Abstract
Neuronal apoptosis induced by survival factor deprivation is strongly regulated at the transcriptional level. Notably, the nuclear factor of activated T cell (NFAT) transcription factors have an important role in the control of the survival/death fate of neurons. However, the mechanisms that regulate NFAT activity in response to apoptotic stimuli and the target genes that mediate their effect on neuronal apoptosis are mostly unknown. In a previous study, we identified Trim17 as a crucial E3 ubiquitin ligase that is necessary and sufficient for neuronal apoptosis. Here, we show that Trim17 binds preferentially SUMOylated forms of NFATc3. Nonetheless, Trim17 does not promote the ubiquitination/degradation of NFATc3. NFAT transcription factors are regulated by calcium/calcineurin-dependent nuclear-cytoplasmic shuttling. Interestingly, Trim17 reduced by twofold the calcium-mediated nuclear localization of NFATc3 and, consistent with this, halved NFATc3 activity, as estimated by luciferase assays and by measurement of target gene expression. Trim17 also inhibited NFATc4 nuclear translocation and activity. NFATc4 is known to induce the expression of survival factors and, as expected, overexpression of NFATc4 protected cerebellar granule neurons from serum/KCl deprivation-induced apoptosis. Inhibition of NFATc4 by Trim17 may thus partially mediate the proapoptotic effect of Trim17. In contrast, overexpression of NFATc3 aggravated neuronal death, whereas knockdown of NFATc3 protected neurons from apoptosis. This proapoptotic effect of NFATc3 might be due to a feedback loop in which NFATc3, but not NFATc4, induces the transcription of the proapoptotic gene Trim17. Indeed, we found that overexpression or silencing of NFATc3, respectively, increased or decreased Trim17 levels, whereas NFATc4 had no significant effect on Trim17 expression. Moreover, we showed that NFATc3 binds to the promoter of the Trim17 gene together with c-Jun. Therefore, our results describe a novel mechanism regulating NFAT transcription factors beyond the calcium/calcineurin-dependent pathway and provide a possible explanation for the opposite effects of NFATc3 and NFATc4 on neuronal apoptosis.
Similar content being viewed by others
Main
Neuronal apoptosis is crucial for normal development of the nervous system and aberrant apoptosis may participate in both acute and chronic neurodegenerative diseases.1, 2 Apoptosis is robustly regulated at the transcriptional level in neurons.3 Indeed, transcription inhibitors have been shown to prevent neuronal death in several in vitro models,4, 5, 6 and many transcription factors controlling neuronal apoptosis have been identified. Notably, the nuclear factor of activated T cell (NFAT) transcription factors have an important role in the development of the nervous system7, 8 and in the control of the survival/death fate of neurons.9, 10, 11, 12, 13, 14
The NFAT family comprises four calcium/calcineurin-dependent transcription factors that are encoded by four closely related genes.15, 16, 17 NFAT proteins are expressed in most mammalian tissues, with the different members of the family being present in distinct but overlapping sets of cell types.18 NFATc3 (NFAT4/NFATx) and NFATc4 (NFAT3) are the predominant NFAT genes expressed in neurons.10, 19 Owing to their high sequence similarity, NFAT proteins have relatively redundant functions. However, non-redundant roles are evident in the phenotypes observed in individual NFAT knockout mice.18 NFAT-dependent gene regulation mediates a wide variety of cellular processes, such as survival, apoptosis, differentiation and proliferation. Both NFATc3 and NFATc4 have been shown to have either proapoptotic or antiapoptotic effects, depending on the physiologic and cellular context.9, 10, 11, 12, 13, 14, 20, 21, 22 However, the mechanisms that regulate their activity in response to apoptotic stimuli and the target genes that mediate their differential effects on neuronal apoptosis are mostly unknown.
Under resting conditions, NFATs are heavily phosphorylated, which results in their cytosolic retention. Upon increase in intracellular calcium, the calcium/calmodulin-dependent protein phosphatase calcineurin is activated and dephosphorylates NFATs leading to their nuclear import.15, 16, 17 Once inside the nucleus, NFATs cooperate with multiple transcriptional partners, including activator protein 1 (AP-1), to regulate gene expression. Nuclear import of NFATs is opposed by rapid export induced by rephosphorylation mediated by several protein kinases.16 Although the critical role of phosphorylation/dephosphorylation on NFAT activity is widely accepted, the exact mechanism of cytoplasmic retention of phosphorylated NFAT transcription factors is poorly understood. SUMOylation was shown to have an important role in regulating nuclear localization and activity of NFATc1 (NFAT2/NFATc)23 and NFATc2 (NFAT1/NFATp).24 However, the mechanisms mediating these effects of SUMO (small ubiquitin-like modifier) are mostly unknown.
Here we describe a novel mechanism regulating the activity of NFATc3 beyond the calcium/calcineurin-dependent pathway. We found that NFATc3 interacted in a SUMO-dependent manner with Trim17, an E3 ubiquitin ligase necessary for neuronal apoptosis.25 Although Trim17 did not induce NFATc3 ubiquitination, this interaction inhibited the activity of NFATc3 by preventing its nuclear localization. Moreover, we found that NFATc3 had a proapoptotic effect in cerebellar granule neurons (CGNs), whereas NFATc4 was neuroprotective. This may be due to a negative feedback loop in which NFATc3, but not NFATc4, induced the expression of Trim17. Taken together, our data not only provide new insight into the mechanisms of action and regulation of the proapoptotic protein Trim17 but also give a possible explanation for the opposite effects of NFATc3 and NFATc4 on neuronal apoptosis.
Results
Trim17 interacts with NFATc3 in a SUMO-dependent manner
In a previous study, we showed that Trim17 is both necessary and sufficient for neuronal apoptosis.25 To investigate the mechanisms of action of Trim17, a yeast two-hybrid (Y2H) screen was performed by Hybrigenics SA (Paris, France) using full-length mouse Trim17 as bait, and a mouse embryo brain (E10.5–E12.5) cDNA library as prey. One positive clone was identified as NFATc3 (Figure 1a). We confirmed this interaction, using an in situ proximity ligation assay (PLA), between overexpressed Trim17 and endogenous NFATc3 in Neuro2A cells, and showed that it occurs predominantly in the cytoplasm. Treating the cells with the calcium ionophore A23187 had no significant effect on the interaction (Figure 1b). We next studied the Trim17/NFATc3 interaction by co-immunoprecipitation. Trim17-GFP (green fluorescent protein) was, furthermore, co-immunoprecipitated with both hemagglutinin tag (HA)-NFATc3 (Figure 1c, left panel) and HA-NFATc4 (Figure 1c, right panel), using anti-HA antibody. In both cases, the interaction was not modified by A23187 (Figure 1c). Inversely, both HA-NFATc3 and HA-NFATc4 were precipitated together with Trim17-GFP using GFP-Trap beads (Figure 1d). Importantly, the NFATc3 fragment isolated in the Y2H screen contains a SUMO consensus site (Figure 1a). Moreover, Trim17 and NFATc3 could be efficiently co-precipitated only in the presence of N-ethylmaleimide (NEM), a potent alkylating agent that inhibits SUMO-specific proteases and deubiquitinases, or SUMO2-VS, a more specific inhibitor of SUMO-specific proteases (Figure 1d, left panel). In contrast, the interaction between Trim17 and NFATc4 was not altered in the absence of NEM or when the cell lysate was treated with the SUMO-specific protease SENP1 (Figure 1d, right panel). Therefore, these data suggest that Trim17 interacts with NFATc3 in a SUMO-dependent manner, whereas its interaction with NFATc4 is SUMO-independent.
