Abstract
The tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL) is a potent inducer of tumor cell apoptosis, but concerns of considerable liver toxicity limit its uses in human cancer therapy. Here, we show that i.v. injected Escherichia coli DH5α (E. coli DH5α) specifically replicates in solid tumors and metastases in live animals. E. coli DH5α does not enter tumor cells and suits for being the vector for soluble TRAIL (sTRAIL), which induces apoptosis by activating cell-surface death receptors. With the high ‘tumor-targeting’ nature, we demonstrate that intratumoral (i.t.) and intravenous injection of sTRAIL-expressing E. coli DH5α results in the tumor-targeted release of biologically active molecules, which leads to a dramatic reduction in the tumor growth rate and the prolonged survival of tumor-bearing mice. TRAIL delivery by E. coli DH5α did not cause any detectable toxicity to any organs, suggesting that E. coli DH5α-delivered sTRAIL protein therapy may provide a feasible and effective form of treatment for solid tumors.
Similar content being viewed by others
Introduction
The tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL) is a type-II transmembrane protein that was initially identified based on homology of the extracellular domain to CD95 L, TNF and LTα.1, 2 The membrane-bound and the soluble extracellular domains of TRAIL (amino acids 95–281 or 114–281) induce apoptosis in a wide variety of tumor cell lines. Approximately two-thirds of tumor cell lines tested are sensitive to the cytotoxic effects of TRAIL in vitro,3, 4 suggesting that TRAIL may be exploited as a powerful antitumor agent. Repeated intravenous administration of recombinant, biologically active TRAIL (rTRAIL) can induce tumor cells apoptosis, suppress tumor progression and improve survival in tumor-bearing mice.5, 6 Unfortunately, the use of TRAIL as a therapeutic agent is limited by severe cytotoxicity in normal hepatocytes,7, 8 esophageal epithelial cells,9 prostate epithelial cells10 and keratinocytes11 in vitro, and its short half-life after systemic administration in vivo.12
Gene therapy may enable continuous production of TRAIL in substantial amounts for a relatively long period of time. This strategy would overcome the necessity for repeated injections of recombinant TRAIL and may also result in increased antitumor effect. The transfer and expression of TRAIL into cells by adenovirus or adeno-associated virus induces apoptosis and apoptotic bystander effects in several human cancer cells in vitro and in xenograft models of human tumors.13, 14, 15 However, adenovirus or adeno-associated virus-based vectors may cause hepatotoxicity through innate and cell-mediated immune responses, and have less targeting to the tumor site when systemically administrated.16, 17
It has been known for more than 60 years that anaerobic bacteria can selectively grow in tumors.18, 19, 20, 21 Several approaches to developing anaerobic bacteria for tumor therapies have been described. The anaerobic species Bifidobacterium longum22, 23 and Clostridum novyi24, 25 could selectively grow in the hypoxic environment of large solid tumors. Recently, Salmonella typhimurium with attenuated lipid-A was evaluated in a phase-I clinical trial.26, 27 To overcome toxicity, Zhao et al.28, 29, 30 mutagenized S. typhimurim and selected an amino-acid auxotrophic strain, which selectively grew in and killed tumor cells. Non-pathogenic Escherichia coli is a facultative anaerobic bacterium that naturally resides in the digestive tracts of humans and other animals. There have been no reports to date describing the use of E. coli as an anticancer protein-delivery agent to repress tumor growth in vivo.31, 32
Minimizing side effects while maximizing tumor targeting is the greatest challenge of current tumor therapeutic protocols. Here, we show that intravenously (i.v.) injected E. coli specifically replicates in solid tumors and metastases in live animals. With this high ‘tumor-targeting’ nature, we describe a novel treatment of injecting TRAIL-expressing E. coli so that it can grow loco-regionally inside tumors and release the biologically active soluble TRAIL (sTRAIL) protein at relatively high levels, thereby achieving maximal therapeutic effects while sparing potential systemic side effects.
Materials and methods
Cell lines and cell culture
Murine B16-F10-luc+ melanoma cells and human NCI-H460 lung tumor cells were obtained from Xenogen (Hopkinton, MA, USA) and ATCC (Rockville, MD, USA), respectively. B16-luc+ were grown in Dulbecco's modified Eagle's medium (Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum, 1% non-essential amino acids, L-glutamine, and sodium pyruvate (all from Hyclone, Ogden, UT, USA); MEM vitamin solution (Invitrogen); and 1% penicillin and streptomycin. NCI-H460 cells were grown in RPMI-1640 medium supplemented with 10% fetal bovine serum, and 1% penicillin and streptomycin. All the cells were cultured at 37 °C in a humidified atmosphere of 5% CO2. H460 cells were transfected with pcDNA3.1-Luc+ vector containing firefly luciferase using Lipofectamine 2000 (Invitrogen). Stable clones expressing luc were isolated in the presence of 300 μg ml−1 geneticin (G418) and the clone with the highest level of luc expression (as determined by bioluminescence) was selected by using D-luciferin and an In Vivo Imaging System (IVIS) (Xenogen).
Bacterial strains and plasmids
E. coli DH5α were routinely grown in LB medium and cultured in a shaker at 37 °C. The plasmid pMK luxABCDE, also called pXen-1 (Xenogen), which contains bacterial luciferase, was electroporated into DH5α and positive transformants were screened with an IVIS.33, 34 The clone with the highest bioluminescence was grown overnight at 37 °C in LB medium containing 100 μg ml−1 ampicillin. The bioluminescent clones were stored at −80 °C and thawed before use. Bacterial concentration was estimated spectrophotometrically by determining absorbance at 600 nm, and cell numbers were verified by plating dilutions of inoculum onto LB agar plate (1 OD≈8(108 c.f.u. ml−1).
