Abstract
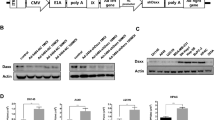
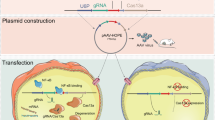
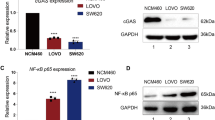
Tumor cells secrete a variety of cytokines to outgrow and evade host immune surveillance. In this context, transforming growth factor-β1 (TGF-β1) is an extremely interesting cytokine because it has biphasic effects in cancer cells and normal cells. TGF-β1 acts as a growth inhibitor in normal cells, whereas it promotes tumor growth and progression in tumor cells. Overexpression of TGF-β1 in tumor cells also provides additional oncogenic activities by circumventing the host immune surveillance. Therefore, this study ultimately aimed to test the hypothesis that suppression of TGF-β1 in tumor cells by RNA interference can have antitumorigenic effects. However, we demonstrated here that the interrelation between TGF-β isotypes should be carefully considered for the antitumor effect in addition to the selection of target sequences with highest efficacy. The target sequences were proven to be highly specific and effective for suppressing both TGF-β1 mRNA and protein expression in cells after infection with an adenovirus expressing TGF-β1 short hairpin RNA (shRNA). A single base pair change in the shRNA sequence completely abrogated the suppressive effect on TGF-β1. Surprisingly, the suppression of TGF-β1 induced TGF-β3 upregulation, and the suppression of TGF-β2 induced another unexpected downregulation of both TGF-β1 and TGF-β3. Taken together, this information may prove useful when considering the design for a novel cancer immunogene therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Abbreviations
- TGF-β:
-
transforming growth factor-β
- shRNA:
-
short hairpin RNA
- GFP:
-
green fluorescence protein
- ELISA:
-
enzyme-linked immunosorbent assay
References
Massague J . The TGF-beta family of growth and differentiation factors. Cell 1987; 49: 437–438.
Moore LD, Isayeva T, Siegal GP, Ponnazhagan S . Silencing of transforming growth factor-beta1 in situ by RNA interference for breast cancer: implications for proliferation and migration in vitro and metastasis in vivo. Clinical Cancer Res 2008; 14: 4961–4970.
Annes JP, Munger JS, Rifkin DB . Making sense of latent TGFbeta activation. J Cell Sci 2003; 116 (Part 2): 217–224.
Massague J . TGFbeta in Cancer. Cell 2008; 134: 215–230.
Pardali K, Moustakas A . Actions of TGF-beta as tumor suppressor and pro-metastatic factor in human cancer. Biochim Biophys Acta 2007; 1775: 21–62.
Massague J, Blain SW, Lo RS . TGFbeta signaling in growth control, cancer, and heritable disorders. Cell 2000; 103: 295–309.
Elliott RL, Blobe GC . Role of transforming growth factor beta in human cancer. J Clin Oncol 2005; 23: 2078–2093.
Kim S, Buchlis G, Fridlender ZG, Sun J, Kapoor V, Cheng G et al. Systemic blockade of transforming growth factor-beta signaling augments the efficacy of immunogene therapy. Cancer Res 2008; 68: 10247–10256.
Ivanovic V, Demajo M, Krtolica K, Krajnovic M, Konstantinovic M, Baltic V et al. Elevated plasma TGF-beta1 levels correlate with decreased survival of metastatic breast cancer patients. Clin Chim Acta 2006; 371: 191–193.
Conroy H, Galvin KC, Higgins SC, Mills KH . Gene silencing of TGF-beta1 enhances antitumor immunity induced with a dendritic cell vaccine by reducing tumor-associated regulatory T cells. Cancer Immunol Immunother 2012; 61: 425–431.
Wei H, Liu P, Swisher E, Yip YY, Tse JH, Agnew K et al. Silencing of the TGF-beta1 gene increases the immunogenicity of cells from human ovarian carcinoma. J Immunother 2012; 35: 267–275.
Ding Y, Kim JK, Kim SI, Na HJ, Jun SY, Lee SJ et al. TGF-{beta}1 protects against mesangial cell apoptosis via induction of autophagy. J Biol Chem 2010; 285: 37909–37919.
Dogar AM, Towbin H, Hall J . Suppression of latent transforming growth factor (TGF)-beta1 restores growth inhibitory TGF-beta signaling through microRNAs. J Biol Chem 2011; 286: 16447–16458.
Schafer H, Struck B, Feldmann EM, Bergmann F, Grage-Griebenow E, Geismann C et al. TGF-beta1-dependent L1CAM expression has an essential role in macrophage-induced apoptosis resistance and cell migration of human intestinal epithelial cells. Oncogene 2013 doi:10.1038/onc.2012.44; epub ahead of print 20 February 2012.
Almeida R, Allshire RC . RNA silencing and genome regulation. Trends Cell Biol 2005; 15: 251–258.