Trim17 interacts with NFATc3 and NFATc4. (a) Schematic representation of mouse NFATc3 depicting its different domains (TAD: strong transactivation domain; NHR: NFAT-homology region, regulatory domain containing many serine/threonine residues that can be phosphorylated by various protein kinases; RHR: Rel-homology DNA-binding region), its three SUMOylation consensus sites (*) and the cDNA clone isolated in the Y2H screen containing the coding region for amino acids 681–858 (Y2H clone). (b) Neuro2A cells were transfected with Flag-Trim17 for 24 h and its interaction with endogenous NFATc3 detected by in situ PLA, using anti-Flag and anti-NFATc3 antibodies. Each bright spot represents one interaction between the two proteins. Negative control was obtained by omitting anti-Flag antibody. When indicated (+A23187), cells were deprived of serum for 3 h, and were subsequently treated with 1 μM A23187 and 100 nM PMA in serum-free medium for 2 h. 4',6-Diamidino-2-phenylindole (DAPI) staining and merge pictures indicate that the interaction takes place mostly in the cytoplasm. (c) Neuro2A cells were transfected with Trim17-GFP together with either HA-NFATc3, HA-NFATc4 or pCS2-HA empty vector (as a negative control) for 24 h. Cells were then treated for 5 h with 20 μM MG-132. When indicated (A23187), cells were deprived of serum for 3 h, and were subsequently treated with 1 μM A23187 and 100 nM PMA in serum-free medium for 2 h. The cells were subsequently harvested and lysates were subjected to immunoprecipitation, in the presence of 10 mM NEM and phosphatase inhibitors, using anti-HA antibody. Immunoprecipitates and total lysates were analyzed by western blot using anti-HA and anti-GFP antibodies. (d) Neuro2A cells were transfected with GFP (negative control) or Trim17-GFP together with either HA-NFATc3 (left panel) or HA-NFATc4 (right panel) for 24 h, treated for 6 h with 20 μM MG-132 and then harvested and subjected to immunoprecipitation using GFP-trap beads, in lysis buffer containing 50 mM NEM, 400 ng recombinant SUMO-specific protease SENP1-GST or 5 μM SUMO2-VS as indicated, in the presence of phosphatase inhibitors. Immunoprecipitates and total lysates were analyzed by western blot using anti-HA and anti-GFP antibodies
SUMOylation is known to occur on an acceptor Lys within a ΨKX(E/D) consensus motif, where Ψ is a large hydrophobic residue.26 The environments of Lys435, Lys704 and Lys1013 of mouse NFATc3 fit this consensus with high confidence scores, and are, furthermore, conserved between mouse and human (Figure 1a). In addition, the Lys704 and Lys1013 motifs are present in other members of the NFAT family (Figure 2a) and are, de facto, SUMOylated in human NFATc123 and NFATc2.24 To determine whether Lys435, Lys704 and Lys1013 of murine NFATc3 can be SUMOylated, they were mutated to Arg. The resulting NFATc3 K/R mutant constructs were subjected to in vitro SUMOylation using SUMO1. Immunoblotting of wild-type (WT) NFATc3 revealed two higher-molecular bands, which likely correspond, respectively, to mono- and di-SUMOylated forms (Figure 2b). The K435R substitution did not significantly affect NFATc3 in vitro SUMOylation. In contrast, the di-SUMOylated band of NFATc3 disappeared when either Lys704 or Lys1013 was mutated, and almost no SUMOylation of NFATc3 could be detected when they were both replaced by Arg (Figure 2b), suggesting these two lysine residues to be the main SUMOylation sites of NFATc3. The different SUMO site-deficient constructs of NFATc3 were tested in co-immunoprecipitation with Trim17. The interaction between the two proteins was not significantly altered by single K/R mutations, whereas it was strongly decreased by Lys704 and Lys1013 double substitution (Figure 2c). This suggests that SUMOylation of Lys704 and Lys1013 in NFATc3 is necessary for its interaction with Trim17.
Trim17 interacts preferentially with the SUMOylated forms of NFATc3. (a) SUMOylation consensus motifs of murine NFATc3 and corresponding sequences in murine NFATc1, NFATc2 and NFATc4. SUMOylation consensus motifs are in bold type, seuences identical to NFATc3 are underlined. (b) Proteins produced by the in vitro transcription–translation of the various HA-NFATc3 constructs were subjected to in vitro SUMOylation in the presence or the absence of recombinant SUMO1. Reaction products were analyzed by western blot using anti-HA antibody. Because SUMO1 does not form polySUMO chains, the two bands that migrate with an apparent molecular weight greater than that of unmodified NFATc3 likely correspond to mono- and di-SUMOylated forms of HA-NFATc3 (indicated, respectively, by * and **). (c and d) BHK cells were transfected with the different variants of HA-NFATc3 together with WT Trim17-GFP or GFP (as a negative control) (c), or with WT HA-NFATc3 together with the different variants of Trim17-GFP or GFP (d). After 24 h, cells were treated for 8 h with 10 μM MG-132, before harvesting. GFP-fusion proteins were immunoprecipitated in the presence of 10 mM NEM and phosphatase inhibitors using GFP-Trap beads. Immunoprecipitates and lysates were analyzed by western blot using anti-HA and anti-GFP antibodies
Proteins interacting with SUMO generally harbor SUMO-interacting motifs (SIMs) that typically consist of a hydrophobic core containing 3–4 aliphatic residues (most often Val and/or Ile).27 We identified two conserved motifs fulfilling these criteria in Trim17, which we named SIM1 (117-ICVV-120) and SIM2 (390-VVQL-393), respectively. We generated Trim17 constructs in which the two Val residues in an individual or both SIMs were replaced by Ala residues (mSIM1, mSIM2 and mSIM1-2, respectively). These mutations are predicted to disrupt their SUMO-binding ability.28 We found that Trim17(mSIM1) and Trim17(ΔBB), which is deleted of the B-Box domain containing SIM1, co-immunoprecipitated with NFATc3 with a similar efficiency as WT Trim17 (Figure 2d). In contrast, mutation of SIM2, or both SIMs, strongly decreased the interaction of Trim17 with NFATc3 (Figure 2d). Taken together, these data further confirm that the interaction between Trim17 and NFATc3 is SUMO-dependent.
Trim17 is not a physiologic E3 ubiquitin ligase for NFATc3
As Trim17 is an E3 ubiquitin ligase,25, 29, 30 we examined whether Trim17 could mediate NFATc3 ubiquitination. Unexpectedly, the protein level of NFATc3 progressively increased when co-transfected with increasing amounts of Trim17 in Neuro2A cells (Supplementary Figure 1a). Consistently, when co-transfected with His-tagged ubiquitin, the ubiquitination level of NFATc3 was decreased by Trim17 in a dose-dependent manner but was not significantly altered by an inactive mutant of Trim17 (ΔRING; Supplementary Figure 1b). These results thus indicate that Trim17 does not promote the ubiquitin-mediated degradation of NFATc3 but rather prevents it.