A sense primer (5′-GCGGATCCGACCTCTGAGGAAACCATTTC-3′) and antisense primer (5′-CCGCTCGAGTTAGCCAACTAAAAAGGCCC-3′) were used to amplify a 561-bp DNA fragment representing the sTRAIL from mammary library by PCR. The products were gel-purified, digested with BamHI and XhoI restriction enzymes, repurified and inserted into the BamHI and XhoI sites of the prokaryotic expression vector pGEX-KG (Amersham Biosciences, Piscataway, NJ). The resulting vector pGEX-sTRAIL was electroporated into E. coli DH5α and the resulting clone was termed as E. coli DH5α (sTRAIL). Protein expression was induced by 0.1 mM isopropyl-β-thiogalacto pyranoside in culture and protein production was analyzed by Coomassie brillant blue (CBB) staining.
Recipient animals and tumor models
Female BALB/c and nude mice were housed under aseptic conditions in micro-isolator cages. The animals used in the studies were approximately 4–6 weeks of age and weight ranged between 20±5 and 18±2 g. All studies involving mice were approved by the institute's Animal Care and Use Committee.
B16-luc+ or H460-luc+ tumor cells (1 × 106 or 1 × 107, respectively) were injected subcutaneously into the dorsal flanks of the BALB/c and nude mice and tumor growth was monitored twice a week by in vivo imaging or external caliper measurement. Mice bearing subcutaneous melanoma tumors and subcutaneous lung tumors were grown about 7–10 days until the tumor size was approximately 200 mm3. A total of 5 × 105 B16 cells were injected into nude mice through the tail vein and extensive lung metastases occurred within 2 weeks after cellular implantation.
Bioluminescence imaging
B16-F10 melanoma and H460 lung carcinoma xenografts were established as above. Luminescent bacteria were grown and harvested at late logarithmic phase, washed and diluted with sterile normal saline and injected via the tail vein into tumor-bearing or non-tumor-bearing mice at various doses ranging from 1 × 106 to 1 × 1010 c.f.u. per mouse. To analyze bacterial luciferase activity and trace bioluminescent bacteria, luminescence was quantified from the in vivo signals emitted from the dorsal or ventral views of each mouse prior to killing and from ex vivo images taken of excised tissue immediately after being killed. Total photon emission from different parts within the images of each mouse was quantified at different time points using the Living Image software (Xenogen).
For recording tumor growth by in vivo imaging, the animals were injected intraperitoneally with D-luciferin at 150 mg kg−1 in Dulbecoo's phosphate buffered saline and anesthetized with 1–3% isoflurane. The mice were then placed on a warmed stage inside a light-tight camera box with continuous exposure to 1–3% isoflurane. Imaging time ranged from 1 s to 3 min, depending on the tumor model and time points. Low levels of light emitted from the bioluminescent tumors were detected by an IVIS, and were integrated, digitized and quantified as photons/second using the Living Image software.
Cytotoxicity assay
H460 cells were seeded in 24-well plates at a density of 1 × 105 cells per well in 500 μl of complete culture medium. After 12–18 h of adherence, the cells were treated with E. coli DH5α (sTRAIL) and E. coli DH5α (empty vector) at various multiplicities of infection (MOIs) (50:1, 100:1, 200:1 and 400:1) in fresh culture medium containing 5% penicillin and streptomycin to inhibit bacterial growth. Cell morphology was observed by light microscopy after 12 h of incubation. For blocking assay, H460 cells were incubated with 0 to 200 ng of death receptor4 (DR4):Fc or TNF receptor:Fc (both from Alexis Biochemicals, San Diego, CA, USA) per milliliter in addition to E. coli DH5α (sTRAIL) or E. coli DH5α (empty vector) at an MOI of 100:1 in the same setting, and the results were documented as photographs. Unless otherwise specified, E. coli DH5α (empty vector) was used as vector control and the cell culture medium as mock control.
Cells plated in 12-well dishes were infected with E. coli DH5α (sTRAIL) at different MOIs and stained with crystal violet. The medium was removed 12 h after co-incubation and the cells were fixed for 3 min in 4% paraformaldehyde at room temperature prior to crystal violet staining. Fixed cells were washed with phosphate-buffered saline and incubated for 3 min in 1% crystal violet in 70% ethanol. Cells were rinsed three times with water, air-dried and photographed. Cells in a duplicate plate were trypsinized and stained with trypan blue and the percentage of viable cells was counted by light microscopy.
Flow cytometry
H460 cells were infected with E. coli DH5α (sTRAIL) at an MOI of 200 for 2, 4 or 8 h. The cells were harvested by trypsinization, followed by washing with cold phosphate-buffered saline containing 10% fetal bovine serum and fixation in 70% ethanol. The cells were then stained with 10 μg ml−1 propidium iodide and analyzed using a FACScan instrument.
Bacterial infection
In order to study bacterial infection on non-tumor bearing or tumor-bearing mice in vivo, tissue samples were obtained from liver, spleen and tumors after injection as described above. Normal tissues and tumors were excised, weighed and c.f.u. were determined at different time points after homogenizing and plating supernatants on LB agar plates.
For intratumoral (i.t.) injection, E. coli DH5α (sTRAIL) induced by IPTG were harvested, washed and diluted with sterile normal saline. Bacteria were injected directly into the central area of the tumors at a dose of 1 × 109 c.f.u. per 50 μl of sterile normal saline on the first two successive days (total dose of 2 × 109 c.f.u.). For intravenous injection, a total of 5 × 107 c.f.u. of E. coli DH5α (sTRAIL) were injected into the tail vein of tumor-bearing nude mice. One week later, the second injection was performed as described as above. Unless otherwise specified, E. coli DH5α (empty vector) was used as a vector control, and sterile normal saline was used as a mock control.