Aagaard L, Rossi JJ . RNAi therapeutics: principles, prospects and challenges. Adv Drug Deliv Rev 2007; 59: 75–86.
Whitehead KA, Langer R, Anderson DG . Knocking down barriers: advances in siRNA delivery. Nature reviews. Drug Discov 2009; 8: 129–138.
Zhang H, Kolb FA, Jaskiewicz L, Westhof E, Filipowicz W . Single processing center models for human dicer and bacterial RNase III. Cell 2004; 118: 57–68.
Rand TA, Ginalski K, Grishin NV, Wang X . Biochemical identification of Argonaute 2 as the sole protein required for RNA-induced silencing complex activity. Proc Natl Acad Sci USA 2004; 101: 14385–14389.
Kumar LD, Clarke AR . Gene manipulation through the use of small interfering RNA (siRNA): from in vitro to in vivo applications. Adv Drug Deliv Rev 2007; 59: 87–100.
Sharma A, Tandon M, Bangari DS, Mittal SK . Adenoviral vector-based strategies for cancer therapy. Curr Drug Ther 2009; 4: 117–138.
Douglas JT . Adenoviral vectors for gene therapy. Mol Biotechnol 2007; 36: 71–80.
Schlingensiepen KH, Jaschinski F, Lang SA, Moser C, Geissler EK, Schlitt HJ et al. Transforming growth factor-beta 2 gene silencing with trabedersen (AP 12009) in pancreatic cancer. Cancer Sci 2011; 102: 1193–1200.
Olivares J, Kumar P, Yu Y, Maples PB, Senzer N, Bedell C et al. Phase I trial of TGF-beta 2 antisense GM-CSF gene-modified autologous tumor cell (TAG) vaccine. Clin Cancer Res 2011; 17: 183–192.
De Crescenzo G, Hinck CS, Shu Z, Zuniga J, Yang J, Tang Y et al. Three key residues underlie the differential affinity of the TGFbeta isoforms for the TGFbeta type II receptor. J Mol Biol 2006; 355: 47–62.
Laverty HG, Wakefield LM, Occleston NL, O’Kane S, Ferguson MW . TGF-beta3 and cancer: a review. Cytokine Growth Factor Rev 2009; 20: 305–317.
Hinck AP, Archer SJ, Qian SW, Roberts AB, Sporn MB, Weatherbee JA et al. Transforming growth factor beta 1: three-dimensional structure in solution and comparison with the X-ray structure of transforming growth factor beta 2. Biochemistry 1996; 35: 8517–8534.
Bocharov EV, Blommers MJ, Kuhla J, Arvinte T, Burgi R, Arseniev AS . Sequence-specific 1H and 15N assignment and secondary structure of transforming growth factor beta3. J Biomol NMR 2000; 16: 179–180.
Akhurst RJ, Lehnert SA, Faissner A, Duffie E . TGF beta in murine morphogenetic processes: the early embryo and cardiogenesis. Development 1990; 108: 645–656.
Bierie B, Moses HL . Transforming growth factor beta (TGF-beta) and inflammation in cancer. Cytokine Growth Factor Rev 2010; 21: 49–59.
Li C, Wang J, Wilson PB, Kumar P, Levine E, Hunter RD et al. Role of transforming growth factor beta3 in lymphatic metastasis in breast cancer. Int J Cancer 1998; 79: 455–459.
Acknowledgements
This work was supported by the Industrial Strategic Technology Development program (10035562: Development of nucleic acid-based anticancer drugs overcoming the immunotherapy resistance) funded by the Ministry of Knowledge Economy (MKE, Korea). This work was also supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) and funded by the Ministry of Education. Science and Technology (2012-0002108). S Oh and E Kim are funded by the Brain Korea 21 project for Medical Science, Yonsei University, College of Medicine, Seoul, South Korea.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Cancer Gene Therapy website
Rights and permissions
About this article
Cite this article
Oh, S., Kim, E., Kang, D. et al. Transforming growth factor-β gene silencing using adenovirus expressing TGF-β1 or TGF-β2 shRNA. Cancer Gene Ther 20, 94–100 (2013). https://doi.org/10.1038/cgt.2012.90
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cgt.2012.90
Keywords
This article is cited by
-
Immunotherapy for lung cancer combining the oligodeoxynucleotides of TLR9 agonist and TGF-β2 inhibitor
Cancer Immunology, Immunotherapy (2023)
-
Bmi1 drives hepatocarcinogenesis by repressing the TGFβ2/SMAD signalling axis
Oncogene (2020)
-
TGF-β downregulation-induced cancer cell death is finely regulated by the SAPK signaling cascade
Experimental & Molecular Medicine (2018)
-
Naringenin prevents TGF-β1 secretion from breast cancer and suppresses pulmonary metastasis by inhibiting PKC activation
Breast Cancer Research (2016)
-
Oxidative stress mediates the conversion of endothelial cells into myofibroblasts via a TGF-β1 and TGF-β2-dependent pathway
Laboratory Investigation (2014)