Trim17 favors the cytoplasmic localization of NFATc3 and NFATc4
Because NFATc3 is known to be regulated by nuclear-cytoplasmic shuttling, we examined the effect of Trim17 on its subcellular localization. As expected, WT NFATc3 expressed in baby hamster kidney (BHK) cells was predominantly cytoplasmic (Figure 3a, left panel). Upon addition of the calcium ionophore A23187, NFATc3 translocated into the nucleus in the presence of GFP but remained mainly in the cytoplasm when coexpressed with Trim17-GFP (Figure 3a, right panel). Indeed, NFATc3 was detected in the nucleus of about 70% of cells in the presence of GFP versus 40% in the presence of Trim17-GFP, following A23187 treatment (Figure 3b). Interestingly, the inhibitory effect of Trim17 on NFATc3 nuclear translocation was abolished by deletion of its RING domain or mutation of its SIM2 motif (Figure 3c), as well as by mutation of the different SUMOylation sites in NFATc3 (Figure 3d). These results suggest that the E3 ubiquitin-ligase activity of Trim17 conferred by its RING domain25 and the SUMO-dependent interaction between Trim17 and NFATc3 are involved in the cytoplasmic retention of NFATc3 mediated by Trim17. However, the K435R substitution was sufficient to abolish the effect of Trim17 on the nuclear translocation of NFATc3 (Figure 3d), whereas it did not significantly alter the SUMOylation of NFATc3 (Figure 2b) nor the Trim17/NFATc3 interaction (Figure 2c). It is thus possible that an undetectable alteration of NFATc3 SUMOylation, resulting in an undetectable reduction of its interaction with Trim17, might have a significant impact on the Trim17-mediated cytoplasmic retention of NFATc3. Alternatively, the K435R substitution might influence NFATc3 nuclear translocation independently of SUMOylation or Trim17.
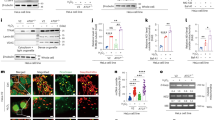
Trim17 prevents NFATc3 nuclear localization in BHK cells. (a) BHK cells were transfected with HA-NFATc3 together with GFP (negative control) or Trim17-GFP for 24 h. They were then deprived of serum for 3 h and treated with 1 μM of the calcium ionophore A23187 in serum-free medium for an additional 30 min before fixation. NFATc3 was detected using an anti-HA antibody and visualized by confocal microscopy. GFP or Trim17-GFP was visualized by GFP fluorescence and nuclei were stained with 4',6-diamidino-2-phenylindole (DAPI). (b) Quantification of the nuclear localization of NFATc3 in the experiment presented in (a). The percentage of cells showing NFATc3 mainly in the nucleus was determined among the population of cells expressing both HA-NFATc3 and Trim17-GFP. Data are the means±S.E.M. of four independent experiments. ***P<0.001 significantly different from the corresponding value obtained in cells transfected with GFP instead of Trim17-GFP (one-way analysis of variance (ANOVA) followed by Bonferroni multiple comparison test). (c and d) BHK cells were transfected with WT HA-NFATc3 together with GFP or different forms of Trim17-GFP (c) or with different forms of HA-NFATc3 together with GFP or WT Trim17-GFP (d). After 24 h, cells were treated as in (a). Nuclear localization of NFATc3 was determined as in (b). Data are the means±S.E.M. of at least three independent experiments. **P<0.01; *P<0.05 significantly different from the control GFP (one-way ANOVA followed by Dunnett’s multiple comparison test) (c). ***P<0.001 significantly different from the corresponding value obtained in cells transfected with GFP instead of Trim17-GFP (one-way ANOVA followed by Bonferroni multiple comparison test) (d)
We confirmed the effect of Trim17 on NFATc3 subcellular localization in primary cultures of CGNs, in which we initially identified the proapoptotic effect of Trim17.25 Indeed, endogenous NFATc3 accumulated in the cytoplasm of neurons transfected with Trim17-GFP (Figure 4a). Interestingly, overexpression of Trim17-mCherry also resulted in the accumulation of NFATc4-GFP in the cytoplasm of transfected neurons (Figure 4b). These data thus suggest that Trim17 favors the cytoplasmic localization of both NFATc3 and NFATc4.
Trim17 prevents NFATc3 and NFATc4 nuclear localization in primary neurons. (a) CGN primary cultures were transfected with GFP (as a negative control) or Trim17-GFP for 16 h. Following fixation, endogenous NFATc3 was detected by immunofluorescence, GFP or Trim17-GFP was visualized by GFP fluorescence and nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). Coverslips were analyzed by confocal microscopy. Note that endogenous NFATc3 accumulates in the cytoplasm upon Trim17 overexpression. (b) CGNs were transfected with NFATc4-GFP together with mCherry (as a negative control) or Trim17-mCherry for 16 h. Proteins were visualized by GFP or mCherry fluorescence and nuclei were stained with DAPI. Coverslips were analyzed by confocal microscopy. Note that NFATc4-GFP accumulates in the cytoplasm upon Trim17 overexpression. The effect of Trim17 on the subcellular localization of endogenous NFATc4 could not be assessed because overexpression of Trim17 significantly decreased the level of endogenous NFATc4
Trim17 inhibits the activity of NFATc3 and NFATc4
To test whether the effect of Trim17 on the nuclear translocation of NFATc3 has an impact on its activity, we performed NFAT-luciferase expression reporter assays. As expected, overexpression of NFATc3 increased luciferase activity in BHK cells transfected with the NFAT-luciferase reporter plasmid and subsequently treated with A23187 and phorbol 12-myristate 13-acetate (PMA) (Figure 5a). Importantly, co-transfection of WT Trim17 decreased the NFATc3-mediated luciferase expression in a dose-dependent manner, whereas Trim17(mSIM1-2) and Trim17(ΔRING) had no significant effect (Figure 5a). Moreover, Trim17 overexpression provoked a twofold decrease in the mRNA levels of endogenous brain-derived neurotrophic factor (BDNF) triggered by NFATc3 (Figure 5b), confirming that Trim17 inhibits NFATc3 activity. Interestingly, Trim17 overexpression also provoked a twofold decrease in the reporter gene activity mediated by NFATc4 (Figure 5c).
Trim17 inhibits the activity of NFATc3 and NFATc4. (a) BHK cells were transfected with an NFAT-luciferase reporter plasmid (pNFAT-TA, 300 ng) and pRL-TK (100 ng), together with empty plasmid (−) or pCS2-HA-NFATc3 (200 ng), and with increasing amounts of pCI-Trim17-GFP (0, 200, 400 and 600 ng), or pCI-Trim17(mSIM1-2)-GFP (100 ng), or pCI-Trim17(ΔRING)-GFP (100 ng). After 24 h, cells were deprived of serum for 8 h, and were subsequently treated with 1 μM A23187 and 100 nM PMA in serum-free medium for 30 min before measurement of luciferase activity. Luciferase activity was expressed relative to that assessed in cells transfected with pNFAT-TA alone. Data are the means±S.E.M. of three independent experiments. **P<0.01 significantly different from the value from cells transfected with NFATc3 in the absence of Trim17 (one-way analysis of variance (ANOVA) followed by Dunnett’s multiple comparison test). (b) Neuro2A cells were transfected with HA-NFATc3, Trim17-GFP or both for 24 h and then deprived of serum for 5 h. Total RNA was extracted and the mRNA level of Bdnf was estimated by quantitative RT-PCR. Fold change was calculated by comparison with cells transfected with empty plasmid. Data are the means±S.E.M. of three independent experiments. **P<0.01, *P<0.05, significantly different from cells transfected with NFATc3 alone (one-way ANOVA followed by Student–Newman–Keuls multiple comparison test). (c) BHK cells were transfected with pNFAT-TA (300 ng) and pRL-TK (100 ng) together with either empty plasmid, pCS2-HA-NFATc3 (200 ng) or pCS2-HA-NFATc4 (200 ng) in the presence or the absence of pCI-Trim17-GFP (400 ng). Cells were treated and luciferase activity was measured as in (a). Data are the means±S.E.M. of three independent experiments. *P<0.05 significantly different from the corresponding value from cells transfected with GFP instead of Trim17-GFP (one-way ANOVA followed by Bonferroni multiple comparison test)
NFATc3 and NFATc4 have opposite effects on neuronal apoptosis
NFATc4 has been shown to sustain survival in developing CGNs,9 cortical neurons10 and hippocampal neurons.11 In contrast, the role of NFATc3 in neuronal survival has been poorly studied. To address this point, we first overexpressed NFATc3 in primary cultures of CGNs. These cultures represent one of the best-characterized in vitro models of neuronal apoptosis.31 Cultured CGNs survive in the presence of serum and depolarizing levels of extracellular KCl (25 mM) that mimic the neuronal activity required for developing CGN survival in vivo.32 Withdrawal of serum, accompanied by the lowering of KCl to 5 mM (K5), triggers CGN apoptosis,4 recapitulating the programmed cell death that occurs in the cerebellum during postnatal development.33 As previously reported by others,9 we found that NFATc4 is located in the nucleus of CGNs in survival conditions and accumulates in the cytoplasm following serum/KCl deprivation (Supplementary Figures 2b and 3b). Interestingly, we showed that NFATc3 is also nuclear in survival conditions, but remains in the nucleus even after serum/KCl deprivation (Supplementary Figures 2a and 3a).
