS and CCL2 were enriched in immune response and COL4A2 was involved in the ECM (extracellular matrix) pathway, whereas CXCL5 was related to cytokine–cytokine receptor pathway. Taken together, FOS, CCL2, COL4A2 and CXCL5 might participate in the pathogenesis of RI or UI, and could serve as potential diagnosis targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Westerlaan HE, Van Dijk J, Jansen-van der Weide MC, de Groot JC, Groen RJ, Mooij JJA et al. Intracranial aneurysms in patients with subarachnoid hemorrhage: CT angiography as a primary examination tool for diagnosis—systematic review and meta-analysis 1. Radiology 2011; 258: 134–145.
Vlak MH, Algra A, Brandenburg R, Rinkel GJ . Prevalence of unruptured intracranial aneurysms, with emphasis on sex, age, comorbidity, country, and time period: a systematic review and meta-analysis. Lancet Neurol 2011; 10: 626–636.
Risselada R, De Vries L, Dippel D, van Kooten F, van der Lugt A, Niessen W et al. Incidence, treatment, and case-fatality of non-traumatic subarachnoid haemorrhage in the Netherlands. Clin Neurol Neurosurg 2011; 113: 483–487.
Nakaoka H, Tajima A, Yoneyama T, Hosomichi K, Kasuya H, Mizutani T et al. Gene expression profiling reveals distinct molecular signatures associated with the rupture of Intracranial aneurysm. Stroke 2014; 45: 2239–2245.
Lylyk P, Ferrario A, Pabón B, Miranda C, Doroszuk G . Buenos Aires experience with the Neuroform self-expanding stent for the treatment of intracranial aneurysms. J Neurosurg 2005; 102: 235–241.
Weinsheimer S, Lenk GM, Van der Voet M, Land S, Ronkainen A, Alafuzoff I et al. Integration of expression profiles and genetic mapping data to identify candidate genes in intracranial aneurysm. Physiol Genom 2007; 32: 45–57.
Krischek B, Kasuya H, Tajima A, Akagawa H, Sasaki T, Yoneyama T et al. Network-based gene expression analysis of intracranial aneurysm tissue reveals role of antigen presenting cells. Neuroscience 2008; 154: 1398–1407.
Shi C, Awad IA, Jafari N, Lin S, Du P, Hage ZA et al. Genomics of human intracranial aneurysm wall. Stroke 2009; 40: 1252–1261.
Pera J, Korostynski M, Krzyszkowski T, Czopek J, Slowik A, Dziedzic T et al. Gene expression profiles in human ruptured and unruptured intracranial aneurysms. What is the role of inflammation? Stroke 2010; 41: 224–231.
Shabalin AA, Tjelmeland H, Fan C, Perou CM, Nobel AB . Merging two gene-expression studies via cross-platform normalization. Bioinformatics 2008; 24: 1154–1160.
Rudy J, Valafar F . Empirical comparison of cross-platform normalization methods for gene expression data. BMC Bioinform 2011; 12: 467.
Benjamini Y, Hochberg Y . Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 1995; 57: 289–300.
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A et al. STRING v9. 1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 2013; 41: D808–D815.
Kohl M, Wiese S, Warscheid B . Cytoscape: software for visualization and analysis of biological networks. Methods Mol Biol 2011; 696: 291–303.
Ruan J, Dean AK, Zhang W . A general co-expression network-based approach to gene expression analysis: comparison and applications. BMC Syst Biol 2010; 4: 8.
Langfelder P, Horvath S . WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 2008; 9: 559.
Li S, Fu H, Wang Y, Tie Y, Xing R, Zhu J et al. MicroRNA− 101 regulates expression of the v− fos FBJ murine osteosarcoma viral oncogene homolog (FOS) oncogene in human hepatocellular carcinoma. Hepatology 2009; 49: 1194–1202.
Tao T, Wang Y, Luo H, Yao L, Wang L, Wang J et al. Involvement of FOS-mediated miR-181b/miR-21 signalling in the progression of malignant gliomas. Eur J Cancer 2013; 49: 3055–3063.
Walter I, Wolfesberger B, Miller I, Mair G, Burger S, Gallè B et al. Human osteosarcoma cells respond to sorafenib chemotherapy by downregulation of the tumor progression factors S100A4, CXCR4 and the oncogene FOS. Oncol Rep 2014; 31: 1147–1156.
Langer S, Singer C, Hudelist G, Dampier B, Kaserer K, Vinatzer U et al. Jun and Fos family protein expression in human breast cancer: correlation of protein expression and clinicopathological parameters. Eur J Gynaecol Oncol 2005; 27: 345–352.
Shaulian E, Karin M . AP-1 in cell proliferation and survival. Oncogene 2001; 20: 2390–2400.
Wagner EF, Eferl R . Fos/AP− 1 proteins in bone and the immune system. Immunol Rev 2005; 208: 126–140.