PCR assay and tissue immunohistochemistry
Established H460 tumors were allowed to grow until they were approximately 200 mm3 in size. A total of 5 × 107 c.f.u. per mouse of E. coli DH5α (sTRAIL) was injected i.v. through the tail vein into H460 tumor-bearing nude mice. Four days after injection, animals were killed and tumors, spleens, livers and kidneys were aseptically removed, homogenized, diluted and plated onto LB agar plates supplemented with 50 μg ml−1 penicillin to determine the c.f.u. of E. coli DH5α (sTRAIL). Several clones were randomly picked and PCR was performed using the corresponding sTRAIL primers. At the same time, normal tissues and tumors from nude mice were fixed, paraffin-embedded and sectioned. Tissue immunohistochemistry were performed with an anti-hTRAIL monoclonal antibody (clone 2H12) purchased from R&D Systems (San Francisco, CA) using standard methods.
Western blot analysis
H460 tumor cells were lysed in buffer (25 mM Tris–Cl (pH 7.4), 50 mM NaCl, 0.5% Na-deoxycholate, 2% Nonidet P-40, 0.2% SDS and 10% protease inhibitor cocktail) and cell lysates were subjected to electrophoresis on 15% SDS–PAGE gels and probed with rabbit anti-hDR4/DR5 polyclonal antibody (Santa Cruz Biotechnology, Inc., Santa Cruz, CA).
Tumor size recording and animal survival rate
Mice were imaged along the dorsal view twice a week from day 0 to day 20 after bacteria injection using an IVIS. Bioluminescence was recorded as photons/second. Tumor volume (mm3) was measured by external calipers twice weekly and calculated using the formula L × W2 × 0.5, where L and W represent length and width of the tumor in millimeters, respectively. All mice were monitored for survival as previously described.
Pathological section and H&E staining
The tumors were removed from nude mice on day 3 after bacteria injection. The tissue was fixed with 10% buffered formalin and processed for paraffin sectioning and hematoxylin–eosin (H&E) staining using standard methods.
Toxicity of sTRAIL in vivo
The toxicity of sTRAIL treatment delivered by E. coli DH5α was examined following intravenous injection of 5 × 107 c.f.u. of E. coli DH5α (sTRAIL) to tumor-bearing mice. Seven days later, a second injection was performed as described above. At 4 and 30 days after infection, serum from all mice was collected and aminotransferase/aspartate aminotransferase was analyzed as indicator of liver injury using standard method. Livers, spleens, kidneys and hearts were harvested on day 30 after bacteria injection and fixed in 10% buffered formalin. The tissues were sectioned, stained with H&E and histopathological changes in organs were examined.
Statistical analysis
The in vivo tumor growth data were analyzed by analysis of variance using the Statistica software (StatSoft, Tulsa, OK). The survival rates were analyzed by log-rank test using Statistica. P⩽0.05 was considered significant.
Results
E. coli DH5α specifically resides in tumors
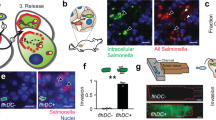
A total of 1 × 108 c.f.u. of luminescent E. coli DH5α were i.v. injected into tumor-bearing BALB/c or nude mice (n>10). Photon collection (3 min) using the IVIS at 18 h after bacterial injection showed that detectable light emission occurred only at the tumor site, indicating presence of luminescent E. coli (Figure 1a). Luminescent E. coli DH5α are also present in B16-F10 lung metastases in nude mice (Figure 1a). We performed a dose escalation test of E. coli DH5α and found that a dose of 1 × 108 or 5 × 107 c.f.u. in BALB/c mice or nude mice, respectively, was optimal for observing bacterial distribution without bacteremia. Unless otherwise specified, these doses were used in all subsequent animal experiments.
E. coli DH5α specifically resides in tumors. (a) Bacteria in tumors can be detected as early as 18 h after injection of 1 × 108 c.f.u. E. coli DH5α through the tail vein (left, B16-F10 murine melanoma in BALB/c mice; middle, H460 human lung carcinoma in nude mice; right, B16-F10 murine lung metastases in nude mice). (b) Tissue distribution of E. coli DH5α in immunocompetent BALB/c mice and immunocompromised nude mice using IVIS at different time points. Bars: photons/second (c) Various organs and tumors were removed from the mice and imaged using the IVIS without any processing at 30 min and day 4 after bacteria injection. (d) At different time points after infection, mice were killed. The livers, spleens and tumors were excised, homogenized and assayed for bacterial titers. ND, not detectable. Each value represents mean±s.d. from 10 mice. Photon collection was for 3 min at different time points after bacteria injection using the Xenogen IVIS 50 imaging system. C.f.u., colony-forming units; IVIS, In Vitro Imaging System.
To follow the fate of bacteria injected i.v. into the non-tumor-bearing animals, we monitored each animal with the IVIS at different time intervals as indicated in Figure 1b (n>10). Injection of E. coli DH5α immediately resulted in light emission localized mainly in the lungs, followed by accumulation of bioluminescent bacteria in the liver, which was present for several hours. Imaging of the same animals at 24 h post-injection revealed that detectable light emission from the earlier time points diminished (Figure 1b). These findings, together with complete absence of bacteria in the blood (data not shown), indicate that the light-emitting bacteria were probably eliminated by the host's immune system. Bacterial clearance was independently confirmed by absence of light emission in all of the excised organs of the same animal (data not shown).
