Abstract
Zinc-finger proteins (ZNFs) are one of the most abundant groups of proteins and have a wide range of molecular functions. Given the wide variety of zinc-finger domains, ZNFs are able to interact with DNA, RNA, PAR (poly-ADP-ribose) and other proteins. Thus, ZNFs are involved in the regulation of several cellular processes. In fact, ZNFs are implicated in transcriptional regulation, ubiquitin-mediated protein degradation, signal transduction, actin targeting, DNA repair, cell migration, and numerous other processes. The aim of this review is to provide a comprehensive summary of the current state of knowledge of this class of proteins. Firstly, we describe the actual classification of ZNFs, their structure and functions. Secondly, we focus on the biological role of ZNFs in the development of organisms under normal physiological and pathological conditions.
Similar content being viewed by others
Facts
-
Zinc-finger proteins (ZNFs) are involved in several cellular processes acting through different molecular mechanisms.
-
ZNFs have key role in development and differentiation of several tissues.
-
ZNFs are involved in tumorigenesis, cancer progression and metastasis formation.
-
Alterations in ZNFs are involved in the development of several of diseases such as neurodegeneration, skin disease and diabetes.
Open questions
-
ZNFs may act both as oncogene or tumor suppressor gene; can restoration or depletion of ZNFs expression be a new challenge in cancer drug design?
-
Could ZNFs be used as a prognostic factor for cancer, neurodegeneration, or other diseases?
ZNF structure, classification, and molecular functions
The first ZNF was identified in the late 1980s. The first ZNF was Transcription Factor IIIa (TFIIIa) from Xenopus laevis. This gave rise to the discovery of a new group of transcriptional activator proteins with a 30 amino acid repeating region. This new class of proteins was able to bind specific sequences of DNA.1,2 The zinc-finger structure (extensively reviewed in refs 3–7) is maintained by the zinc ion, which coordinates cysteine and histidine in different combinations. In classical C2H2 zinc-finger proteins, two cysteines in one chain and two histidines in other one are coordinated by a zinc ion. Crystallographic studies revealed that classical zinc-finger domains have two β-sheets and one α-helix.8
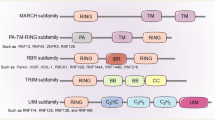
Non-classical types of zinc-finger differ in cysteine/histidine combinations, such as C2–H2, C2–CH, and C2–C2. Currently, 30 types of ZNFs are approved by The HUGO Gene Nomenclature Committee,9 and ZNF classification is based on the zinc-finger domain structure. A complete list of ZNF types with a description of the zinc-finger domain structure, the number of genes included, and the most studied members is summarized in Table 1. The most important and abundant types of zinc-finger domain proteins include C2H2, really interesting new gene (RING), plant homeodomain (PHD), and Lin-ll, Isl-1, and Mec-3 (LIM domains). Their protein structures are presented in Figure 1a.
Structure, molecular functions, and subcellular localization of ZNFs. (a) A schematic representation of the structure of C2H2, RING, PHD, and LIM zinc-finger domains. (b) A schematic representation of the structure of some ZNFs with multiple zinc-finger domains. (c) Gene ontology analysis of 1723 annotated ZNFs according to molecular function, log10(P-val)<(−5). (d) Schematic representation of the subcellular localization of different ZNFs.
Among C2H2 ZNFs, there are a large number of transcription factors with the C-x-C-x-H-x-H motif, which mediates direct interaction with DNA. One of the C2H2 members, ZNF217, contains multiple C2H2 domains. These domains bind a specific DNA sequence (T/A)(G/A)CAGAA(T/G/C), repressing the expression of target genes.10
The group of RING domain proteins include numerous E3-ubiquitin ligases. The RING-motif structure is C-x-C-x-C-x-H-x-C-x-C-x-C-x-C. One of the most important E3-ubiquitin ligases, Mouse Double Minute 2 (MDM2), which is involved in cancer progression, has a RING domain. This domain mediates its interaction with itself and Mouse Double Minute 4 (MDM4). This domain is also important for E3-ubiquitin ligase activity.11
The PHD zinc-finger domains are involved in the regulation of epigenetic modifications via their chromatin-remodelling ability. The PHD motif has the following primary structure: C-x-C-x-C-x-C-x-H-x-C-x-C-x-C. One of the ZNFs with a PHD domain is Lysine Demethylase 2A (KDM2A), which mediates nucleosome recognition.12
The LIM-type of ZNF was identified in transcription factors Lin-ll, Isl-1, and Mec-3.13 Currently, this class of ZNFs contains proteins important for actin targeting, cytoskeleton interaction, and focal adhesion. One important member, Paxillin, has four LIM motifs. The LIM motif structure is C-x-C-x-H-x-C-x-C-x-C-x-C-x-(C,H,D).14 The zinc-finger domain of Paxillin mediates β-catenin interaction at focal adhesion sites15 and stress fibers.16
Interestingly, many ZNFs contain multiple and different types of zinc-finger domains (Figure 1b). For example, two lysine demethylases, Lysine Demethylase 4A (KDM4A), a novel target for antitumor therapy,17 and KDM2A, which is required for DNA damage response,18 exhibit different zinc-finger compositions. Nevertheless, an important acetyltransferase, Lysine Acetyltransferase 6A (KAT6A), which regulates cell cycle progression,19 has the same zinc-finger pattern as KDM4A but exhibits a different molecular function. Furthermore, RANBP2-Type and C3HC4-Type Zinc-Finger Containing 1 (RBCK1), Ubiquitin Like With PHD And Ring Finger Domains 1 (UHRF1), and Roquin-1 contain a RING-type zinc-finger domain that possesses E3-ubiquitin ligase activity. These proteins also possess additional different zinc-finger domains. For example, RBCK1 contains a RAN-binding protein 2 (RanBP2) domain and has an important role in the immune response.20 UHFR1 also contains a PHD domain that is important for its repressive activity on gene promoters.21 Finally, Roquin-1 possesses a C3H1 domain that targets RNA.22
Gene ontology analysis of 1723 annotated human ZNFs revealed that this class of proteins has numerous functions (Figure 1c). ZNFs localize in different cell compartments (Figure 1d). Indeed, chromatin-remodelling ZNFs (for example, KDM2A, Lysine Methyltransferase 2B (KMT2B) and AT-Rich Interaction Domain 2 (ARID2)) and transcription factors (for example, ZNF750, Kruppel Like Factor 4 (KLF4) and GATA Binding Protein 2 (GATA2)) are localized in the nucleus. Cbl Proto-Oncogene (CBL) and TNF Receptor Associated Factor 4 (TRAF4) are membrane proteins. MDM2, Praja Ring Finger Ubiquitin Ligase (PJA2), and Autocrine Motility Factor Receptor (AMFR) belong to the E3-ubiquitin ligase family are mainly localized in the cytoplasm. However, MDM2 has been also show to localize in the nucleus.23,24 Rho/Rac Guanine Nucleotide Exchange Factor 2 (ARHGEF2) and Actin-Binding LIM Protein 1 (ABLIM1) are associated with the cytoskeleton, and Paxillin with ZNF185 is localized to focal adhesion sites.25
The zinc-finger domain is one of the most frequently utilized DNA-binding motif found in eukaryotic transcriptional factors. The binding of a zinc-finger domain to its target site juxtaposes three base pairs on DNA to a few amino acids in the α-helix structure. The identity of the aminonoacids at the contact site defines the DNA sequence recognition specificity of zinc fingers. Thus, by changing these amino acids, a high degree of selectivity can be achieved toward a given three base-pair DNA sequence.26 Exploiting this recognition mechanism, protein modules containing multiple zinc-finger motifs, each one recognizing a specific three base-pair DNA sequence, have been engineered which bind to specific DNA sequences.27 Fusing this recognition module with a sequence-independent endonuclease was the first successful strategy to introduce breaks at specific sites of genomic DNA.28 Precise genome editing was more recently achieved with other techniques based on transcription activator-like effector nucleases (TALEN)29 and clustered regularly interspaced short palindromic repeats (CRISPR) Cas930 whose description is beyond the scope of this review.
Physiological role of ZNFs
Skin
Through their ability to regulate gene expression, ZNF proteins participate in numerous physiological processes, including cell proliferation, differentiation, and apoptosis, thereby maintaining tissue homeostasis. For example, a recent study suggested that the zinc-finger proteins ZFP36 (RING-type, not transcription factor) an RNA binding protein, also known as Tristetrapolin31 and ZFP36L1 have a key role in the regulation of several aspects of keratinocyte biology, such as cell proliferation, differentiation and apoptosis. In fact, inhibition of the expression of these two proteins in cultured keratinocytes caused apoptosis and cell cycle arrest at the G2/M phase. In addition, Zfp36 knockdown in these cells also results in increased expression of the differentiation marker Keratin 10, suggesting the possible involvement of this protein in keratinocyte differentiation.32
Another zinc-finger protein that has a crucial role in keratinocyte differentiation, is the transcription factor, KLF4 (C2H2-type, transcription factor).33 Indeed, in the epidermis, KLF4 is mainly expressed in suprabasal layers, where it modulates the expression of genes involved in keratinocyte differentiation (ECM1, SPINK5, CDSN, FLG, and LCE3)34 (Figure 2a). In Klf4−/− mice, the absence of this ZNF protein results in altered skin barrier formation, causing embryonic death soon after birth.35 Conversely, the ectopic expression of KLF4 in basal keratinocytes of transgenic mice epidermis accelerates the differentiation process, resulting in early epidermal barrier formation.36
Molecular pathways regulated by ZNFs in physiological conditions. (a) ZNF750 regulates keratinocytes terminal differentiation by interacting with KLF4 and chromatin regulators. This interaction leads to the positive regulation of genes (PPL, PKP1) involved in differentiation. In addition, ZNF750 interacts with KDM1A and negatively regulates progenitor gene expression (RBBP8, HOMER3). ZNF750 directly regulates the expression of KLF4, which subsequently modulates the expression of the indicated genes. (b) KLF4 regulates epithelial cell differentiation by interacting with β-catenin and repressing the WNT signalling pathway. (c) KLF5 is involved in myoblast differentiation, acting as a co-factor for MyoD. This action leads to the upregulation of the indicated genes. (d) The presence of SLUG on the PPARG promoter reduces HDAC1 recruitment, leading to C/EBP-mediated activation of PPARG expression. This effect promotes adipogenesis.