In CGNs maintained in the initial culture medium (control), overexpression of NFATc3-GFP or NFATc4-GFP did not significantly induce apoptosis, although NFATc3-GFP slightly increased basal apoptosis (Figure 6a). Importantly, following 8 h serum/KCl deprivation, NFATc3-GFP significantly aggravated apoptosis, whereas NFATc4-GFP clearly protected neurons (Figure 6a). Consistently, we found that transduction with a lentivirus expressing a short hairpin RNA (shRNA) against NFATc3 strongly protected CGNs from serum/KCl deprivation (Figure 6b). In contrast, NFATc4 deletion11 or knockdown10 has been reported to induce neuronal apoptosis, notably in CGNs.9 Taken together, these data suggest that NFATc4 has a neuroprotective role, whereas NFATc3 induces neuronal apoptosis.
Differential effects of NFATc3 and NFATc4 on neuronal apoptosis. (a) CGN primary cultures were transfected with GFP (as a negative control), NFATc3-GFP or NFATc4-GFP for 16 h. Then, neurons were switched to serum-free medium containing 5 mM KCl (K5) for 8 h or were left untreated (control). Following fixation, nuclei were visualized by with 4′,6-diamidino-2-phenylindole (DAPI) staining and proteins fused to GFP were detected by fluorescent microscopy. Arrows indicate GFP-positive neurons with thick arrows for neurons undergoing apoptosis and thin arrows for healthy neurons. Scale bar is 20 μM. The percentage of transfected, GFP-positive neurons undergoing apoptosis was assessed by examining cell morphology and nuclear condensation. Data are the means±S.E.M. of four independent experiments. **P<0.01 significantly different from the corresponding value obtained in neurons transfected with GFP (one-way analysis of variance (ANOVA) followed by Bonferroni multiple comparison test). (b) CGNs were transduced with lentiviral particles expressing a non-targeting control shRNA (sh-ctrl) or an shRNA specifically targeting NFATc3 (sh-NFATc3) one day after plating. At 6 DIV, neurons were incubated for 8 h in K5 medium, fixed and stained with DAPI. The percentage of apoptotic neurons was estimated by examining nuclear condensation. Data are the means±S.E.M. of three independent experiments. ***P<0.001 was significantly different from neurons transduced with the control shRNA (one-way ANOVA followed by Bonferroni multiple comparison test)
NFATc3, but not NFATc4, partially mediates Trim17 induction during CGN apoptosis
The unexpected proapoptotic effect of NFATc3 could be due to the transcriptional induction of a strong proapoptotic protein. In fact, we observed a clear increase in the mRNA levels of endogenous Trim17 in Neuro2A cells treated with A23187 and PMA (Figure 7a, left panel), suggesting that Trim17 expression could be stimulated by NFAT transcription factors. Consistently, overexpression of NFATc3, but not NFATc4, increased Trim17 mRNA levels in Neuro2A cells treated with A23187 and PMA (Figure 7a, right panel). Moreover, the protein level of endogenous Trim17 also increased in CGN transfected with NFATc3-GFP, but not with NFATc4-GFP, both in control and apoptotic conditions (Figure 7b).
Expression of Trim17 is induced by NFATc3 but not by NFATc4. (a) Left panel: Neuro2A cells were deprived of serum for 3 h, and were subsequently treated with 1 μM A23187 and 100 nM PMA in serum-free medium for 2 h or left in serum-free medium (control). Right panel: Neuro2A cells were transfected with empty plasmid (control), HA-NFATc3 or HA-NFATc4 for 24 h, deprived of serum for 3 h and then treated with 1 μM A23187 and 100 nM PMA in serum-free medium for 2 h. Following the different treatments, total RNA was extracted and the mRNA levels of Trim17 was estimated by quantitative RT-PCR. Fold change was calculated by comparison with control cells. Data are the means±S.E.M. of three independent experiments. **P<0.01 was significantly different from control cells (one-way analysis of variance (ANOVA) followed by Dunnett’s multiple comparison test). (b) CGN primary cultures were transfected with GFP (negative control), NFATc3-GFP or NFATc4-GFP for 16 h, and then switched to K5 medium for 5 h or maintained in the initial culture medium (control). Following fixation, Trim17 expression was measured by immunofluorescence. The intensity of Trim17-associated fluorescence (AU: arbitrary unit) was measured specifically in GFP-positive neurons. Data are the means±S.E.M. of three independent experiments. (c and d) CGNs were transduced with lentiviral particles expressing a control shRNA or shRNA sequences targeting specifically NFATc3 (c) or NFATc4 (d), one day after plating. At DIV6, neurons were incubated for 4 h in K5 medium or maintained in the initial culture medium (control). Total RNA was extracted and the mRNA levels of Trim17, Nfatc3 (c) and Nfatc4 (d) were estimated by quantitative RT-PCR. Fold change was calculated by comparison with neurons transduced with control shRNA and maintained in control medium. Data are the means±S.E.M. of three independent experiments. ***P<0.001 significantly different from neurons transduced with the control shRNA in the corresponding condition (one-way ANOVA followed by Student–Newman–Keuls multiple comparison test)
We initially identified Trim17 as one of the most highly upregulated genes in CGN following KCl deprivation.34 To determine whether NFATc3 could be involved in this transcriptional induction, CGNs were transduced with two different shRNAs, which efficiently reduced endogenous NFATc3 expression (Figure 7c, left panel). Importantly, the increase in Trim17 mRNA levels triggered by serum/KCl deprivation was significantly reduced by these two shRNAs (Figure 7c, right panel). In contrast, the knocking down of NFATc4 did not impair the induction of Trim17 during CGN apoptosis (Figure 7d). Consistently, knocking down of NFATc3 (Supplementary Figure 4a), but not NFATc4 (Supplementary Figure 4b), significantly decreased endogenous Trim17 expression in Neuro2A cells. Taken together, these data suggest that Trim17 is a target gene of NFATc3 but not of NFATc4.