To further determine the spatial and temporal progression of E. coli DH5α infection in animals with implanted tumors, luminescent E. coli DH5α were i.v. injected into BALB/c mice (n>10) with melanoma (∼200 mm3) that had been growing in the right hind leg. The animals were monitored each day for 15 days with a low-light imager. Bioluminescence distribution patterns determined immediately after injection were similar to distribution patterns in non-tumor-bearing animals. However, at day 1 after injection, no luminescence outside of the tumor region was detected. Continued monitoring of the mice showed that after an initial increase, luminescence decreased until day 4, and then dramatically increased in the tumors (Figure 1b), indicating efficient replication of the bacteria. Furthermore, i.v. injected luminescent E. coli DH5α also replicated in the human lung xenografts on nude mice (n>10; Figure 1b). At 30 min and 4 days post-infection, five of the tumor-bearing animals were killed. Luminescence in the excised liver and lung tissue was detected at the 30-min time point as indicated in Figure 1c. In contrast, detectable light emission only existed within the tumor site at day 4 post-infection, indicating that distribution of E. coli DH5α into tumors via the bloodstream did not result in significant infection of healthy organs (Figure 1c). The results of the bacterial count using homogenized samples at day 4 post-infection indicate that E. coli DH5α preferentially accumulates in tumors rather than in the spleens and livers at a ratio of >10 000:1. The numbers of bacteria detected in tumors was maintained as long as 18 days, while those in spleens and livers gradually disappeared (Figure 1d).
Cytotoxicity of sTRAIL-expressing E. coli DH5α in vitro
We then constructed a strain of sTRAIL-expressing E. coli DH5α, which could specifically target sTRAIL proteins into tumors. To determine whether sTRAIL-expressing E. coli DH5α had cytotoxic effects on human tumor cell lines in vitro, H460 lung tumor cells were incubated with E. coli DH5α (empty vector) and sTRAIL-expressing E. coli DH5α at various MOIs for 12 h. The cells were visualized by light microscopy for signs of apoptosis. Cells incubated with E. coli DH5α (empty vector) showed slight toxic effects, whereas cells incubated with sTRAIL-expressing E. coli DH5α underwent rapid and massive apoptosis (Figure 2a). A fusion protein having DR4 extracellular domain fused to the immunoglobulin Fc region (DR4: Fc) was added to the cell cultures which reduced, in a concentration-dependent manner, the amount of apoptotic cells and blocked morphological changes associated with apoptosis. As a negative control, 200 ng of TNF receptor:Fc per ml, which binds TNF but not TRAIL, was added to the cells and failed to block DH5α (sTRAIL)-induced apoptosis (Supplementary Figure S1).
Cytotoxicity of E. coli DH5α expressing sTRAIL in vitro. (a) H460 cells were infected with E. coli DH5α (sTRAIL) and E. coli DH5α (empty vector) at various MOIs and the morphological features of each cell were analyzed with a phase-contrast inverted microscope. (b) Cells were treated as described in panel a for 12 h, fixed and stained with crystal violet and photographed. (c) Cell survival was determined by trypan blue exclusion assay. The values represent the means of the values from triplicate wells. (d) The efficacy of sTRAIL-expressing E. coli DH5α on H460 tumor cells in vitro by FACS analysis with an MOI of 200 for 2, 4 and 8 h. The percentage of apoptotic cells was determined by quantifying sub-G1 cells after treatment. The representative results of three independent experiments are shown. Mock Control: Cells treated with cells culture medium; Vector Control: cells treated with E. coli DH5α (empty vector). FACS, fluorescence-activated cell sorting; MOI, multiplicity of infection; sTRAIL, soluble TRAIL; TRAIL, tumor necrosis factor (TNF)-related apoptosis-inducing ligand.
Further analyses demonstrated that the sTRAIL expressed by E. coli DH5α induces apoptosis in a dose-dependent fashion (Figures 2b and c). Apoptosis was characterized by FACS analysis in Figure 2d. The sub-G1 cell population increased in the E. coli DH5α (sTRAIL) -treated group in a time-dependent manner. These results indicate that E. coli DH5α-expressed sTRAIL effectively induces tumor cells apoptosis.
Antitumor activity of sTRAIL-expressing E. coli DH5α in vivo
These in vitro results encouraged us to evaluate whether tumor apoptosis by sTRAIL-expressing E. coli DH5α could be demonstrated in vivo. Nude mice (n=20) with luc+-labeled H460 lung tumors approximately 200 mm3 in size were intratumorally injected with E. coli DH5α (empty vector) or E. coli DH5α (sTRAIL). Tumor growth was monitored by two-dimensional caliper measurement and by in vivo tumor imaging after intraperitoneal injection of luciferin. As indicated in Figures 3a and b, H460 tumors in the mock control group grew exponentially, increasing 20-fold in size over the time period examined. Tumor growth was significantly inhibited at 1 week post-injection both in the vector control and in the E. coli DH5α (sTRAIL) groups, compared with that in the mock control group (P<0.0001), and the difference between the vector control and E. coli DH5α (sTRAIL) groups was not significant (P=0.29). In the vector control group, all tumors underwent regrowth during the subsequent 3 weeks, reaching a 17-fold increase as compared with the original tumor size. These results suggest that E. coli DH5α alone had little effect on the growth of H460 tumors in mice, with less than 10% inhibition on day 30 post-injection. In the E. coli DH5α (sTRAIL)-treated group, tumor growth was substantially suppressed as compared with that in control mice. The difference in tumor volume on day 30 between the sTRAIL-treated group and the mock or vector control groups is statistically significant (P<0.0001; Figure 3b).
The antitumor activity of sTRAIL-expressing E. coli DH5α when administered intratumorally. (a) Whole-body imaging of the efficacy of sTRAIL-expressing E. coli DH5α on the growth of H460 tumors after i.t. injection. Tumors were imaged at the indicated time points after bacteria injection. Bar: photons/second. (b) Tumor volume (mm3) was also measured by external calipers twice per week and calculated as described under Materials and methods. Each point represents the mean for n=20 animals. The results represent the mean±s.d. The experiment was repeated independently at least three times with similar results. The arrows indicate time points at which treatment was administered. (c) Necrotic areas and more condensed nuclei were seen in H460 tumors on day 3 after injection with sTRAIL-expressing E. coli DH5α. Scale bars, 50 μm. sTRAIL, soluble TRAIL; TRAIL, tumor necrosis factor (TNF)-related apoptosis-inducing ligand.