The zing-finger protein ZNF750 (C2H2-type, transcription factor) also acts as an essential regulator of keratinocyte differentiation. During epidermal differentiation, TP63 transcription regulates ZNF750 expression, which subsequently directly activates KLF4 expression.37 However, more recently, a layer of complexity has been added to the underlying mechanism by which ZNF750 regulates terminal differentiation of keratinocytes. ZNF750 represses the expression of progenitor genes (RBBP8, HOMER3) by interacting with the chromatin modifiers REST corepressor 1 (RCOR1), lysine demethylase 1A (KDM1A) and C-terminal binding protein 1/2 (CTBP1/2) and activates differentiation genes (PPL, PKP1) by interacting with RCOR1, KLF4, and CTBP1/238 (Figure 2a). Interestingly, as described below, alteration of ZNF750 transcriptional regulation network during keratinocyte differentiation, caused by ZNF750 mutations, is involved in the development of diseases, such as psoriasis.39,40
Intestine
ZNF proteins are also involved in intestinal epithelium biology. For example, in addition to its well-documented role in skin homeostasis, KLF4 also has a key role in the intestines. In this tissue, KLF4 is expressed in the terminally differentiated epithelial cells (luminal surface) and goblet cells (crypts), where it promotes differentiation and inhibits proliferation.41–44 In particular, KLF4 represses intestinal epithelium proliferation by interacting with β-catenin and inhibiting β-catenin-mediated gene expression (Figure 2b). In addition, Klf4−/− mice lack goblet cells, indicating that KLF4 has an essential role in goblet cell differentiation.45
Another Krüppel-like factor, KLF5 (C2H2-type, transcription factor), is crucial for regulation of proliferation in the intestinal epithelium, exerting an opposing function to KLF4.42 KLF5, which is expressed in basal epithelial cells of the crypts, is activated by the Wnt signalling pathway, promoting cell proliferation.41
The GATA family members GATA4 (GATA-type, transcription factor) and GATA6 (GATA-type, transcription factor) have important roles in differentiation and homeostasis of the small intestinal epithelium. GATA4 expression is detected in the proximal but not the distal small intestine, having an important role in the maintenance of jejunal-ileal specifications. Indeed, in the jejuna of inducible intestine-selective GATA4 knockout mice, the inactivation of GATA4 results in downregulation of genes specifically expressed in the jejunum and increased expression of specific ileum genes.46 GATA6 is expressed in the entire small intestine, where is required for intestinal proliferation, secretory cell differentiation and absorptive enterocyte gene expression.47
Muscle
ZNFs have a regulatory function in muscle differentiation. For example, SET and MYND Domain Containing 1 (SMYD1), which is specifically expressed in striated muscle, acts as an essential regulator of myogenesis.48 SMYD1 (MYND-type, not transcription factor) deletion impaired myoblast differentiation, decreasing myofibre formation and reducing muscle-specific gene expression. Moreover, inhibition of KLF5 expression in cultured C2C12 myoblasts suppresses myotube formation, suggesting that this zinc-finger protein is required for myogenic differentiation. In particular, at a molecular level, KLF5 promotes myoblast differentiation into myotubes by recruiting MyoD to muscle-specific target genes (MYOG, MYBPH, MYL4, and MYOM2)49 (Figure 2c). Further examples of zinc-finger proteins involved in the regulation of skeletal myogenesis include CXXC Finger Protein 5 (CXXC5) and Early Growth Response 3 (EGR3). In fact, CXXC5 facilitates myocyte differentiation by positively regulating skeletal muscle differentiation genes,50 whereas EGR3 promotes myoblast proliferation by stimulating nuclear factor kappa B (NF-кB) signaling.51 By contrast, myogenic cellular differentiation is negatively regulated by murine zinc-finger Zfp637 (C2H2-type, not transcription factor). Although its transcriptional activity has not been fully investigated, Zfp637 overexpression inhibits differentiation and promotes proliferation of myoblasts, potentially regulating murine telomerase reverse transcriptase (mTERT) expression.52
Adipose tissue
Recent studies revealed an increased number of ZNFs as key transcriptional regulators involved in adipogenesis.53 For example, ZNF638 (Matrin-type, not transcription factor) seems to positively regulate this process given that its expression increases during preadipocyte differentiation. Indeed, ectopic expression of ZNF638 results in increased adipogenesis in vitro. On the other hand, inhibition of ZNF638 expression decreases differentiation by inhibiting the expression of adipocyte-specific genes. Specifically, ZNF638 promotes adipogenesis by acting as a transcriptional co-factor of CCAAT/enhancer-binding protein (C/EBP) and results in the expression of peroxisome proliferator-activated receptor γ (PPARG), which regulates adipocyte differentiation.54,55
The transcription factor SLUG (C2H2-type, transcription factor) is also involved in adipocyte differentiation in vitro and in vivo.56 Indeed, SLUG knockout mice exhibit decreased white adipose tissue (WAT) mass compared with wild-type mice, whereas the WAT size is increased in Slug-overexpressing mice. Accordingly, SLUG-deficient mouse embryonic fibroblasts (MEFs) exhibit impaired adipogenesis compared with wild-type MEFs. SLUG potentially controls WAT development by affecting Histone deacetylase 1 (HDAC1) recruitment to the PPARG promoter, favouring a more accessible chromatin state for PPARG transcriptional activators (Figure 2d).
In contrast to ZNF638 and SLUG, the GATA transcription factors GATA2 (GATA-type, transcription factor) and GATA3 (GATA-type, transcription factor) act as negative regulators of adipocyte differentiation.57 Indeed, their expression is detected in preadipocytes and is decreased during differentiation. Consistently, the ectopic expression of GATA2 and GATA3 in preadipocytes inhibits their transition to adipocytes by binding to the PPARG promoter, inhibiting PPARG expression. In addition, GATA2 and GATA3 also interact with C/EBPα, and C/EBPβ, suppressing their transcriptional activities. Both molecular mechanisms are required to negatively regulate adipogenesis.
Cellular stemness regulation
In mice, Zfp281 (C2H2-type, transcription factor), the murine homolog of ZNF281, has an important role in regulating cellular stemness by binding the promoter of Nanog and inhibiting its transcription.58 Further work demonstrated that Nanog transcriptional repression requires the coordinated activity of NANOG itself and Zfp281, which recruits the NuRD repressor complex.59 Functionally, modulation of Nanog expression by Zfp281 is an efficient mechanism to fine-tune reprogramming activity in stem cells.
Role of ZNFs in diseases
Tumour suppressor and oncogenic functions of ZNFs
Recent findings have highlighted the importance of ZNFs in cancer onset and progression. The zinc-finger family includes both tumour suppressor genes and oncogenes.60,61 ZNFs are involved in all the principal pathways of cancer progression from carcinogenesis to metastasis formation. Furthermore, ZNFs are involved in cancer via their transcription factor function. In addition, emerging evidence indicates the importance of zinc-finger proteins as recruiters of chromatin modifiers or as structural proteins that regulate cancer cell migration and invasion.
ZNF281
In recent years, several experimental studies revealed a role of ZNF281 (C2H2-type) in tumorigenesis and tumour invasion. ZNF281 is involved in two crucial processes in cancer: the DNA damage response (DDR)62–65 and the epithelial–mesenchymal transition (EMT). ZNF281 expression is increased upon DNA damage induced by drugs in several cancer types. In particular, the expression of several proteins involved in the DDR, including XRCC2, XRCC4, and Nucleolin, is regulated by ZNF281.66 Interestingly, two molecular mechanisms have been proposed: i) ZNF281 acts as transcription factor and directly regulates the transcription of XRCC2 and XRCC4; ii) ZNF281 also indirectly regulates Nucleolin expression, acting as a co-factor of c-Myc and producing an additive effect46 (Figure 3c).
Molecular mechanisms underlying the role of ZNFs in cancer biology (a) ZNF185 interacts with actin filaments in focal adhesion sites to regulate migration and invasion. (b) TGF-β induces the expression of ZEB1, which represses CDH1 expression, hence inducing EMT. (c) ZNF281 regulates expression of genes involved in the DDR and EMT. (d) ZNF750 acts as tumour suppressor gene by inducing the expression of the lncRNA TINCR, which inhibits cancer cell proliferation. In addition, ZNF750 represses LAMC2 expression, inhibiting cancer cell migration. (e) ZBP89 represses VIM and ODC1 expression by recruiting HDAC1 to the promoters of these genes. Moreover, ZBP89 induces MMP3 expression. (f) MDM2 interacts with p53 to induce proteasomal degradation and impair p53 to exert its function.