Gentilini D, Perino A, Viganò P, Chiodo I, Cucinella G, Vignali M et al. Gene expression profiling of peripheral blood mononuclear cells in endometriosis identifies genes altered in non-gynaecologic chronic inflammatory diseases. Hum Reprod 2011; 26: 3109–17.
Choi HJ, Yun HS, Kang HJ, Ban H-J, Kim Y, Nam H-Y et al. Human transcriptome analysis of acute responses to glucose ingestion reveals the role of leukocytes in hyperglycemia-induced inflammation. Physiol Genom 2012; 44: 1179–1187.
Marinković G, Hibender S, Hoogenboezem M, van Broekhoven A, Girigorie AF, Bleeker N et al. Immunosuppressive drug azathioprine reduces aneurysm progression through inhibition of Rac1 and c-Jun-terminal-N-kinase in endothelial cells. Arterioscler Thromb Vasc Biol 2013; 33: 2380–2388.
Kanematsu Y, Kanematsu M, Kurihara C, Tada Y, Tsou T-L, van Rooijen N et al. Critical roles of macrophages in the formation of intracranial aneurysm. Stroke 2011; 42: 173–178.
Zhang L, Zhao J, Kuboyama N, Abiko Y . Low-level laser irradiation treatment reduces CCL2 expression in rat rheumatoid synovia via a chemokine signaling pathway. Lasers Med Sci 2011; 26: 707–717.
Chalouhi N, Ali MS, Jabbour PM, Tjoumakaris SI, Gonzalez LF, Rosenwasser RH et al. Biology of intracranial aneurysms: role of inflammation. J Cerebr Blood Flow Metab 2012; 32: 1659–1676.
Gunda B, Mine M, Kovács T, Hornyák C, Bereczki D, Várallyay G et al. COL4A2 mutation causing adult onset recurrent intracerebral hemorrhage and leukoencephalopathy. J Neurol 2014; 261: 500–503.
Kurban G, Duplan E, Ramlal N, Hudon V, Sado Y, Ninomiya Y et al. Collagen matrix assembly is driven by the interaction of von Hippel–Lindau tumor suppressor protein with hydroxylated collagen IV alpha 2. Oncogene 2007; 27: 1004–1012.
Peters DG, Kassam AB, Feingold E, Heidrich-O’Hare E, Yonas H, Ferrell RE et al. Molecular anatomy of an intracranial aneurysm coordinated expression of genes involved in wound healing and tissue remodeling. Stroke 2001; 32: 1036–1042.
Tazzyman S, Niaz H, Murdoch C . Neutrophil-mediated tumour angiogenesis: subversion of immune responses to promote tumour growth. Semin Cancer Biol 2013; 23: 149–158.
Li A, Strieter R, Sugi M, Reber H, Eibl G, Hines O . Cxcl5 [Ena-78] mediates pancreatic cancer derived angiogenesis. Pancreas 2008; 37: 479.
Beitelshees AL, Aquilante CL, Allayee H, Langaee TY, Welder GJ, Schofield RS et al. CXCL5 polymorphisms are associated with variable blood pressure in cardiovascular disease-free adults. Hum Genom 2012; 6: 9.
Yan J, Tingey C, Lyde R, Gorham T, Choo D, Muthumani A et al. Novel and enhanced anti-melanoma DNA vaccine targeting the tyrosinase protein inhibits myeloid-derived suppressor cells and tumor growth in a syngeneic prophylactic and therapeutic murine model. Cancer Gene Ther 2014; 21: 507–517.
Nowicki KW, Hosaka K, He Y, McFetridge PS, Scott EW, Hoh BL . Novel high-throughput in vitro model for identifying hemodynamic-induced inflammatory mediators of cerebral aneurysm formation. Hypertension 2014; 64: 1306–1313.
Kalwitz G, Endres M, Neumann K, Skriner K, Ringe J, Sezer O et al. Gene expression profile of adult human bone marrow-derived mesenchymal stem cells stimulated by the chemokine CXCL7. Int J Biochem Cell Biol 2009; 41: 649–658.
Skirgaudas M, Awad IA, Kim J, Rothbart D, Criscuolo G . Expression of angiogenesis factors and selected vascular wall matrix proteins in intracranial saccular aneurysms. Neurosurgery 1996; 39: 537–547.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Zheng, X., Xue, C., Luo, G. et al. Identification of crucial genes in intracranial aneurysm based on weighted gene coexpression network analysis. Cancer Gene Ther 22, 238–245 (2015). https://doi.org/10.1038/cgt.2015.10
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cgt.2015.10
This article is cited by
-
A Transcriptomic Comparative Study of Cranial Vasculature
Translational Stroke Research (2023)
-
Weighted gene co-expression network analysis revealed key biomarkers associated with the diagnosis of hypertrophic cardiomyopathy
Hereditas (2020)
-
Identification of key gene modules for human osteosarcoma by co-expression analysis
World Journal of Surgical Oncology (2018)
-
Gene co-expression network reconstruction: a review on computational methods for inferring functional information from plant-based expression data
Plant Biotechnology Reports (2017)