We next examined whether in vivo delivery of sTRAIL by E. coli DH5α into tumor tissues induces tumor cell death via apoptosis. To address this question, we injected the mock control, the E. coli DH5α vector control and sTRAIL-expressing E. coli DH5α into established tumors, excised the tumor tissues 3 days later and performed histological analysis on tissue sections. As shown in Figure 3c, H&E staining reveals that tumors injected with sTRAIL-expressing E. coli DH5α showed much more areas of necrosis and numerous cells with condensed nuclei as compared with the control groups. These data clearly indicate that sTRAIL-expressing E. coli DH5α suppresses tumor growth by inducing tumor cell apoptosis.
In general, local therapies such as i.t. injection have limited efficacy in clinical treatments. Therefore, the antitumor efficacy of sTRAIL-expressing E. coli DH5α was tested following systemic delivery. As a result, tail-vein injection of E. coli DH5α had little effect on tumor regression. In contrast, tumor growth was significantly retarded in mice treated with sTRAIL-expressing E. coli DH5α. The mean tumor volumes in mice treated with sTRAIL-expressing E. coli DH5α was dramatically lower than that of tumors treated with sterile normal saline on day 30 (P<0.0001). The difference between the sTRAIL-treated group and the vector control group is also statistically significant (P=0.008; Figure 4a). Moreover, survival of mice treated with sTRAIL-expressing E. coli DH5α was significantly prolonged as compared with that of mock- or vector control-treated mice. The survival of mock- and vector control-treated animals was about 40% at day 30 post-injection, whereas all the mice treated with sTRAIL-expressing E. coli DH5α were still alive at the same time point (Figure 4b).
Efficacy of sTRAIL-expressing E. coli DH5α on tumor growth and animal survival in vivo when administered systematically. (a) Quantitative efficacy of sTRAIL-expressing E. coli DH5α on the growth of H460 tumors in nude mice after intravenous injection. Tumor volume (mm3) was measured by external calipers twice per week and calculated as described under Materials and methods. Each point represents the mean for n=20 animals. The experiment was repeated independently at least three times with similar results. The arrows indicate time points at which treatment was given. (b) Effect of sTRAIL-expressing E. coli DH5α on the survival of animals bearing subcutaneous H460 xenografts. Animals (n=20) were treated as described under Materials and methods. sTRAIL, soluble TRAIL; TRAIL, tumor necrosis factor (TNF)-related apoptosis-inducing ligand.
Validation and toxicity
To further confirm that sTRAIL-expressing E. coli DH5α localizes preferentially in tumors, we performed PCR on bacterial colonies obtained from tumor tissues. H460 tumors were allowed to grow in nude mice until they were approximately 200 mm3 in size, and then 5 × 107 c.f.u. of sTRAIL-expressing E. coli DH5α were injected through the tail vein. Four days after injection the animals were killed and tumors, spleens, livers and kidneys were aseptically removed, homogenized, diluted and plated onto LB agar plates supplemented with 50 μg ml−1 penicillin to determine the c.f.u. of sTRAIL-expressing E. coli DH5α. As shown in Figure 5a, sTRAIL-expressing E. coli DH5α was found preferentially within the tumor tissue. PCR analysis using sTRAIL primers of randomly picked bacterial colonies showed that the bacteria cultured from the tumors contained the sTRAIL plasmids. Immunohistochemical analysis of tumor tissues also indicated positive sTRAIL staining (Figure 5b). In addition, sTRAIL protein was undetectable by a TRAIL-specific ELISA in mice serum on day 1, 4 and 7 after 5 × 107 c.f.u. of DH5α (sTRAIL) were injected i.v. (data not shown).
Validation and toxicity. (a) Comparison of the numbers of sTRAIL-expressing E. coli DH5α in both tumor and normal tissues at 4 days post-injection of viable bacteria into tumor-bearing mice through the tail vein. The experiments were repeated at least three times. Bacterial clones from the tumor plate were determined by PCR assay using the corresponding sTRAIL primers. ‘—’ indicates negative control with E. coli DH5α (empty vector) as PCR template. (b) Immunohistochemical staining for sTRAIL protein on different organs obtained from day 4 post-injection of sTRAIL-expressing E. coli DH5α. Scale bars, 50 μm. (c) Hepatotoxicity in tumor-bearing nude mice after systemic administration of E. coli DH5α-expressed sTRAIL. Serum samples were collected 4 and 30 days after tail-vein injection and aminotransferase/aspartate aminotransferase was measured using standard method. The values represent the mean±s.d. for 10 animals. (d) H&E-stained liver sections from the same animals as in panel c on day 30 after tail vein-injection. Scale bars, 50 μm. H&E, hematoxylin–eosin; sTRAIL, soluble TRAIL; TRAIL, tumor necrosis factor (TNF)-related apoptosis-inducing ligand.
To determine whether E. coli DH5α (sTRAIL) caused hepatocellular toxicity in vivo, we measured the serum levels of two indicators of hepatocellular damage, alanine aminotransferase and aspartate aminotransferase, at 4 and 30 days after infection. As shown in Figure 5c, the values for serum aminotransferase and aspartate aminotransferase were within normal ranges, and no significant difference was found among the groups. Furthermore, histological analysis of liver sections performed on day 30 showed that expression of sTRAIL did not cause any pathological changes as compared with those in mock- and vector control-treated mice (Figure 5d). Histological analysis of other tissues produced similar results to those obtained from liver (data not shown). Notably, mice injected with 5 × 107 c.f.u. of sTRAIL-expressing E. coli DH5α once per week for two consecutive weeks tolerated the infection and survived longer than mock- and vector control-treated mice (Figure 4b). We also examined the effect of DH5α (sTRAIL) on the TRAIL sensitivity of tumor cells and demonstrated that H460 cells aseptically separated from the tumor-bearing mice treated with DH5α (sTRAIL) are still susceptible to TRAIL-induced apoptosis (Supplementary Figure S2a). Consistently, expression of TRAIL receptor DR4/DR5 was not obviously affected by DH5α or DH5α (sTRAIL) administration (Supplementary Figure S2b). All these results suggest that E. coli DH5α-mediated sTRAIL expression kills tumor cells without detectable toxicity on normal cells.