Moreover, ZNF281 has also a role in metastasis in colorectal cancer (CRC) through regulation of the EMT67 (Figure 4a). During the EMT, ZNF281 expression is induced by SNAIL and inhibited at the post-transcriptional level by miR-34a.68–70 The expression of miR-34a is subsequently promoted by p53, indicating that ZNF281 in CRC is controlled by a feed-forward loop (Figure 4c). In addition, modulation of ZNF281 expression in CRC regulates the EMT through the activation of SNAIL expression. However, ZNF281 can also directly bind the promoters of EMT effector genes, such as CDH-1, OCLN, and CLDN-7 (Figure 3c). Interestingly, ZNF281 expression was upregulated in patient tumour samples, confirming an important role of ZNF281 in CRC. These data strongly suggest that ZNF281 acts as an oncogene by regulating metastasis.
Transcriptional regulation of some ZNFs transcription and their roles in cancer. (a) SNAIL promotes the EMT by positively regulating the expression of ZNF281 and negatively regulating the expression of the tumour suppressor miR-34a. (b) p63 induces ZNF750 expression, which subsequently represses cell proliferation and migration. (c) ZNF185 expression is regulated by Brg-1 and the SWI/SNF complex. Its activation represses cell invasion and tumour growth. (d) TCF/ β-catenin induces ZBP89 expression, promoting tumour growth and metastasis.
ZNF750
ZNF750 is another member of the family that is involved in cancer. Indeed, ZNF750 has been described as tumour suppressor gene in squamous cell carcinomas (SCCs) of the oesophagus, lung and cervix.71,72 ZNF750 is mutated in SCCs, and truncation and missense mutations represent the most common mutations. These mutations are located in the C2H2 zinc-finger domain, suggesting the importance of the zinc-finger domain in mediating the tumour suppressor activity of ZNF750. In addition, ZNF750 is expressed at much lower levels in SCC patients compared with normal tissue. Hence, ZNF750 overexpression in vitro inhibits cell proliferation and migration (Figure 4b). Interestingly, overexpression of the C2H2 ZNF750 mutant is not able to suppress tumour growth, demonstrating that the C2H2 zinc-finger domain is essential for the tumour suppressor activity of ZNF750. At the molecular level, ZNF750 regulates a set of genes involved in cell migration, proliferation and adhesion. Particularly, ZNF750 directly induces the expression of the long non-coding RNA TINCR, through which it regulates cancer cell proliferation and tumour growth and represses the expression of LAMC2, a component of Laminin-332. Collectively, these actions regulate cancer cell migration73,74 (Figure 3d). Accordingly, low expression of ZNF750 has been observed in head and neck SCC and lung SCC patient datasets, and this expression pattern is associated with poor prognosis. Moreover, high levels of ZNF750 are associated with a good response to chemoradiotherapy, suggesting that ZNF750 could serve as a novel candidate biomarker for chemoradiotherapy sensitivity.75
ZNF185
ZNF185 (LIM-type, not transcription factor) is a zinc-finger protein that contains a LIM domain necessary for protein–protein interactions and an ATD (Actin Targeting Domain) domain with actin-binding activity.76 Proteins that contain LIM domains can be localized both in the nucleus and cytoplasm, exerting their molecular function through protein–protein interactions rather than DNA binding. The importance of ZNF185 in cancer progression is highlighted by its reduced expression in intermediate, high-grade, and metastatic prostate tumours compared with normal tissue. Interestingly, ZNF185 expression is reduced in prostate cancer owing to DNA methylation. In fact, prostate cancer cell lines treated with a DNA Methyl Transferase 1 (DNMT1) inhibitor exhibit increased ZNF185 expression.77 Indeed, deregulation of ZNF185 expression seems to be a recurring event in different human cancers, including prostate cancer, primary lung tumours, colon cancer and HNSCC.78,79 These data suggest a putative tumour suppressor function for ZNF185 by regulating cell proliferation and differentiation.80 Moreover, in lung cancer, BRG1, a component of the human switch/sucrose non-fermenting complex (SWI/SNF), regulates ZNF185 expression (Figure 4c). A possible mechanism by which ZNF185 exerts its function in cancer biology is through the interaction with actin filaments. Indeed, ZNF185 is associated with multiple actin-regulated structures, such as focal adhesion sites, and possesses growth inhibitory activity (Figure 3a). Localization of ZNF185 to the actin–cytoskeleton is mediated via its ATD domain, which is also required for its growth-suppressing activity. Furthermore, in prostate cancer, in addition to actin stress fibres, ZNF185 co-localizes with several cytoskeletal-related components, such as focal adhesion sites and filopodia/lamellipodia.81 These data suggest that ZNF185 may act as a novel tumour suppressor gene, having a key role in cancer onset and progression.
ZBP89
ZBP89 (C2H2-type, transcription factor), also known as ZNF148, is a well-characterized zinc-finger factor involved in cancer growth and apoptosis. Indeed, several tumours, such as breast cancer, melanoma and gastric cancer, exhibit increased ZBP89 expression compared with normal tissues, suggesting an oncogene function for ZBP89.82–84 However, ZBP89 may act as tumour suppressor gene in colorectal cancer by repressing cell proliferation and inducing apoptosis.85,86 ZBP89 exerts its molecular function via two different mechanisms. First, it may act as autonomous transcription factor by regulating the expression of MMP387 (Matrix Metallopeptidase 3) (Figure 3e) a protein involved in tumour development and metastasis.88,89 Second, ZBP89 inhibits ODC (Ornithine Decarboxylase)90 and Vimentin91 expression through recruitment of HDAC1 to the promoter of these genes92,93 (Figure 3e). ODC is involved in tumour development, and Vimentin has a role in cell migration and invasion. These findings suggest a role for ZBP89 in the inhibition of both neoplastic transformation and metastasis formation. Moreover, ZBP89 facilitates the recruitment of HDAC3 to the promoter of CDKN2A to restrain cellular senescence, facilitating lung cancer cell proliferation.94 Recently, it has been demonstrated that ZBP89 regulates the β-catenin pathway, supporting the hypothesis that ZBP89 is involved in cancer metastasis. Indeed, in colorectal cancer, the binding of ZBP89 to the promoter of CTNNB (β-catenin) results in increased gene expression. Interestingly, the inhibition of β-catenin expression resulted in a strong reduction in ZBP89 protein expression (Figure 4d). These data suggests that β-catenin accumulation initiates a cell proliferation program through the activation of its target genes, including Zbp89. Furthermore, the induction of ZBP89 contributes to sustaining β-catenin levels, further promoting cancer cell proliferation.95
MDM2
MDM2 (RANBP2-type; RING-motif, not transcription factor) is a zinc-finger protein that does not act as a transcription factor. Nevertheless, MDM2 has a very important role in tumour biology (extensively reviewed in Oliner et al.96). Its importance in cancer is attributed to its regulatory function on the tumour suppressor activity of p53. Indeed, MDM2 regulates p53 activity via three different mechanisms. First, given that MDM2 exhibits E3-ubiquitin ligase activity, it can ubiquitinate p53 to promote its proteasomal degradation. Second, MDM2 interacts with p53 to prevent the binding of p53 to its target genes, which mediate the tumour suppressor function of p53.97–99 Third, MDM2 binds to the N-terminus of p53, promoting the translocation of p53 into the cytoplasm and therefore blocking the activation of p53 target genes (Figure 3f).100–102 The importance of MDM2 in tumorigenesis is also provided by overexpression experiments. In fact, MDM2 overexpression induces spontaneous tumour formation.103,104 In addition, analysis of 28 tumour types performed on approximately 4000 patients revealed that the MDM2 gene is amplified in 7% of human cancers.105 Particularly, the percentage of MDM2 amplification is increased in liposarcomas (>80%), osteosarcomas (16%), soft tissue tumours (20%), and oesophageal carcinomas (13%).106 Moreover, point mutations affecting the zinc-finger of MDM2 have been described in human tumors.107 In vitro experiments show that these mutations disrupt the interaction of MDM2 with the ribosomal protein L5 and L11 and the ability to degrade p53.108 Given its importance, MDM2 is considered a putative target for therapies. An effort has been made to develop compounds that may prevent the interaction between MDM2 and p53, blocking the oncogenic activity of MDM2.109 As extensively reviewed by Wang et al.,110 three compounds (RG7112, RG7388 and SAR405838) exhibited relevant anti-tumoural activity in patients with p53 wild type in phase I clinical trials. Given that the anti-cancer activity of these compounds is attributed to the activation of wild type p53, and these compounds are expected be effective only in patients with wild type p53.111
ZEB1
ZEB1 (C2H2-type, transcription factor) is one of the most important zinc-finger proteins involved in tumour invasion and metastasis. Indeed, ZEB1 is one of the master regulators of the EMT112 (extensively reviewed in Zhang et al.113). ZEB1 expression is regulated by several signalling pathways, such as Wnt, TGF-β, NF-κB, and HIF signalling, and miRNA.114 The oncogenic role of ZEB1 is due to the repression of E-Cadherin expression, which is one of the most important cell–cell adhesion proteins. ZEB1 exerts its molecular function on E-Cadherin by interacting with several chromatin-remodelling factors, such as CtBP115 and the SWI/SNF complex.116 On the other hand, ZEB1 also directly activates the promoter of genes involved in the EMT. ZEB1 interacts with SMAD protein or with p300-P/CAF and activates TGF-β responsive genes to promote the EMT.117,118 Among ZEB1-activated genes, CDH2 (N-cadherin), a mesenchymal cadherin, is important in cancer progression (Figure 3b) given that altered expression of ZEB1 is observed in several human cancers, including pancreatic cancer, lung cancer, liver cancer, osteosarcoma, breast cancer, and colon cancer.109,119–122 Furthermore, the overexpression of ZEB1 in several cancer lines induces the EMT and promotes cell invasion.123,124