Trim17 is a target gene of NFATc3 and c-Jun
NFATs generally act in cooperation with unrelated transcription factors to regulate gene expression.15 AP-1, which consists of heterodimers of the Fos and Jun family proteins, is the main transcriptional partner of NFATs. The NFAT:Fos:Jun complexes contact an ≈15–20 bp stretch of DNA, in which the NFAT and AP-1 elements are apposed to create a ‘composite’ site.35 Analysis of the Trim17 promoter sequences allowed us to identify a region around the transcription start site containing two conserved AP-1 binding sites and one conserved composite NFAT:AP-1 element (Figure 8a).
Trim17 is a target gene of NFAT and c-Jun transcription factors. (a) Alignment of the mouse and human genomic sequences around the transcription start site of mouse Trim17. The region shown is entirely included in exon 1 of human TRIM17. The asterisks indicate identical nucleotides. Two potential AP-1 binding sites and one potential composite NFAT-AP-1 site that are conserved between mouse and human are boxed. Sequences of the primers used for quantitative PCR of ChIP samples are highlighted. (b) CGN primary cultures were transduced with lentiviral particles expressing the shRNA NFATc3 no. 2 (+) or a control shRNA (−), one day after plating. At DIV6, neurons were incubated for 4 h in the initial culture medium (control) or in K5 medium, both containing either 10 μM SP600125 (+) or the same volume of dimethyl sulfoxide (DMSO) (−). Total RNA was extracted and the mRNA levels of Trim17, NFATc3 and c-Jun were estimated by quantitative RT-PCR. Fold change was calculated by comparison with neurons transduced with control shRNA and maintained in control medium containing only DMSO. Data are the means±S.D. of triplicate determinations and are representative of three independent experiments. (c) CGNs were incubated for 4 h in the initial culture medium (control) or in K5 medium before cross-linking. ChIP assays were performed using antibodies specific to NFATc3, c-Jun or H3K4me3. Rabbit IgG were used as a negative control. Quantitative PCR was carried out on the immunoprecipitates and the input chromatin using primers flanking the transcription start site of mouse Trim17 (described in a). The amount of Trim17 promoter DNA precipitated by each antibody is expressed as the percentage of the input used for each immunoprecipitation. Data are representative of three independent experiments. (d) Proposed model for the reciprocal regulation of NFATc3 and Trim17 in neuronal apoptosis. Trim17 inhibits both NFATc3 and NFATc4 by preventing their nuclear translocation. In the case of NFATc3, this effect depends on the SUMO-dependent interaction between Trim17 and NFATc3. The inhibition NFATc4 by Trim17 could contribute to the proapoptotic effect of Trim17, as we have shown that NFATc4 promotes neuronal survival. Conversely, NFATc3 overexpression induces neuronal apoptosis. This could be due to a feedback loop in which NFATc3 induces the transcription of the Trim17 gene in cooperation with c-Jun. The black arrow indicates induction of transcription. Gray lines indicate inhibition by prevention of nuclear localization
The c-Jun N-terminal kinase (JNK)/c-Jun signaling pathway is known to have a crucial role in neuronal apoptosis in different models,3 notably in CGNs deprived of serum and KCl.36 To determine whether c-Jun cooperate with NFATc3 in the transcriptional regulation of Trim17, we measured Trim17 mRNA levels in CGNs treated with the JNK inhibitor SP600125, following transduction with control or NFATc3-specific shRNAs, in control and apoptotic conditions. Interestingly, JNK inhibition prevented Trim17 induction after serum/KCl deprivation, to a similar extent as NFATc3 knockdown. Moreover, this effect was further increased when JNK was inhibited and NFATc3 was silenced at the same time (Figure 8b). These results support the idea that NFATc3 and c-Jun cooperate to increase the expression of Trim17 during neuronal apoptosis.
To demonstrate that c-Jun and NFATc3 indeed bind the Trim17 regulatory region that we identified, we performed chromatin immunoprecipitation (ChIP) assays using primary CGNs. As expected, we could amplify the region described in Figure 8a following immunoprecipitation with antibodies against NFATc3 and c-Jun but not with control IgG. Interestingly, the binding of NFATc3 and c-Jun to the Trim17 promoter strongly increased following serum/KCl deprivation (Figure 8c). The trimethylation of Lys4 of histone H3 (H3K4me3) associated with the Trim17 regulatory region was also enhanced in apoptotic neurons (Figure 8c), consistently with the strong increase in Trim17 mRNA levels in apoptotic CGNs (Figure 8b). Taken together, our results strongly suggest that NFATc3 induces the transcription of the proapoptotic gene Trim17 in association with c-Jun, possibly explaining why NFATc3 promotes neuronal apoptosis (Figure 6).
Discussion
Trim17 is a member of the TRIM family that represents one of the largest classes of single-protein RING-containing E3 ubiquitin ligases.37, 38 We have previously shown that Trim17 is necessary for neuronal apoptosis25 and contributes to Mcl-1 ubiquitination in CGNs.29 In the present study, we provide further insight into the mechanism of action and the transcriptional regulation of Trim17.
We found that Trim17 bound and inhibited NFATc3 and NFATc4 transcription factors. Interestingly, the interaction between Trim17 and NFATc3 appears to be SUMO-dependent as it was impaired either by mutations of SUMOylation consensus sites of NFATc3 or by alteration of Trim17 SIMs. Importantly, Trim17 provoked a twofold decrease in the calcium-mediated nuclear localization of NFATc3. This effect seems to depend on the physical interaction between Trim17 and NFATc3, because it was abolished by mutations that prevented the co-immunoprecipitation of the two proteins. Trim17 also induced a twofold decrease in NFATc3 activity, as measured by luciferase assays and the increase in BDNF mRNA levels. Moreover, Trim17(mSIM1-2) and Trim17(ΔRING), which were unable to prevent NFATc3 nuclear translocation, did not modify NFATc3 activity. Taken together, these results strongly suggest that Trim17 inhibits NFATc3 by binding it and thereby preventing its nuclear localization. Although the interaction between Trim17 and NFATc4 appeared to be SUMO-independent, similar mechanisms seem to mediate the inhibition of NFATc4 by Trim17, as we found that Trim17 also favors its cytoplasmic localization. As such, we describe a novel mechanism regulating the activity of NFATc3 and NFATc4 beyond calcium-calcineurin signaling.
Data obtained by others in knockdown experiments9, 10 or in NFATc4-deficient mice11 suggest that NFATc4 sustains survival in different types of neurons, notably by inducing the transcription of survival factors.10 Consistent with this, we showed that transfection of NFATc4 protected CGNs from apoptosis. The inhibition of the prosurvival factor NFATc4 by Trim17 may thus partially mediate the proapoptotic effect of Trim17 in CGNs (Figure 8d). Although the role of NFATc3 is less documented, Vashishta et al.10 noted that knockdown of NFATc3 protected cortical neurons from phosphatidyl inositol-3-OH kinase (PI3K) inhibition-induced apoptosis. Here, we show that silencing of NFATc3 protects CGNs from serum/KCl deprivation. Moreover, we found that overexpression of NFATc3 aggravated apoptosis. Therefore, our data further confirm that NFATc3 favors neuronal apoptosis, whereas NFATc4 protects neurons from survival factor withdrawal-induced apoptosis. More importantly, we provide a possible explanation for these opposite effects. Indeed, we found that NFATc3 overexpression increased Trim17 levels, and that NFATc3 knockdown reduced the induction of Trim17 during CGN apoptosis. Moreover, we found that NFATc3 binds the promoter of the Trim17 gene together with c-Jun, and cooperates with c-Jun to induce the transcription of Trim17. In contrast, overexpression or knocking down of NFATc4 had no significant effect on Trim17 level. Therefore, these data strongly suggest that NFATc3 induces neuronal apoptosis by increasing the expression of the proapoptotic protein Trim17 (Figure 8d).