Discussion
Specifically inducing tumor cells to undergo apoptosis is a promising therapeutic approach for cancer. In this study, we showed that non-pathogenic E. coli carrying a soluble TRAIL protein administered systemically, selectively localized in tumors and resulted in persistent tumor growth inhibition in mice. This treatment combines the advantages of tumor-targeting E. coli and the potent antitumor effects of sTRAIL.
The sTRAIL protein in various forms has been demonstrated to induce apoptosis of many cancer cells in vitro and inhibit growth of many tumors in rodent models. The sTRAIL protein acts by activating the cell-surface DR4 and DR5.35, 36 In our study, inhibition of tumor growth was mediated by sTRAIL protein expressed in E. coli, since E. coli carrying the empty vector did not have tumor inhibition activity and a DR4:Fc fusion molecule could block the sTRAIL-harboring E. coli from inducing H460 cell apoptosis in vitro.
Bacteria, including S. typhimurium, have recently gained attention as tumor-targeting vectors.37, 38 These bacteria selectively localize in tumor tissues when administered systemically, while the widely used adenoviral vector and adeno-associated virus vector do not have inherent tumor tropism and their tumor specificity is usually achieved via i.t. vector delivery.39 Otherwise, side effects associated with systemic administration of these viral vectors and the transgene products expressed would be a significant concern.40 The tumor targeting in mice by E. coli, as observed in this study, is impressively specific and efficient. E. coli DH5α administered i.v. localized specifically in several types of solid tumors in mice (murine B16 melanoma in BALB/c mice, human H460 lung carcinoma in nude mice) and also in multiple tiny metastatic sites of murine B16 melanoma in nude mice. This specific tumor targeting is possibly a mechanism related to the unique leaking tumor vasculature, the immune-suppression environment or the rich nutrient environment of tumor tissues.41 The tumor-targeting property of S. typhimurium has been confirmed in dogs and humans in addition to mice, although with varied efficiency.27, 42, 43, 44 Nevertheless, these findings offer cautious optimism for using bacteria as tumor gene therapy vectors.
Another prominent feature of bacterial gene delivery vectors is that they are fully replication-competent while adenovirus or adeno-associated virus gene delivery vectors used for in vivo therapy purpose are usually replication-incompetent. This feature provides a basis for the in vivo persistence of the bacterial vectors and the therapeutic gene they carry. In our experiments, E. coli could be detected from the tumor tissue up to 1 month after administration (data not shown). The recombinant sTRAIL protein has a short half-life in vivo: from 3 to 5 min in mouse and 30 min in cynomolgus monkeys or chimpanzees.12 But the E. coli-expressed sTRAIL protein could be detected in tumor tissue at 4 days after administration. sTRAIL expression at later time points had not been determined, but the flat tumor growth curve at 3 weeks after the last administration suggests that sTRAIL expression in tumor might have lasted much longer. Moreover, E. coli DH5α delivery of sTRAIL protein results in diffusion throughout tumor tissues through the bloodstream. This may lead to increased extensive tumor killing, transcending the limitations of current gene therapies by affecting infected cells and adjacent areas.45
Compared with other bacterial vectors with potential for tumor gene delivery, the E. coli vector employed here has certain advantages. First, E. coli is an extracellular bacterium while S. typhimurium is mainly intracellular. Since the sTRAIL molecule acts by interacting with the extracellular portion of its receptor, it would be advantageous and more efficient to have sTRAIL released by an extracellular bacterium. Secondly, as a facultative anaerobic bacterium, E. coli can grow in both small and large tumors, whereas anaerobic bacterium can only grow in large tumors with a necrotic center.
Bacterial vectors administered systematically would inevitably result in some complications, including certain level of bacteremia and induction of circulating proinflammatory cytokines due to their endotoxin. In clinical trials of the attenuated S. typhimurium strain VNP20009, tumor localization was only found in some patients receiving the bacteria at high doses where dose-limiting toxicity was found.27 E. coli will probably share some of these complications. However, many E. coli species are harbored by humans and other mammalian species in their digestive tract in a symbiotic relationship; thus non-pathogenic E. coli bacteria are likely to evoke a lower immune response and toxicity response in animals as compared with that by attenuated S. typhimurium. In this study, the endotoxin levels in mice plasma on day 0.5, 1, 3 and 7 after intravenous injection of 5 × 107 c.f.u. DH5α were found to differ non-significantly between the bacteria-injected mice and normal saline-injected mice (data not shown). Thus, in terms of safety profile and efficiency in tumor localization in humans, E. coli might hold more potential than other bacteria. Furthermore, E. coli are the most thoroughly studied and known bacteria; numerous well-characterized strains and tools are available for their manipulation, which will certainly facilitate related studies.
To our knowledge, this is the first report of tumor suppression in vivo achieved by using E. coli as the protein delivery vector. This E. coli DH5α (sTRAIL) therapy could serve as a prototype for further development of a feasible and effective form of treatment for solid tumors. In addition, our study has general implications for the application of non-pathogenic E. coli as human gene therapy vectors.
Abbreviations
- TRAIL:
-
tumor necrosis factor-related apoptosis-inducing ligand
- E. coli :
-
Escherichia coli
- IVIS:
-
In Vivo Imaging System
- c.f.u.:
-
colony-forming units
- i.v.:
-
intravenously
- i.t.:
-
intratumoral
- MOI:
-
multiplicity of infection
References
Smith CA, Farrah T, Goodwin RG . The TNF receptor superfamily of cellular and viral proteins: activation, costimulation, and death. Cell 1994; 76: 959–962.