ZNF family members in neurodegenerative diseases
ZPR1
In recent years, ZNFs have been demonstrated to have an important role in the pathogenesis of neuronal diseases. Spinal muscular atrophy (SMA) is a rare neuromuscular disorder characterized by loss of α-motor neurons in the anterior horn of the spinal cord and progressive muscle wasting, often leading to early death.125 The cause of the disease is a mutation in the Survival Motor Neurons 1 (SMN1) gene that results in reduced expression of the full-length SMN protein, which is necessary for survival of motor neurons.126 The first evidence of a possible involvement of ZPR1 (C4-type, not transcription factor) in SMA came from the experimental observation that the SMN protein interacts with ZPR1. The consequence of this interaction is a redistribution of the complex from the cytoplasm to the nucleus. Interestingly, this process is hampered in patients affected by SMA type I. In addition, this observation is also corroborated by evidence demonstrating that ZPR1 is expressed at low levels in patients with severe SMA.127 Furthermore, it has been reported that, mutation of ZPR1 resulted in embryonic lethality in mice. Moreover, the reduction of ZPR1 expression in mice, results in increased loss of spinal motor neuron, a similar phenotype observed in mice with reduced Smn gene, suggesting that the lower ZPR1 expression observed in SMA patients, can contribute to the gravity of SMA.128
ZNF179
Zinc-Finger Protein 179 (ZNF179) (C4-type, not transcription factor) belongs to the RING finger class, and its expression is restricted in the brain, suggesting a possible role in the central nervous system.129 Indeed, inhibition of ZNF179 expression reduced neuronal differentiation in P19 cells and primary culture of cerebellar granule cells by inhibiting cell cycle progression through the regulation of p35 expression and the accumulation of p27 protein. More recently, it has been shown that ZNF179 has an anti-apoptotic role in astrocytes derived from the mouse APPtg model of Alzheimer’s disease. This effect is in part due to the inhibition of IGFBP3 and BIK expression.130,131
ZNF746
Recently, a novel role for ZNF746 (C2H2-type, not transcription factor), also known as Parkin Interacting Substrate (PARIS), has been identified in the pathogenesis of Parkinson’s disease (PD).132 Human ZNF746 is protein that contains C2HC and C2H2-type zinc-finger domains at the C-terminus. This protein is regulated by the proteasome system, in particular by ubiquitination mediated by Parkin, an E3-ubiquitin ligase. PD-associated mutations in the PARK2 gene lead to the loss of its E3 ligase function, resulting in ZNF746 accumulation in human PD brain.133 ZNF746 overexpression results in the loss of dopaminergic neurons in the substantia nigra by repressing the expression of peroxisome proliferator-activated receptor gamma (PPARγ) coactivator-1α (PPARGC1A)132 (Figure 5a).
Regulation of ZNFs target genes in human diseases. (a) ZNF746 represses the expression of PGC-1α, resulting in the loss of dopaminergic neurons in the substantia nigra of Parkinson’s patients. (b) ZNF750 regulates the expression of epidermal differentiation markers, such as FLG, LOR, SPINK5, ALOX12B, and DSG, which are altered in human skin diseases. (c) Glis1 regulates transcription of several genes involved in the differentiation of epidermal keratinocytes, including cornifin, involucrin, and transglutaminase 1. The expression of these genes is altered in psoriasis. (d) Glis3 modulates expression of the insulin gene, contributing to the pathogenesis of neonatal diabetes and hypothyroidism. (e) Troponin C and I and myosin light chain-3 genes are induced during cardiac hypertrophy due to overexpression of the GATA4 transcription factor. (f) The expression of SEMA3C and its receptor PLXNA2 is downregulated by GATA6 mutations, resulting in the development of OFT defects associated with CHDs.
ZNFs in other human diseases
ZNF750
Increasing evidence confirms the important roles of ZNFs in psoriasis. Psoriasis is a chronic inflammatory disorder of the skin, which varies in severity and clinical manifestation. ZNF750 is associated with a seborrhea-like dermatitis with psoriasiform elements.40 In particular, the 56_57dupCC mutation in ZNF750 has been identified in psoriasis patients and results in a frameshift mutation. This mutation leads to the production of a truncated protein that does not contain the zinc-finger domain. Downregulation of ZNF750 leads to reduced expression of genes involved in epidermal differentiation and skin barrier formation, such as Filaggrin (FLG), loricrin (LOR), serine protease inhibitor Kazal-type 5 (SPINK5), Arachidonate 12-Lipoxygenase, 12R Type (ALOX12B) and desmoglein1 (DSG1) (Figure 5b). These genes are mutated in various human skin diseases. In fact, the clinical manifestations of skin diseases derived from ZNF750 human mutations result from a combination of mutations in some of those downstream genes. ZNF750 and its downstream genes could be important targets for the treatment of skin diseases.39,134
GLIS1
Gli-similar protein 1 (GLIS1) (C2H2-type, transcription factor) is Krüppel-like zinc-finger protein involved in the pathogenesis of psoriasis. Indeed, GLIS1 is significantly overexpressed in psoriatic epidermis.135 GLIS1 mRNA is present only in the suprabasal layers of psoriatic skin, whereas normal human epidermis does not express GLIS1. These data suggest that GLIS1 that could be involved in the regulation of abnormal differentiation observed in psoriatic epidermis. Consistently, microarray analysis reveals that ectopic expression of GLIS1 transcriptionally regulates the expression of several genes involved in the differentiation of epidermal keratinocytes, including S100A9, KLK7, small proline-rich proteins (SPRRs), involucrin (IVL), and transglutaminase 1 (TGM1) (Figure 5c). GLIS1 contains both a repressor domain at its amino terminus and an activation domain at its carboxy terminus, resulting in both transcriptional repressor and transactivator functions. GLIS1 regulates transcription of target genes through binding to oligonucleotides containing the Gli-binding site consensus sequence, GACCACCCAC, as demonstrated by electrophoretic mobility shift assays. In addition, GLIS1 is expressed in different temporal and spatial patterns during the embryonic development, thus regulating gene expression at different stages of the developmental process as demonstrated by whole mount in situ hybridization studies performed on mouse embryos.136
GLIS3
Several reports indicate that the ZNF family might have a role in the development of diabetes. For example, GLIS3 (C2H2-type, transcription factor), a member of Kruppel-like Zinc-Finger proteins, is highly expressed in human pancreatic β-cells, and mutations in the GLIS3 gene have been identified in neonatal diabetes and congenital hypothyroidism (NDH).137 A human GLIS3 mutation that results in a truncated protein at its C-terminal domain has been identified, but the specific mechanism by which this mutation leads to the development of NDH has not been investigated to date. GLIS3 modulates the expression of the insulin through both direct and indirect mechanisms: binding to the INS promoter (Figure 5d) or modulating the activity of other β-cell-enriched transcription factors, such as MafA, Nkx6-1, and Pax6. Recently, a GLIS3-deficient (Glis3−/−) mouse model has been generated, exhibiting high blood sugar levels, pancreatic defects and premature death. These phenotypes resemble human neonatal diabetes caused by GLIS3 mutations.138 This murine model could be very useful for studying novel therapeutic applications in human diabetes. Novel clinical manifestations for patients with neonatal diabetes caused by GLIS3 mutations have been identified, such as osteopenia associated with skeletal deformity and fractures, bilateral sensorineural deafness and exocrine pancreatic dysfunction. These clinical features were not previously described, demonstrating great variability in GLIS3 mutated phenotype given that different genetic mutations result in tissue-specific expression of GLIS3 mRNA.139
GATA4
ZNFs are involved also in the pathogenesis of congenital heart diseases (CHDs). CHDs are the most common developmental anomaly affecting new-borns. For example, GATA4 is essential for proper cardiac morphogenesis. Indeed, GATA4 mutations are implicated in human congenital heart disease. A heterozygous G296S missense mutation of GATA4 has been identified140 that causes reduced transcriptional activity and DNA-binding affinity of GATA4. Furthermore, the GATA4 mutation prevents the physical interaction between GATA4 and TBX5, a T-box protein responsible for a subset of syndromic cardiac septal defects.141,142 Overexpression of GATA4 is associated with cardiac hypertrophy, where directly it regulates the expression of several cardiac specific proteins, such as troponin C and I and myosin light chain-3 (Figure 5e). Interestingly, these genes are induced during cardiac hypertrophy.143 In addition, expression of several other proteins, including Na+/Ca2+-exchanger, acetylcholine receptor-M2, cardiac-restricted ankyrin repeat protein (CARP), and adenosine receptor-A1 and carnitine palmitoyltransferase-1β, is regulated by GATA4.144,145 These findings suggest that GATA transcription factors could be an attractive therapeutic target for the treatment of cardiovascular diseases.