The first question raised by this model is how can NFATc3 be activated after serum/KCl withdrawal. Indeed, the protein kinases glycogen synthase kinase 3 (GSK3), JNK and p38, which promote NFAT nuclear export,15, 16, 17 have all been shown to be activated in CGNs deprived of serum and KCl.29, 36, 39, 40, 41 Moreover, KCl deprivation should result in calcineurin inactivation by reducing voltage-gated calcium influx. Therefore, NFAT transcription factors should be highly phosphorylated and maintained in the cytoplasm of CGNs following serum/KCl deprivation. This has actually been shown for NFATc4,9 and we have confirmed these results in our culture conditions for both endogenous and overexpressed NFATc4. However, our results also clearly show that NFATc3 contributes to the increase in Trim17 mRNA levels 4 h after serum/KCl deprivation, indicating that NFATc3 is still active in these conditions. Consistently, we found that NFATc3 is still located in the nucleus of CGNs up to 8 h after serum/KCl deprivation. Our results thus suggest that NFATc3 and NFATc4 are differentially regulated in response to the same stimulus. This is in line with an elegant recent study reporting that NFATc3 translocates to the nucleus upon depolarization of dorsal root ganglion (DRG) and hippocampal neurons, more rapidly, with a higher rate and in response to lower KCl concentrations than NFATc4.42 Moreover, knocking down of GSK3 did not significantly affect NFATc3 nuclear import, whereas it strongly increased nuclear localization of NFATc4 upon depolarization.42 This is consistent with our results showing that NFATc3 is still active in deprived CGN, in spite of GSK3 activation.
The second question is how NFATc3 and NFATc4, which share very similar DNA-binding domains,15, 35 can have differential effects on Trim17 expression. This could be because of the relative expression of NFATc3 and NFATc4. Indeed, we found that the mRNA level of NFATc3 is 10 times higher than that of NFATc4 in CGNs (not shown), as previously reported in cortical10 and DRG19 neurons. In addition, NFATc4 has been found to be inactivated in CGNs deprived of serum and KCl,9 and we confirmed that NFATc4 is excluded from the nucleus in these conditions. Finally, distinct NFAT proteins may have differential affinities for various nuclear partners, thereby controlling distinct sets of genes.
We have shown that both NFATc3 and c-Jun bind a regulatory region of the Trim17 gene around the transcription start site. Moreover, we previously showed that GSK3 inhibition blocked both apoptosis and the increase of Trim17 mRNA levels in deprived CGNs.25, 29 Other studies reported that GSK3 is required for the full induction of AP-1-responsive proapoptotic target genes in CGNs.43, 44 Therefore, it is possible that GSK3 induces Trim17 expression by promoting c-Jun activity. The strong induction of Trim17 during CGN apoptosis may thus result from the combination of JNK/c-Jun, PI3K/Akt/GSK3 and NFATc3 signaling pathways integrated at the level of NFAT:AP-1 sites in the Trim17 gene.
Materials and Methods
Materials
SUMO2-VS was purchased from Boston Biochem (Cambridge, MA, USA). Culture media were from GIBCO Life Technologies (Carlsbad, CA, USA). Poly-D-lysine was from Becton Dickinson Biosciences (Franklin Lakes, NJ, USA). Fetal calf serum, other culture reagents, protease inhibitor cocktail, DAPI, NEM, MG-132, PMA, A23187 and other chemicals were from Sigma-Aldrich (St. Louis, MO, USA). Protein G-agarose and protein A-agarose beads were from Roche (Basel, Switzerland). GFP-Trap-A beads were from Chromotek (Planegg-Martinsried, Germany). Rat monoclonal anti-HA antibody (clone 3F10) and mouse monoclonal anti-GFP antibody (clones 7.1 and 13.1) were from Roche. Mouse monoclonal antibody against actin (clone C4) was from Chemicon International (Billerica, MA, USA; MAB1501). Rat monoclonal anti-Flag antibody (clone M2), mouse monoclonal anti-tubulin (clone DM1A) and rabbit IgG were from Sigma-Aldrich (F3165, T6199 and I5006, respectively). Rabbit polyclonal antibody against mouse NFATc3 (M-75) and NFATc4 (H-74) were from Santa Cruz (Dallas, TX, USA; sc-8321 and sc-13036, respectively). Rabbit monoclonal antibody against c-Jun (60A8) was from Cell Signaling Technology (Denvers, MA, USA; no. 9165). Rabbit polyclonal antibody against H3K4me3 was from Abcam (Cambridge, MA, USA; ab8580). Fluorescent and horseradish peroxidase-conjugated goat anti-rabbit, anti-rat and anti-mouse secondary antibodies were from Molecular Probes-Life Technologies (Carlsbad, CA, USA) and Jackson ImmunoResearch Laboratories Inc., (West Grove, PA, USA) respectively. The HIV-derived lentiviral vectors pLKO.1 containing control shRNAs (SHC002, SHC003, SHC005) and the shRNAs TRCN0000377122 (shRNA NFATc3 no. 1), TRCN0000377129 (shRNA NFATc3 no. 2), TRCN0000232310 (shRNA NFATc4 no. 1) and TRCN0000232313 (shRNA NFATc4 no. 2) were obtained from Sigma-Aldrich.
Y2H screen
Y2H screening was performed by Hybrigenics SA (http://www.hybrigenics-services.com). The coding sequence for amino acids 2–447 of the mouse Trim17 protein (GenBank accession number gi: 141802118 ) was PCR amplified and cloned into pB27 as a C-terminal fusion to LexA (N-LexA-Trim17-C). The construct was checked by sequencing the entire insert and used as a bait to screen a random-primed mouse embryo brain (E10.5–E12.5) cDNA library constructed into pP6. pB27 and pP6 derive from the original pBTM11645 and pGADGH46 plasmids, respectively. Eighty-four million clones (11-fold the complexity of the library) were screened using a mating approach with Y187 (matα) and L40ΔGal4 (mata) yeast strains as described previously.47 Eighty-five His+ colonies were selected on a medium lacking tryptophan, leucine and histidine. The prey fragments of the positive clones were amplified by PCR and sequenced at their 5′ and 3′ junctions. The resulting sequences were used to identify the corresponding interacting proteins in the GenBank database (NCBI) using a fully automated procedure. A confidence score (PBS, for Predicted Biological Score) was attributed to each interaction as previously described.48
Protein sequence analysis
The sequence of the proteins NFATc3 and Trim17 were analyzed using the prediction software JASSA (Joined Advanced SUMOylation site and SIM Analyser, http://www.jassa.sitesgb.info/), to identify consensus SUMOylation sites and putative SUMO-interacting motifs.
Cell cultures and transient transfection
Lenti-X 293 T (Clontech), Neuro2A and BHK cell lines were grown in Dulbecco’s modified Eagle’s medium containing 4.5 g/l glucose supplemented with 5–10% fetal bovine serum and penicillin–streptomycin 100 IU/ml–100 μg/ml. Cells were transfected with GenJet in vitro transfection reagent (version II, SignaGen Laboratories, Ijamsville, MD, USA) preoptimized and conditioned for transfecting Neuro2A and BHK-21 cells, respectively according to the manufacturer’s instructions.