Armitage RJ . Tumor necrosis factor receptor superfamily members and their ligands. Curr Opin Immunol 1994; 6: 407–413.
Wiley SR, Schooley K, Smolak PJ, Din WS, Huang CP, Nicholl JK et al. Identification and characterization of a new member of the TNF family that induces apoptosis. Immunity 1995; 3: 673–682.
Griffith TS, Chin WA, Jackson GC, Lynch DH, Kubin MZ . Intracellular regulation of TRAIL-induced apoptosis in human melanoma cells. J Immunol 1998; 161: 2833–2840.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M et al. Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 1999; 5: 157–163.
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA et al. Safety and antitumor activity of recombinant soluble Apo2 ligand. J Clin Invest 1999; 104: 155–162.
Jo M, Kim TH, Seol DW, Esplen JE, Dorko K, Billiar TR et al. Apoptosis induced in normal human hepatocytes by tumor necrosis factor-related apoptosis-inducing ligand. Nat Med 2000; 6: 564–567.
Lawrence D, Shahrokh Z, Marsters S, Achilles K, Shin D, Mounho B et al. Differential hepatocyte toxicity of recombinant Apo2L/TRAIL versions. Nat Med 2001; 7: 383–385.
Kim SH, Kim KH, Kwagh JG, Dicker DT, Herlyn M, Rustgi AK et al. Death induction by recombinant native TRAIL and its prevention by a caspase-9 inhibitor in primary human esophageal epithelial cells. J Biol Chem 2004; 279: 40044–40052.
Nesterov A, Ivashchenko Y, Kraft AS . Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) triggers apoptosis in normal prostate epithelial cells. Oncogene 2002; 21: 1135–1140.
Leverkus M, Neumann M, Mengling T, Rauch CT, Brocher EB, Krammer PH et al. Regulation of tumor necrosis factor-related apoptosis-inducing ligand sensitivity in primary and transformed human keratinocytes. Cancer Res 2000; 60: 553–559.
Kelley SK, Harris LA, Xie D, DeForge L, Totpal K, Bussiere J et al. Preclinical studies to predict the disposition of Apo2L/tumor necrosis factor-related apoptosis-inducing ligand in humans: characterization of in vivo efficacy, pharmacokinetics, and safety. J Pharmacol Exp Ther 2001; 299: 31–38.
Wang Y, Huang F, Cai H, Zhong S, Liu X, Tan WS . Potent antitumor effect of TRAIL mediated by a novel adeno-associated viral vector targeting to telomerase activity for human hepatocellular carcinoma. J Gene Med 2008; 10: 518–526.
Shashkova EV, Kuppuswamy MN, Wold WS, Doronin K . Anticancer activity of oncolytic adenovirus vector armed with IFN-alpha and ADP is enhanced by pharmacologically controlled expression of TRAIL. Cancer Gene Ther 2008; 15: 61–72.
Wohlfahrt ME, Beard BC, Lieber A, Kiem HP . A capsid-modified, conditionally replicating oncolytic adenovirus vector expressing TRAIL leads to enhanced cancer cell killing in human glioblastoma models. Cancer Res 2007; 67: 8783–8790.
Lin TY, Gu J, Zhang LD, Huang XF, Stephens LC, Curley SA et al. Targeted expression of green fluorescent protein/ tumor necrosis factor-related apoptosis-inducing ligand fusion protein from human telomerase reverse transcriptase promoter elicits antitumor activity without toxic effects on primary human hepatocytes. Cancer Res 2002; 62: 3620–3625.
Armeanu S, Lauer UM, Smirnow I, Schenk M, Weiss TS, Gregor M et al. Adenoviral gene transfer of tumor necrosis factor-related apoptosis-inducing ligand overcomes an impaired response of hepatoma cells but cause severe apoptosis in primary human hepatocytes. Cancer Res 2003; 63: 2369–2372.
Parker RC, Plummer HC, Siebenmann CO, Chapman MG . Effect of histolyticus infection and toxin on transplantable mouse tumors. Proc Soc Exp Biol Med 1947; 66: 461–467.
Mose JR, Mose G . Oncolysis by clostridia. I. Activity of Clostridium butyricum (M-55) and other nonpathogenic Clostridia against the Ehrlich carcinoma. Cancer Res 1964; 24: 212–216.
Gericke D, Engelbart K . Oncolysis by Clostridia. II. Experiments of a tumour spectrum with a variety of Clostridia in combination with heavy metal. Cancer Res 1964; 24: 217–221.
Thiele EH, Arison R, Boxer GE . Oncolysis by Clostridia. III. Effects of Clostridia and chemotherapeutic agents on rodent tumors. Cancer Res 1964; 24: 222–233.
Xu YF, Zhu LP, Hu B, Fu GF, Zhang HY, Wang JJ et al. A new expression plasmid in Bifidobacterium longum as a delivery system of endostatin for cancer gene therapy. Cancer Gene Ther 2007; 14: 151–157.
Sasaki T, Fujimori M, Hamaji Y, Hama Y, Ito K, Amano J et al. Genetically engineered Bifidobacterium longum for tumor-targeting enzyme–prodrug therapy of autochthonous mammary tumors in rats. Cancer Sci 2006; 97: 649–657.
Fox ME, Lemmon MJ, Mauchline ML, Davis TO, Giaccia AJ, Minton NP et al. Anaerobic bacteria as a delivery system for cancer gene therapy: in vitro activation of 5-fluorocytosine by genetically engineered Clostridia. Gene Ther 1996; 3: 173–178.
Agrawal N, Bettegowda C, Cheong I, Geschwind JF, Drake CG, Hipkiss EL et al. Bacteriolytic therapy can generate a potent immune response against experimental tumors. Proc Natl Acad Sci USA 2004; 101: 15172–15177.
Low KB, Ittensohn M, Le T, Platt J, Sodi S, Amoss M et al. Lipid A mutant Salmonella with suppressed virulence and TNFα induction retain tumor-targeting in vivo. Nat Biotechnol 1999; 17: 37–41.