GATA6
Another GATA zinc-finger transcription factor expressed in the developing heart is GATA6. Mutations in this gene have been identified in patients with CHDs.146 Recent studies demonstrated that downregulation of GATA6 in neural crest-derived smooth muscle causes defects of the cardiac outflow tract (OFT) and in aortic arch arteries.147,148 GATA6 regulates neurovascular guiding molecule semaphorin 3C (SEMA3C) and its receptor plexin A2 (PLXNA2) expression (Figure 5f), which is important for a normal OFT. GATA6 mutations result in downregulation of these genes, disrupting semaphorin–plexin signalling and contributing to OFT defects, which accounts for 30% of CHDs.
Similar to GATA4, GATA6, and Tbx5 are co-expressed in the embryonic heart, and their interaction is necessary to activate the atrial natriuretic factor promoter during cardiac morphogenesis. The interaction between the GATA family of transcription factors and Tbx5 is necessary for proper cardiac function. Indeed, mutations in GATA4, GATA6, and TBX5 genes disrupt these interactions, contributing to the pathogenesis of CHDs.149
These data contribute to the identification of GATA mutations as a major genetic cause of CHDs.
Conclusions
It is now well accepted that ZNFs have a crucial role both in tissue homeostasis and disease. Interestingly, although this class of proteins was initially classified as transcription factors, several studies have highlighted novel functions of ZNFs. In fact, it has been shown that ZNFs could also act as recruiters of chromatin modifiers, as co-factors, or as structural proteins involved in cell migration and invasion.
In particular, the role of ZNFs in cancer development, progression and metastasis is becoming an interesting research issue. In fact, ZNF expression is upregulated or downregulated in cancer patients, demonstrating that ZNFs may act both as tumour suppressors or oncogenes. Furthermore, the functions of several ZNFs seem to be selective for specific tumours. Thus, the design of drugs that target specific ZNFs to avoid or restore abnormal expression of these proteins could be one of the most important challenges in the near future. Moreover, given the high specificity in terms of function and expression of some ZNFs for some tumours, it could be useful to exploit this class of proteins as prognostic factors.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Gibson TJ, Postma JP, Brown RS, Argos P . A model for the tertiary structure of the 28 residue DNA-binding motif ('zinc finger') common to many eukaryotic transcriptional regulatory proteins. Protein Eng 1988; 2: 209–218.
Vrana KE, Churchill ME, Tullius TD, Brown DD . Mapping functional regions of transcription factor TFIIIA. Mol Cell Biol 1988; 8: 1684–1696.
Matthews JM, Bhati M, Lehtomaki E, Mansfield RE, Cubeddu L, Mackay JP . It takes two to tango: the structure and function of LIM, RING, PHD and MYND domains. Curr Pharm Des 2009; 15: 3681–3696.
Hossain MA, Barrow JJ, Shen Y, Haq MI, Bungert J . Artificial zinc finger DNA binding domains: versatile tools for genome engineering and modulation of gene expression. J Cell Biochem 2015; 116: 2435–2444.
Klug A . The discovery of zinc fingers and their development for practical applications in gene regulation and genome manipulation. Q Rev Biophys 2010; 43: 1–21.
Malgieri G, Palmieri M, Russo L, Fattorusso R, Pedone PV, Isernia C . The prokaryotic zinc-finger: structure, function and comparison with the eukaryotic counterpart. FEBS J 2015; 282: 4480–4496.
Eom KS, Cheong JS, Lee SJ . Structural analyses of zinc finger domains for specific interactions with DNA. J Microbiol Biotechnol 2016; 26: 2019–2029.
Zhang W, Xu C, Bian C, Tempel W, Crombet L, MacKenzie F et al. Crystal structure of the Cys2His2-type zinc finger domain of human DPF2. Biochem Biophys Res Commun 2011; 413: 58–61.
Gray KA, Yates B, Seal RL, Wright MW, Bruford EA . Genenames.org: the HGNC resources in 2015. Nucleic Acids Res 2015; 43 (Database issue): D1079–D1085.
Nunez N, Clifton MM, Funnell AP, Artuz C, Hallal S, Quinlan KG et al. The multi-zinc finger protein ZNF217 contacts DNA through a two-finger domain. J Biol Chem 2011; 286: 38190–38201.
Linke K, Mace PD, Smith CA, Vaux DL, Silke J, Day CL . Structure of the MDM2/MDMX RING domain heterodimer reveals dimerization is required for their ubiquitylation in trans. Cell Death Differ 2008; 15: 841–848.
Borgel J, Tyl M, Schiller K, Pusztai Z, Dooley CM, Deng W et al. KDM2A integrates DNA and histone modification signals through a CXXC/PHD module and direct interaction with HP1. Nucleic Acids Res 2017; 45: 1114–1129.
Sanchez-Garcia I, Rabbitts TH . The LIM domain: a new structural motif found in zinc-finger-like proteins. Trends Genet 1994; 10: 315–320.
Turner CE, Miller JT . Primary sequence of paxillin contains putative SH2 and SH3 domain binding motifs and multiple LIM domains: identification of a vinculin and pp125Fak-binding region. J Cell Sci 1994; 107 (Pt 6): 1583–1591.
Dubrovskyi O, Tian X, Poroyko V, Yakubov B, Birukova AA, Birukov KG . Identification of paxillin domains interacting with beta-catenin. FEBS Lett 2012; 586: 2294–2299.
Smith MA, Blankman E, Deakin NO, Hoffman LM, Jensen CC, Turner CE et al. LIM domains target actin regulators paxillin and zyxin to sites of stress fiber strain. PLoS One 2013; 8: e69378.
Wang J, Wang H, Wang LY, Cai D, Duan Z, Zhang Y et al. Silencing the epigenetic silencer KDM4A for TRAIL and DR5 simultaneous induction and antitumor therapy. Cell Death Differ 2016; 23: 1886–1896.
Cao LL, Wei F, Du Y, Song B, Wang D, Shen C et al. ATM-mediated KDM2A phosphorylation is required for the DNA damage repair. Oncogene 2016; 35: 301–313.
Huang F, Abmayr SM, Workman JL . Regulation of KAT6 acetyltransferases and their roles in cell cycle progression, stem cell maintenance, and human disease. Mol Cell Biol 2016; 36: 1900–1907.
Elton L, Carpentier I, Verhelst K, Staal J, Beyaert R . The multifaceted role of the E3 ubiquitin ligase HOIL-1: beyond linear ubiquitination. Immunol Rev 2015; 266: 208–221.
Ashraf W, Ibrahim A, Alhosin M, Zaayter L, Ouararhni K, Papin C et al. The epigenetic integrator UHRF1: on the road to become a universal biomarker for cancer. Oncotarget 2017; 8: 51946–51962.
Fu M, Blackshear PJ . RNA-binding proteins in immune regulation: a focus on CCCH zinc finger proteins. Nat Rev Immunol 2017; 17: 130–143.
Roth J, Dobbelstein M, Freedman DA, Shenk T, Levine AJ . Nucleo-cytoplasmic shuttling of the hdm2 oncoprotein regulates the levels of the p53 protein via a pathway used by the human immunodeficiency virus rev protein. EMBO J 1998; 17: 554–564.
Tao W, Levine AJ . Nucleocytoplasmic shuttling of oncoprotein Hdm2 is required for Hdm2-mediated degradation of p53. Proc Natl Acad Sci USA 1999; 96: 3077–3080.
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A et al. Proteomics: tissue-based map of the human proteome. Science 2015; 347: 1260419.
Peng Y, Clark KJ, Campbell JM, Panetta MR, Guo Y, Ekker SC . Making designer mutants in model organisms. Development 2014; 141: 4042–4054.
Bibikova M, Beumer K, Trautman JK, Carroll D . Enhancing gene targeting with designed zinc finger nucleases. Science 2003; 300: 764.
Urnov FD, Miller JC, Lee YL, Beausejour CM, Rock JM, Augustus S et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 2005; 435: 646–651.
Bedell VM, Wang Y, Campbell JM, Poshusta TL, Starker CG, Krug RG 2nd et al. In vivo genome editing using a high-efficiency TALEN system. Nature 2012; 491: 114–118.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013; 339: 819–823.
Schichl YM, Resch U, Hofer-Warbinek R, de Martin R . Tristetraprolin impairs NF-kappaB/p65 nuclear translocation. J Biol Chem 2009; 284: 29571–29581.
Prenzler F, Fragasso A, Schmitt A, Munz B . Functional analysis of ZFP36 proteins in keratinocytes. Eur J Cell Biol 2016; 95: 277–284.
Evans PM, Liu C . Roles of Krupel-like factor 4 in normal homeostasis, cancer and stem cells. Acta Biochim Biophys Sin 2008; 40: 554–564.
Patel S, Xi ZF, Seo EY, McGaughey D, Segre JA . Klf4 and corticosteroids activate an overlapping set of transcriptional targets to accelerate in utero epidermal barrier acquisition. Proc Natl Acad Sci USA 2006; 103: 18668–18673.