Primary cultures of CGNs were prepared from 7-day-old murine pups (C57Bl/6 J mice) as described previously.25 Briefly, freshly dissected cerebella were dissociated by trypsinization and mechanical disruption, and plated in basal medium Eagle (BME) supplemented with 10% fetal bovine serum, 2 mM L-Gln, 10 mM HEPES, penicillin–streptomycin 100 IU/ml–100 μg/ml and 20 mM KCl. Primary CGNs, grown in 24-well plates, were transfected at 5 days in vitro (DIV) with 2 μg of plasmids using a calcium phosphate protocol optimized for neuronal cultures as previously described.25
In situ PLA
Neuro2A cells, seeded onto gelatin-coated glass coverslips, were transfected with pCI-Flag-Trim17 for 24 h. Following the indicated treatment, cells were fixed with 4% paraformaldehyde for 20 min, washed with PBS and permeabilized with 0.2% Triton X-100 in PBS for 10 min, at room temperature. The interaction between overexpressed Flag-Trim17 and endogenous NFATc3 was detected using the Duolink In Situ Kit (Olink Bioscience, Uppsala, Sweden), according to the manufacturer’s instructions. Briefly, cells were successively incubated with blocking solution for 30 min at 37 °C, with primary antibodies against Flag (1 : 200) and NFATc3 (1 : 100) overnight at 4°C and with secondary antibodies conjugated with oligonucleotides (PLA probe MINUS and PLA probe PLUS) for 1 h at 37 °C. The cells were then incubated with two connector oligonucleotides together with DNA ligase for 30 min at 37 °C. If the two secondary antibodies are in close proximity, this step allows the connector oligonucleotides to hybridize to the PLA probes and form a circular DNA strand after ligation. Incubation, for 100 min at 37 °C, with DNA polymerase consequently leads to rolling circle amplification, the products of which are detected using fluorescently labelled complementary oligonucleotides. Cells were washed with Duolink In Situ Wash Buffers following each of these steps. In the last wash, 1 μg/ml DAPI was added to the cells for 5 min at room temperature to stain the nuclei. Coverslips were mounted onto glass slides with Mowiol (polyvinyl alcohol 4-88; Sigma-Aldrich) and analyzed by confocal fluorescence microscopy.
Co-immunoprecipitation and western blot analysis
Following transfection with the indicated plasmids for 24 h, Neuro2A or BHK cells were incubated for 6–8 h with 10–20 μM MG-132. They were then homogenized in lysis buffer A (50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 10 mM NaF, 5 mM sodium pyrophosphate, 25 mM β-glycerophosphate, 5 mM EDTA, 20 μM MG-132 and protease inhibitor cocktail) containing 1% of NP-40 for immunoprecipitation with anti-HA and 0.5% of NP-40 for immunoprecipitation with GFP-Trap-A. For anti-HA immunoprecipitation, cell lysates (500 μl) were centrifuged at 300 × g for 5 min at 4 °C. The resulting supernatants were precleared by rotation for 1 h at 4 °C with 10 μl protein G-agarose beads and then rotated for 2 h at 4 °C with 20 μl protein G-agarose beads previously incubated with 4 μl anti-HA antibody. The beads were recovered by centrifugation and washed four times with lysis buffer A containing 0.5 M NaCl for HA-NFATc4 and 0.3 M NaCl for HA-NFATc3 (instead of 150 mM NaCl). For GFP-Trap precipitation, cell lysates (200 μl) were diluted with 300 μl dilution buffer (10 mM Tris-HCl (pH 7.5), 150 mM NaCl, 10 mM NaF, 5 mM sodium pyrophosphate, 25 mM β-glycerophosphate, 0.5 mM EDTA, 20 μM MG-132 and protease inhibitor cocktail) and cell debris was removed by centrifugation. Resulting supernatants were rotated for 2 h at 4 °C with 10–25 μl GFP-Trap-A beads to immunoprecipitate proteins fused to GFP. Beads were recovered by centrifugation and washed four times with dilution buffer. In most conditions, lysis buffer A and dilution buffer were supplemented with 10 mM NEM. Material bound to the protein G-agarose or GFP-Trap beads was eluted by the addition of 3 × Laemmli sample buffer and incubation at 95 °C for 5 min. Precipitated proteins were separated by 10% SDS–PAGE and transferred to Immobilon-P PVDF membrane (Millipore, Billerica, MA, USA). Blocking, probing with antibodies and visualization of immunoreactive proteins were performed as previously described.25
In vitro SUMOylation of NFATc3
NFATc3 and the different K/R mutants were first transcribed and translated in vitro. For this, 2 μg of the different pCS2-HA-NFATc3 constructs were incubated for 2 h at 30 °C in 50 μl in the TNT SP6-coupled wheat germ extract system (Promega, Fitchburg, WI, USA), according to the instructions of the manufacturer. Five microlitres of the in vitro translation reaction were incubated for 30 min at 37 °C in the presence of 3 μg recombinant SUMO1, 150 ng recombinant His-tagged Aos1/Uba2 (E1 enzyme), 100 ng recombinant Ubc9 (E2 enzyme) and 300 ng recombinant GST-PIASxα (E3 enzyme) in 20 μl shift assay buffer (20 mM HEPES (pH 7.3), 110 mM KOAc, 2 mM Mg(OAc)2, 0.5 mM EGTA, 1 mM DTT, 0.05% Tween-20, 0.2 mg/ml ovalbumin, 1 μg/ml leupeptin, 1 μg/ml aprotinin, 1 μg/ml pepstatin) supplemented with 1 mM ATP. Recombinant proteins were produced and purified as previously described.49 Reaction products were separated by SDS-PAGE, transferred to PVDF membranes and analyzed by western blot using anti-HA antibody.
Immunofluorescence and subcellular localization
For immunofluorescence, BHK cells were seeded onto gelatin-coated glass coverslips and primary CGNs were cultured on coverslips previously coated with laminin (16.67 μg/ml) and poly-D-lysine (33.3 μg/ml). BHK cells and neurons were transfected and treated, as described in the figure legends, and then fixed with 4% paraformaldehyde. Overexpressed HA-NFATc3, endogenous NFATc3 and endogenous NFATc4 were detected using anti-HA (1 : 500), anti-NFATc3 (1 : 100) and anti-NFATc4 (1 : 100) antibodies, respectively, as described previously.25 GFP- and mCherry-fused proteins were visualized by fluorescence and nuclei were stained with DAPI. Coverslips were analyzed by conventional or confocal microscopy, as mentioned in the figure legends. Image acquisition and analysis were performed on work stations of the Montpellier RIO imaging facility. For quantification of NFATc3 nuclear localization, BHK cells with predominant cytoplasmic or nuclear localization of NFATc3 were counted, in a blinded manner, among double GFP/HA-positive cells. At least 200 double-positive cells were scored for each experiment and each condition.
NFAT-luciferase expression reporter assay
BHK cells plated in 24-well plates were co-transfected with 300 ng of the NFAT-luciferase reporter plasmid (firefly luciferase under the control of tandem triple copies of the NFAT-binding motif; pNFAT-TA-luciferase; Clontech), 100 ng of a plasmid encoding a Renilla reniformis luciferase under the control of a constitutive HSV-thymidine kinase K promoter (pRL-TK (Promega), to control for transfection efficiency) and different combinations of plasmids expressing HA-NFATc3 (200 ng) or HA-NFATc4 (200 ng) and Trim17-GFP or its mutated forms, as described. One day after transfection, cells were cultured in the absence of serum for 8 h before treatment with 1 μM A23187 and 100 nM PMA for 30 min. They were subsequently lysed and analyzed using the Dual-Luciferase Assay Kit (Promega), according to the manufacturer’s instructions. NFAT-luciferase reporter expression was normalized to that of Renilla luciferase. Data are presented relative to the luciferase activity measured in cells transfected only with the pNFAT-TA-luciferase and pRL-TK plasmids.