Toso JF, Gill VJ, Hwu P, Marincola FM, Restifo NP, Schwartzentruber DJ et al. Phase I study of the intravenous administration of attenuated Salmonella typhimurium to patients with metastatic melanoma. J Clin Oncol 2002; 20: 142–152.
Zhao M, Yang M, Li XM, Jiang P, Baranov E, Li S et al. Tumor-targeting bacterial therapy with amino acid auxotrophs of GFP-expressing Salmonella typhimurium. Proc Natl Acad Sci USA 2005; 102: 755–760.
Zhao M, Yang M, Ma HY, Li XM, Tan XY, Li S et al. Targeted therapy with a Salmonella typhimurium leucine–arginine auxotroph cures orthotopic human breast tumors in nude mice. Cancer Res 2006; 66: 7647–7652.
Zhao M, Geller J, Ma HY, Yang M, Penman S, Hoffman RM . Monotherapy with a tumor-targeting mutant of Salmonella typhimurium cures orthotopic metastatic mouse models of human prostate cancer. Proc Natl Acad Sci USA 2007; 104: 10170–10174.
Yu YA, Shabahang S, Timiryasova TM, Zhang Q, Beltz R, Gentschev I et al. Visualization of tumors and metastases in live animals with bacteria and vaccinia virus encoding light-emitting proteins. Nat Biotechnol 2004; 22: 313–321.
Cheng CM, Lu YL, Chuang KH, Hung WC, Shiea J, Su YC et al. Tumor-targeting prodrug-activating bacteria for cancer therapy. Cancer Gene Ther 2008; 15: 393–401.
Contag CH, Contag PR, Mullins JI, Spilman SD, Stevenson DK, Benaron DA . Photonic detection of bacterial pathogens in living hosts. Mol Microbiol 1995; 18: 593–603.
Francis KP, Joh D, Bellinger-Kawahara C, Hawkinson MJ, Purchio TF, Contag PR . Monitoring bioluminescent Staphylococcus aureus infections in living mice using a novel luxABCDE construct. Infect Immun 2000; 68: 3594–3600.
Pan GH, O’Rourke K, Chinnaiyan AM, Gentz R, Ebner R, Ni J et al. The receptor for the cytotoxic ligand TRAIL. Science 1997; 276: 111–113.
Walczak H, Degli-Esposti MA, Johnson RS, Smolak PJ, Waugh JY, Boiani N et al. TRAIL-R2: a novel apoptosis-mediating receptor for TRAIL. EMBO J 1997; 16: 5386–5397.
Lee CH, Wu CL, Shiau AL . Salmonella choleraesuis as an anticancer agent in a syngeneic model of orthotopic hepatocellular carcinoma. Int J Cancer 2008; 122: 930–935.
Fu W, Chu L, Han X, Liu X, Ren D . Synergistic antitumoral effects of human telomerase reverse transcriptase-mediated dual-apoptosis-related gene vector delivered by orally attenuated Salmonella enterica Serovar Typhimurium in murine tumor models. J Gene Med 2008; 10: 690–701.
Yoo JS, Choi SY, Hwang KS, Cho WK, Jung CR, Kwon ST et al. Adeno-associated virus-mediated gene transfer of a secreted form of TRAIL inhibits tumor growth and occurrence in an experimental tumor model. J Gene Med 2006; 8: 163–174.
Wilson JM . Adenoviruses as gene-delivery vehicles. N Engl J Med 1996; 334: 1185–1187.
Ryan RM, Green J, Lewis CE . Use of bacteria in anticancer therapies. BioEssays 2005; 28: 84–94.
Herbst RS, Hess KR, Tran HT, Tseng JE, Mullani NA, Charnsangavej C et al. Phase I study of recombinant human endostatin in patients with advanced solid tumors. J Clin Oncol 2002; 20: 3792–3803.
Nemunaitis J, Cunningham C, Senzer N, Kuhn J, Cramm J, Litz C et al. Pilot trial of genetically modified, attenuated Salmonella expressing the E. coli cytosine deaminase gene in refractory cancer patients. Cancer Gene Ther 2003; 10: 737–744.
Thamm DH, Kurzman ID, King I, Li ZJ, Sznol M, Dubielzig RR et al. Systemic administration of an attenuated, tumor-targeting Salmonella typhimurium to dogs with spontaneous neoplasia: phase I evaluation. Clin Cancer Res 2005; 11: 4827–4834.
Green NK, Seymour LW . Adenoviral vectors: systemic delivery and tumor targeting. Cancer Gene Ther 2002; 9: 1036–1042.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (no. 30525021, 30672357, 30770471 and 30621063), the Major State Basic Research Development Program of China (973 Program) (no. 2004CB518800) and the National High Technology Research and Development Program of China (863 Program) (no. 2006AA02Z340).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Cancer Gene Therapy website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Zhang, HY., Man, JH., Liang, B. et al. Tumor-targeted delivery of biologically active TRAIL protein. Cancer Gene Ther 17, 334–343 (2010). https://doi.org/10.1038/cgt.2009.76
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cgt.2009.76
Keywords
This article is cited by
-
Promising dawn in tumor microenvironment therapy: engineering oral bacteria
International Journal of Oral Science (2024)
-
A strategy for enhanced tumor targeting of photodynamic therapy based on Escherichia coli-driven drug delivery system
Science China Materials (2021)
-
Bacteria-cancer interactions: bacteria-based cancer therapy
Experimental & Molecular Medicine (2019)
-
Triptolide modulates tumour-colonisation and anti-tumour effect of attenuated Salmonella encoding DNase I
Applied Microbiology and Biotechnology (2019)
-
Secretion of tumoricidal human tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) by recombinant Lactococcus lactis: optimization of in vitro synthesis conditions
Microbial Cell Factories (2018)