Segre JA, Bauer C, Fuchs E . Klf4 is a transcription factor required for establishing the barrier function of the skin. Nat Genet 1999; 22: 356–360.
Jaubert J, Cheng J, Segre JA . Ectopic expression of kruppel like factor 4 (Klf4) accelerates formation of the epidermal permeability barrier. Development 2003; 130: 2767–2777.
Sen GL, Boxer LD, Webster DE, Bussat RT, Qu K, Zarnegar BJ et al. ZNF750 is a p63 target gene that induces KLF4 to drive terminal epidermal differentiation. Dev Cell 2012; 22: 669–677.
Boxer LD, Barajas B, Tao S, Zhang J, Khavari PA . ZNF750 interacts with KLF4 and RCOR1, KDM1A, and CTBP1/2 chromatin regulators to repress epidermal progenitor genes and induce differentiation genes. Genes Dev 2014; 28: 2013–2026.
Birnbaum RY, Hayashi G, Cohen I, Poon A, Chen H, Lam ET et al. Association analysis identifies ZNF750 regulatory variants in psoriasis. BMC Med Genet 2011; 12: 167.
Birnbaum RY, Zvulunov A, Hallel-Halevy D, Cagnano E, Finer G, Ofir R et al. Seborrhea-like dermatitis with psoriasiform elements caused by a mutation in ZNF750, encoding a putative C2H2 zinc finger protein. Nat Genet 2006; 38: 749–751.
Shields JM, Christy RJ, Yang VW . Identification and characterization of a gene encoding a gut-enriched Kruppel-like factor expressed during growth arrest. J Biol Chem 1996; 271: 20009–20017.
McConnell BB, Ghaleb AM, Nandan MO, Yang VW . The diverse functions of Kruppel-like factors 4 and 5 in epithelial biology and pathobiology. Bioessays 2007; 29: 549–557.
Zhang W, Chen X, Kato Y, Evans PM, Yuan S, Yang J et al. Novel cross talk of Kruppel-like factor 4 and beta-catenin regulates normal intestinal homeostasis and tumor repression. Mol Cell Biol 2006; 26: 2055–2064.
Yu T, Chen X, Lin T, Liu J, Li M, Zhang W et al. KLF4 deletion alters gastric cell lineage and induces MUC2 expression. Cell Death Dis 2016; 7: e2255.
Katz JP, Perreault N, Goldstein BG, Lee CS, Labosky PA, Yang VW et al. The zinc-finger transcription factor Klf4 is required for terminal differentiation of goblet cells in the colon. Development 2002; 129: 2619–2628.
Bosse T, Piaseckyj CM, Burghard E, Fialkovich JJ, Rajagopal S, Pu WT et al. Gata4 is essential for the maintenance of jejunal-ileal identities in the adult mouse small intestine. Mol Cell Biol 2006; 26: 9060–9070.
Beuling E, Baffour-Awuah NY, Stapleton KA, Aronson BE, Noah TK, Shroyer NF et al. GATA factors regulate proliferation, differentiation, and gene expression in small intestine of mature mice. Gastroenterology 2011; 140: 1219–1229 e1211-1212.
Nagandla H, Lopez S, Yu W, Rasmussen TL, Tucker HO, Schwartz RJ et al. Defective myogenesis in the absence of the muscle-specific lysine methyltransferase SMYD1. Dev Biol 2016; 410: 86–97.
Hayashi S, Manabe I, Suzuki Y, Relaix F, Oishi Y . Klf5 regulates muscle differentiation by directly targeting muscle-specific genes in cooperation with MyoD in mice. Elife 2016; 5: e17462.
Li G, Ye X, Peng X, Deng Y, Yuan W, Li Y et al. CXXC5 regulates differentiation of C2C12 myoblasts into myocytes. J Muscle Res Cell Motil 2014; 35: 259–265.
Kurosaka M, Ogura Y, Funabashi T, Akema T . Early growth response 3 (Egr3) contributes a maintenance of C2C12 myoblast proliferation. J Cell Physiol 2017; 232: 1114–1122.
Li K, Zhang J, Ren JJ, Wang Q, Yang KY, Xiong ZJ et al. A novel zinc finger protein Zfp637 behaves as a repressive regulator in myogenic cellular differentiation. J Cell Biochem 2010; 110: 352–362.
Meruvu S, Hugendubler L, Mueller E . Regulation of adipocyte differentiation by the zinc finger protein ZNF638. J Biol Chem 2011; 286: 26516–26523.
Park HS, Ju UI, Park JW, Song JY, Shin DH, Lee KH et al. PPARgamma neddylation essential for adipogenesis is a potential target for treating obesity. Cell Death Differ 2016; 23: 1296–1311.
Li Q, Peng H, Fan H, Zou X, Liu Q, Zhang Y et al. The LIM protein Ajuba promotes adipogenesis by enhancing PPARgamma and p300/CBP interaction. Cell Death Differ 2016; 23: 158–168.
Perez-Mancera PA, Bermejo-Rodriguez C, Gonzalez-Herrero I, Herranz M, Flores T, Jimenez R et al. Adipose tissue mass is modulated by SLUG (SNAI2). Hum Mol Genet 2007; 16: 2972–2986.
Tong Q, Tsai J, Tan G, Dalgin G, Hotamisligil GS . Interaction between GATA and the C/EBP family of transcription factors is critical in GATA-mediated suppression of adipocyte differentiation. Mol Cell Biol 2005; 25: 706–715.
Fidalgo M, Shekar PC, Ang YS, Fujiwara Y, Orkin SH, Wang J . Zfp281 functions as a transcriptional repressor for pluripotency of mouse embryonic stem cells. Stem Cells 2011; 29: 1705–1716.
Fidalgo M, Faiola F, Pereira CF, Ding J, Saunders A, Gingold J et al. Zfp281 mediates Nanog autorepression through recruitment of the NuRD complex and inhibits somatic cell reprogramming. Proc Natl Acad Sci USA 2012; 109: 16202–16207.
Jen J, Wang YC . Zinc finger proteins in cancer progression. J Biomed Sci 2016; 23: 53.
Kim SH, Kim EJ, Hitomi M, Oh SY, Jin X, Jeon HM et al. The LIM-only transcription factor LMO2 determines tumorigenic and angiogenic traits in glioma stem cells. Cell Death Differ 2015; 22: 1517–1525.
Pieraccioli M, Nicolai S, Antonov A, Somers J, Malewicz M, Melino G et al. ZNF281 contributes to the DNA damage response by controlling the expression of XRCC2 and XRCC4. Oncogene 2016; 35: 2592–2601.
Malewicz M . The role of 53BP1 protein in homology-directed DNA repair: things get a bit complicated. Cell Death Differ 2016; 23: 1902–1903.
Mistrik M, Bartek J . What a 'Ku'incidence!: parallel discoveries of a new DNA repair factor. Cell Death Differ 2015; 22: 888–889.
Li J, Ao J, Li K, Zhang J, Li Y, Zhang L et al. ZNF32 contributes to the induction of multidrug resistance by regulating TGF-beta receptor 2 signaling in lung adenocarcinoma. Cell Death Dis 2016; 7: e2428.
Nicolai S, Rossi A, Di Daniele N, Melino G, Annicchiarico-Petruzzelli M, Raschella G . DNA repair and aging: the impact of the p53 family. Aging (Albany NY) 2015; 7: 1050–1065.
Zhang L, Zhou Y, Cheng C, Cui H, Cheng L, Kong P et al. Genomic analyses reveal mutational signatures and frequently altered genes in esophageal squamous cell carcinoma. Am J Hum Genet 2015; 96: 597–611.
Agostini M, Knight RA . miR-34: from bench to bedside. Oncotarget 2014; 5: 872–881.
Liu YW, Sun M, Xia R, Zhang EB, Liu XH, Zhang ZH et al. LincHOTAIR epigenetically silences miR34a by binding to PRC2 to promote the epithelial-to-mesenchymal transition in human gastric cancer. Cell Death Dis 2015; 6: e1802.
Hermeking H . The miR-34 family in cancer and apoptosis. Cell Death Differ 2010; 17: 193–199.
Lin DC, Hao JJ, Nagata Y, Xu L, Shang L, Meng X et al. Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat Genet 2014; 46: 467–473.
Hazawa M, Lin DC, Handral H, Xu L, Chen Y, Jiang YY et al. ZNF750 is a lineage-specific tumour suppressor in squamous cell carcinoma. Oncogene 2017; 36: 2243–2254.
Moon YW, Rao G, Kim JJ, Shim HS, Park KS, An SS et al. LAMC2 enhances the metastatic potential of lung adenocarcinoma. Cell Death Differ 2015; 22: 1341–1352.
Marinkovich MP . Tumour microenvironment: laminin 332 in squamous-cell carcinoma. Nat Rev Cancer 2007; 7: 370–380.
Otsuka R, Akutsu Y, Sakata H, Hanari N, Murakami K, Kano M et al. ZNF750 expression as a novel candidate biomarker of chemoradiosensitivity in esophageal squamous cell carcinoma. Oncology 2017; 93: 197–203.
Furukawa D, Chijiwa T, Matsuyama M, Mukai M, Matsuo EI, Nishimura O et al. Zinc finger protein 185 is a liver metastasis-associated factor in colon cancer patients. Mol Clin Oncol 2014; 2: 709–713.
Vanaja DK, Cheville JC, Iturria SJ, Young CY . Transcriptional silencing of zinc finger protein 185 identified by expression profiling is associated with prostate cancer progression. Cancer Res 2003; 63: 3877–3882.
Gonzalez HE, Gujrati M, Frederick M, Henderson Y, Arumugam J, Spring PW et al. Identification of 9 genes differentially expressed in head and neck squamous cell carcinoma. Arch Otolaryngol Head Neck Surg 2003; 129: 754–759.
Medina PP, Carretero J, Ballestar E, Angulo B, Lopez-Rios F, Esteller M et al. Transcriptional targets of the chromatin-remodelling factor SMARCA4/BRG1 in lung cancer cells. Hum Mol Genet 2005; 14: 973–982.
Heiss NS, Gloeckner G, Bachner D, Kioschis P, Klauck SM, Hinzmann B et al. Genomic structure of a novel LIM domain gene (ZNF185) in Xq28 and comparisons with the orthologous murine transcript. Genomics 1997; 43: 329–338.
Zhang JS, Gong A, Young CY . ZNF185, an actin-cytoskeleton-associated growth inhibitory LIM protein in prostate cancer. Oncogene 2007; 26: 111–122.
Taniuchi T, Mortensen ER, Ferguson A, Greenson J, Merchant JL . Overexpression of ZBP-89, a zinc finger DNA binding protein, in gastric cancer. Biochem Biophys Res Commun 1997; 233: 154–160.
Strasberg Rieber M, Zangemeister-Wittke U, Rieber M . p53-Independent induction of apoptosis in human melanoma cells by a bcl-2/bcl-xL bispecific antisense oligonucleotide. Clin Cancer Res 2001; 7: 1446–1451.
Serova M, Calvo F, Lokiec F, Koeppel F, Poindessous V, Larsen AK et al. Characterizations of irofulven cytotoxicity in combination with cisplatin and oxaliplatin in human colon, breast, and ovarian cancer cells. Cancer Chemother Pharmacol 2006; 57: 491–499.
Bandres E, Malumbres R, Cubedo E, Honorato B, Zarate R, Labarga A et al. A gene signature of 8 genes could identify the risk of recurrence and progression in Dukes' B colon cancer patients. Oncol Rep 2007; 17: 1089–1094.
Siavoshian S, Segain JP, Kornprobst M, Bonnet C, Cherbut C, Galmiche JP et al. Butyrate and trichostatin A effects on the proliferation/differentiation of human intestinal epithelial cells: induction of cyclin D3 and p21 expression. Gut 2000; 46: 507–514.
Moran A, Iniesta P, de Juan C, Garcia-Aranda C, Diaz-Lopez A, Benito M . Impairment of stromelysin-1 transcriptional activity by promoter mutations in high microsatellite instability colorectal tumors. Cancer Res 2005; 65: 3811–3814.
Juncker-Jensen A, Romer J, Pennington CJ, Lund LR, Almholt K . Spontaneous metastasis in matrix metalloproteinase 3-deficient mice. Mol Carcinog 2009; 48: 618–625.
Tang CH, Yamamoto A, Lin YT, Fong YC, Tan TW . Involvement of matrix metalloproteinase-3 in CCL5/CCR5 pathway of chondrosarcomas metastasis. Biochem Pharmacol 2010; 79: 209–217.
Law DJ, Du M, Law GL, Merchant JL . ZBP-99 defines a conserved family of transcription factors and regulates ornithine decarboxylase gene expression. Biochem Biophys Res Commun 1999; 262: 113–120.
Dawson MI, Park JH, Chen G, Chao W, Dousman L, Waleh N et al. Retinoic acid (RA) receptor transcriptional activation correlates with inhibition of 12-O-tetradecanoylphorbol-13-acetate-induced ornithine decarboxylase (ODC) activity by retinoids: a potential role for trans-RA-induced ZBP-89 in ODC inhibition. Int J Cancer 2001; 91: 8–21.
Zhang X, Diab IH, Zehner ZE . ZBP-89 represses vimentin gene transcription by interacting with the transcriptional activator, Sp1. Nucleic Acids Res 2003; 31: 2900–2914.
Wieczorek E, Lin Z, Perkins EB, Law DJ, Merchant JL, Zehner ZE . The zinc finger repressor, ZBP-89, binds to the silencer element of the human vimentin gene and complexes with the transcriptional activator, Sp1. J Biol Chem 2000; 275: 12879–12888.
Feng Y, Wang X, Xu L, Pan H, Zhu S, Liang Q et al. The transcription factor ZBP-89 suppresses p16 expression through a histone modification mechanism to affect cell senescence. FEBS J 2009; 276: 4197–4206.
Essien BE, Sundaresan S, Ocadiz-Ruiz R, Chavis A, Tsao AC, Tessier AJ et al. Transcription factor ZBP-89 drives a feedforward loop of beta-catenin expression in colorectal cancer. Cancer Res 2016; 76: 6877–6887.
Oliner JD, Saiki AY, Caenepeel S . The role of MDM2 amplification and overexpression in tumorigenesis. Cold Spring Harb Perspect Med 2016; 6: (pii: a026336).
Fischer M . Census and evaluation of p53 target genes. Oncogene 2017; 36: 3943–3956.
Zhang J, Huang K, O'Neill KL, Pang X, Luo X . Bax/Bak activation in the absence of Bid, Bim, Puma, and p53. Cell Death Dis 2016; 7: e2266.
Shamseddine AA, Clarke CJ, Carroll B, Airola MV, Mohammed S, Rella A et al. P53-dependent upregulation of neutral sphingomyelinase-2: role in doxorubicin-induced growth arrest. Cell Death Dis 2015; 6: e1947.
Wu X, Bayle JH, Olson D, Levine AJ . The p53-mdm-2 autoregulatory feedback loop. Genes Dev 1993; 7: 1126–1132.
Freedman DA, Wu L, Levine AJ . Functions of the MDM2 oncoprotein. Cell Mol Life Sci 1999; 55: 96–107.
Juven-Gershon T, Oren M . Mdm2: the ups and downs. Mol Med 1999; 5: 71–83.
Jones SN, Hancock AR, Vogel H, Donehower LA, Bradley A . Overexpression of Mdm2 in mice reveals a p53-independent role for Mdm2 in tumorigenesis. Proc Natl Acad Sci USA 1998; 95: 15608–15612.
Lundgren K, Montes de Oca Luna R, McNeill YB, Emerick EP, Spencer B, Barfield CR et al. Targeted expression of MDM2 uncouples S phase from mitosis and inhibits mammary gland development independent of p53. Genes Dev 1997; 11: 714–725.
Weaver J, Downs-Kelly E, Goldblum JR, Turner S, Kulkarni S, Tubbs RR et al. Fluorescence in situ hybridization for MDM2 gene amplification as a diagnostic tool in lipomatous neoplasms. Mod Pathol 2008; 21: 943–949.
Weaver J, Goldblum JR, Turner S, Tubbs RR, Wang WL, Lazar AJ et al. Detection of MDM2 gene amplification or protein expression distinguishes sclerosing mesenteritis and retroperitoneal fibrosis from inflammatory well-differentiated liposarcoma. Mod Pathol 2009; 22: 66–70.
Schlott T, Reimer S, Jahns A, Ohlenbusch A, Ruschenburg I, Nagel H et al. Point mutations and nucleotide insertions in the MDM2 zinc finger structure of human tumours. J Pathol 1997; 182: 54–61.
Lindstrom MS, Jin A, Deisenroth C, White Wolf G, Zhang Y . Cancer-associated mutations in the MDM2 zinc finger domain disrupt ribosomal protein interaction and attenuate MDM2-induced p53 degradation. Mol Cell Biol 2007; 27: 1056–1068.
Wu W, Xu C, Ling X, Fan C, Buckley BP, Chernov MV et al. Targeting RING domains of Mdm2-MdmX E3 complex activates apoptotic arm of the p53 pathway in leukemia/lymphoma cells. Cell Death Dis 2015; 6: e2035.
Wang S, Zhao Y, Aguilar A, Bernard D, Yang CY . Targeting the MDM2-p53 protein–protein interaction for new cancer therapy: progress and challenges. Cold Spring Harb Perspect Med 2017; 7: (pii: a026245).
Haupt S, Buckley D, Pang JM, Panimaya J, Paul PJ, Gamell C et al. Targeting Mdmx to treat breast cancers with wild-type p53. Cell Death Dis 2015; 6: e1821.
Lamouille S, Xu J, Derynck R . Molecular mechanisms of epithelial–mesenchymal transition. Nat Rev Mol Cell Biol 2014; 15: 178–196.
Zhang P, Sun Y, Ma L . ZEB1: at the crossroads of epithelial–mesenchymal transition, metastasis and therapy resistance. Cell Cycle 2015; 14: 481–487.
Li X, Gao D, Wang H, Li X, Yang J, Yan X et al. Negative feedback loop between p66Shc and ZEB1 regulates fibrotic EMT response in lung cancer cells. Cell Death Dis 2015; 6: e1708.
Shi Y, Sawada J, Sui G, Affar el B, Whetstine JR, Lan F et al. Coordinated histone modifications mediated by a CtBP co-repressor complex. Nature 2003; 422: 735–738.
Sanchez-Tillo E, Lazaro A, Torrent R, Cuatrecasas M, Vaquero EC, Castells A et al. ZEB1 represses E-cadherin and induces an EMT by recruiting the SWI/SNF chromatin-remodeling protein BRG1. Oncogene 2010; 29: 3490–3500.
Thakur AK, Nigri J, Lac S, Leca J, Bressy C, Berthezene P et al. TAp73 loss favors Smad-independent TGF-beta signaling that drives EMT in pancreatic ductal adenocarcinoma. Cell Death Differ 2016; 23: 1358–1370.
Heldin CH, Vanlandewijck M, Moustakas A . Regulation of EMT by TGFbeta in cancer. FEBS Lett 2012; 586: 1959–1970.
Zhang GJ, Zhou T, Tian HP, Liu ZL, Xia SS . High expression of ZEB1 correlates with liver metastasis and poor prognosis in colorectal cancer. Oncol Lett 2013; 5: 564–568.
Shen A, Zhang Y, Yang H, Xu R, Huang G . Overexpression of ZEB1 relates to metastasis and invasion in osteosarcoma. J Surg Oncol 2012; 105: 830–834.
Lakoma A, Barbieri E, Agarwal S, Jackson J, Chen Z, Kim Y et al. The MDM2 small-molecule inhibitor RG7388 leads to potent tumor inhibition in p53 wild-type neuroblastoma. Cell Death Discov 2015; 1.
Kim ES, Shohet JM . Reactivation of p53 via MDM2 inhibition. Cell Death Dis 2015; 6: e1936.
Spaderna S, Schmalhofer O, Wahlbuhl M, Dimmler A, Bauer K, Sultan A et al. The transcriptional repressor ZEB1 promotes metastasis and loss of cell polarity in cancer. Cancer Res 2008; 68: 537–544.
Liu Y, Zhang N, Wang Y, Xu M, Liu N, Pang X et al. Zinc finger E-box binding homeobox 1 promotes invasion and bone metastasis of small cell lung cancer in vitro and in vivo. Cancer Sci 2012; 103: 1420–1428.
Markowitz JA, Tinkle MB, Fischbeck KH . Spinal muscular atrophy in the neonate. J Obstet Gynecol Neonatal Nurs 2004; 33: 12–20.
Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 1995; 80: 155–165.
Helmken C, Hofmann Y, Schoenen F, Oprea G, Raschke H, Rudnik-Schoneborn S et al. Evidence for a modifying pathway in SMA discordant families: reduced SMN level decreases the amount of its interacting partners and Htra2-beta1. Hum Genet 2003; 114: 11–21.
Ahmad S, Wang Y, Shaik GM, Burghes AH, Gangwani L . The zinc finger protein ZPR1 is a potential modifier of spinal muscular atrophy. Hum Mol Genet 2012; 21: 2745–2758.
Pao PC, Huang NK, Liu YW, Yeh SH, Lin ST, Hsieh CP et al. A novel RING finger protein, Znf179, modulates cell cycle exit and neuronal differentiation of P19 embryonal carcinoma cells. Cell Death Differ 2011; 18: 1791–1804.
Ko CY, Chang LH, Lee YC, Sterneck E, Cheng CP, Chen SH et al. CCAAT/enhancer binding protein delta (CEBPD) elevating PTX3 expression inhibits macrophage-mediated phagocytosis of dying neuron cells. Neurobiol Aging 2012; 33: 422 e411–422 e425.
Li R, Strohmeyer R, Liang Z, Lue LF, Rogers J . CCAAT/enhancer binding protein delta (C/EBPdelta) expression and elevation in Alzheimer's disease. Neurobiol Aging 2004; 25: 991–999.
Shin JH, Ko HS, Kang H, Lee Y, Lee YI, Pletinkova O et al. PARIS (ZNF746) repression of PGC-1alpha contributes to neurodegeneration in Parkinson's disease. Cell 2011; 144: 689–702.
Tanaka K, Suzuki T, Hattori N, Mizuno Y . Ubiquitin, proteasome and parkin. Biochim Biophys Acta 2004; 1695: 235–247.
Cohen I, Birnbaum RY, Leibson K, Taube R, Sivan S, Birk OS . ZNF750 is expressed in differentiated keratinocytes and regulates epidermal late differentiation genes. PLoS One 2012; 7: e42628.
Nakanishi G, Kim YS, Nakajima T, Jetten AM . Regulatory role for Kruppel-like zinc-finger protein Gli-similar 1 (Glis1) in PMA-treated and psoriatic epidermis. J Invest Dermatol 2006; 126: 49–60.
Kim YS, Lewandoski M, Perantoni AO, Kurebayashi S, Nakanishi G, Jetten AM . Identification of Glis1, a novel Gli-related, Kruppel-like zinc finger protein containing transactivation and repressor functions. J Biol Chem 2002; 277: 30901–30913.
Senee V, Chelala C, Duchatelet S, Feng D, Blanc H, Cossec JC et al. Mutations in GLIS3 are responsible for a rare syndrome with neonatal diabetes mellitus and congenital hypothyroidism. Nat Genet 2006; 38: 682–687.
Watanabe N, Hiramatsu K, Miyamoto R, Yasuda K, Suzuki N, Oshima N et al. A murine model of neonatal diabetes mellitus in Glis3-deficient mice. FEBS Lett 2009; 583: 2108–2113.
Dimitri P, Warner JT, Minton JA, Patch AM, Ellard S, Hattersley AT et al. Novel GLIS3 mutations demonstrate an extended multisystem phenotype. Eur J Endocrinol 2011; 164: 437–443.
Molkentin JD, Lin Q, Duncan SA, Olson EN . Requirement of the transcription factor GATA4 for heart tube formation and ventral morphogenesis. Genes Dev 1997; 11: 1061–1072.
Basson CT, Bachinsky DR, Lin RC, Levi T, Elkins JA, Soults J et al. Mutations in human TBX5 [corrected] cause limb and cardiac malformation in Holt-Oram syndrome. Nat Genet 1997; 15: 30–35.
Li QY, Newbury-Ecob RA, Terrett JA, Wilson DI, Curtis AR, Yi CH et al. Holt-Oram syndrome is caused by mutations in TBX5, a member of the Brachyury (T) gene family. Nat Genet 1997; 15: 21–29.
Pikkarainen S, Tokola H, Kerkela R, Ruskoaho H . GATA transcription factors in the developing and adult heart. Cardiovasc Res 2004; 63: 196–207.
Durocher D, Charron F, Warren R, Schwartz RJ, Nemer M . The cardiac transcription factors Nkx2-5 and GATA-4 are mutual cofactors. EMBO J 1997; 16: 5687–5696.
Moore ML, Wang GL, Belaguli NS, Schwartz RJ, McMillin JB . GATA-4 and serum response factor regulate transcription of the muscle-specific carnitine palmitoyltransferase I beta in rat heart. J Biol Chem 2001; 276: 1026–1033.
Kodo K, Nishizawa T, Furutani M, Arai S, Yamamura E, Joo K et al. GATA6 mutations cause human cardiac outflow tract defects by disrupting semaphorin-plexin signaling. Proc Natl Acad Sci USA 2009; 106: 13933–13938.
Koutsourakis M, Langeveld A, Patient R, Beddington R, Grosveld F . The transcription factor GATA6 is essential for early extraembryonic development. Development 1999; 126: 723–732.
Lepore JJ, Mericko PA, Cheng L, Lu MM, Morrisey EE, Parmacek MS . GATA-6 regulates semaphorin 3C and is required in cardiac neural crest for cardiovascular morphogenesis. J Clin Invest 2006; 116: 929–939.
Maitra M, Koenig SN, Srivastava D, Garg V . Identification of GATA6 sequence variants in patients with congenital heart defects. Pediatr Res 2010; 68: 281–285.
Acknowledgements
This work has been supported by the Medical Research Council, UK; grants from Associazione Italiana per la Ricerca contro il Cancro (AIRC): AIRC 2014 IG15653 (to GM), AIRC 5xmille MCO9979 (to GM), Fondazione Roma malattie Non trasmissibili Cronico-Degenerative (NCD) Grant (to GM).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Cassandri, M., Smirnov, A., Novelli, F. et al. Zinc-finger proteins in health and disease. Cell Death Discov. 3, 17071 (2017). https://doi.org/10.1038/cddiscovery.2017.71
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/cddiscovery.2017.71
This article is cited by
-
Endogenous feline leukemia virus long terminal repeat integration site diversity is highly variable in related and unrelated domestic cats
Retrovirology (2024)
-
Metallothionein-3 is a multifunctional driver that modulates the development of sorafenib-resistant phenotype in hepatocellular carcinoma cells
Biomarker Research (2024)
-
Single-cell sequencing reveals increased LAMB3-positive basal keratinocytes and ZNF90-positive fibroblasts in autologous cultured epithelium
Communications Biology (2024)
-
Proteolysis-targeting chimeras with reduced off-targets
Nature Chemistry (2024)
-
Evaluation of the Concentration of Selected Elements in the Serum of Patients with Degenerative Stenosis of the Lumbosacral Spine
Biological Trace Element Research (2024)