RNA preparation and real-time quantitative RT-PCR
Total RNA was extracted using the RNAqueous Kit (Ambion, Carlsbad, CA, USA) and treated with the DNase I from the DNA-free Kit (Ambion) according to the manufacturer’s instructions. RNA was used to perform a two-step reverse transcription-polymerase chain reaction (RT-PCR) as previously described.25 The sequences of the primers used were as follows: mouse Nfatc3 (forward primer: 5′-TTGAGTGCTCTCAGCGATCT-3′; reverse primer: 5′ GACAATGTGAGCCCCTTGAC 3′), mouse Nfatc4 (forward primer: 5′-GGGAGGAGCTAGTGTTGACT-3′; reverse primer: 5′-TCCGTCCATTGGAGACGTAA-3′), mouse Trim17 (forward primer: 5′-AGGGAGTATAAGCTCAAGTTGGA-3′; reverse primer: 5′-CCTGCCACTCAGTTAAGGTCT-3′) and mouse bdnf (forward primer: 5′-ATCCAAAGGCCAACTGAAGC-3′; reverse primer: 5′-TACGATTGGGTAGTTCGGCA-3′) as described previously.25 Data were analyzed and relative amounts of specifically amplified cDNA were calculated with MxPro software (Agilent, Santa Clara, CA, USA) using the β-2 microglobulin (forward primer: 5′-TATGCTATCCAGAAAACCCCTCAA-3′; reverse primer: 5′-GTATGTTCGGCTTCCCATTCTC-3′) and the hmbs (hydroxymethylbilane synthase; forward primer: 5′-ACTCTGCTTCGCTGCATTG-3′; reverse primer: 5′-AGTTGCCCATCTTTCATCACTG-3′) amplicons as references.
Lentivirus preparation and lentiviral transduction of cells
Lentiviruses were produced as described.50 Cells were transduced one day after plating. The lentiviral preparations were added directly to the culture medium for 8 h (∼500 ng p24 per million neurons, 100 ng p24 per million Neuro2A cells). Cells were then replaced in fresh medium. Culture was continued until 6 DIV for neurons. Neuro2A cells were maintained in culture for 48 h after transduction and then selected using 2 μg/ml puromycin for an additional 48 h.
Assessment of neuronal apoptosis and Trim17 immunoreactivity
After 6 DIV, transfected or transduced CGNs were maintained in initial culture medium (control) or were washed once and incubated in serum-free BME supplemented with L-Gln, HEPES, antibiotics and 1 μM (+)-MK-801, and containing 5 mM KCl (K5 medium) for indicated times. For assessment of neuronal apoptosis, CGNs were deprived of serum and KCl for 8 h and then fixed, stained with DAPI and mounted on glass slides using Mowiol. In experiments in which the CGNs were transfected with GFP, NFATc3-GFP or NFATc4-GFP, apoptosis was assessed among GFP-positive neurons, by examining neuronal morphology and nuclear condensation. For each experiment and each condition, at least 200 GFP-positive neurons were scored in a blinded manner. In experiments in which CGNs were transduced with shRNA-expressing lentiviruses, apoptosis was estimated by counting the percentage of condensed nuclei in five random fields for each condition.
To assess the effect of NFATc3 or NFATc4 overexpression on Trim17 expression, transfected CGNs were deprived of serum and KCl for 5 h, fixed and incubated with purified anti-peptide antibody against Trim17 (1:50) as described previously.25 The intensity of Trim17-associated immunofluorescence was measured specifically in neurons transfected with GFP, NFATc3-GFP or NFATc4-GFP using ImageJ64 software (National Institutes of Health, Bethesda, MD, USA). For each condition, Trim17 immunoreactivity was measured in at least 100 GFP-positive neurons.
Genomic sequence analysis
The genomic sequences of the mouse and human Trim17 genes, from 5000 bp upstream to 5000 bp downstream of the transcription start sites, were retrieved from the DBTSS website. They were aligned using the shuffle-LAGAN algorithm at the VISTA website and searched for NFAT, AP-1 and composite NFAT:AP-1 binding sites using TRANSFAC software by setting the threshold to minimize false negatives. A position frequency matrix was created using a set of 11 experimentally verified NFAT:AP-1 composite binding sites15 and matrices from TRANSFAC were used for individual AP-1 and NFAT binding sites.
Chromatin immunoprecipitation
CGNs were incubated in control or K5 medium for 4 h. The neuronal cultures were fixed by the addition of formaldehyde to a final concentration of 1% and incubated for 10 min at 37 °C, before quenching by the addition of an equal volume of glycine 2.5 M for 5 min at room temperature. The neurons were washed in ice-cold PBS, harvested in PBS complemented with protease inhibitor cocktail (Complete; Roche), pelleted by centrifugation at 300 × g for 5 min at 4 °C and incubated successively in wash buffer no. 1 (10 mM HEPES (pH 7.3), 0.25% Triton X-100, 10 mM EDTA, 0.5 mM EGTA, protease inhibitor cocktail) and in wash buffer no. 2 (10 mM HEPES (pH 7.3), 1 mM EDTA, 0.5 mM EGTA, 200 mM NaCl, protease inhibitor cocktail) for 10 min on ice. The cells were then incubated in lysis buffer (200 μl for 15 × 106 neurons; 50 mM Tris-HCl (pH 7.5), 10 mM EDTA, 1% SDS, protease inhibitor cocktail) overnight at 4 °C. Each 200 μl aliquot of lysate was sonicated at high power for 30 min (30 s ON/30 s OFF) using a Bioruptor ultrasonicator (Diagenode, Seraing, Belgium). Lysates were centrifuged two times at 16 000 × g for 5 min at 4 °C. The DNA content of the resulting supernatants was measured and ∼100 μg chromatin (from ∼8 × 106 cells) was used for each immunoprecipitation in each condition. Samples were diluted 1 : 10 in ChIP dilution buffer (16.7 mM Tris-HCl (pH 8), 1.2 mM EDTA, 1.1% Triton X-100, 0.01% SDS, 167 mM NaCl, protease inhibitor cocktail) and precleared for 2 h at 4 °C with protein A agarose beads previously saturated with 0.01% BSA. Then, samples were rotated overnight at 4 °C in the presence of 2 μg of rabbit IgG (as a negative control) or 2 μg of specific antibodies against NFATc3, c-Jun and H3K4me3. Forty microlitres of protein A agarose beads were added to each sample and rotation was continued for 2 h at 4 °C. The beads were recovered by centrifugation at 1500 × g for 5 min at 4°C and washed successively for 10 min at 4 °C with RIPA buffer (50 mM Tris-HCl (pH 8), 0.1% SDS, 0.5% sodium deoxycholate, 1% NP-40, 150 mM NaCl), high salt buffer (50 mM Tris-HCl (pH 8), 0.1% SDS, 1% NP-40, 500 mM NaCl), LiCl buffer (50 mM Tris-HCl (pH 8), 0.5% sodium deoxycholate, 1% NP-40, 250 mM LiCl) and two times with TE buffer (10 mM Tris-HCl (pH 8), 1 mM EDTA). The immune complexes were eluted by incubating the beads two times with 200 μl elution buffer (2% SDS, 100 mM NaHCO3, 1 mM DTT) for 15 min at room temperature and centrifuging them at 1500 × g for 5 min at 4 °C. The supernatants were collected, pooled, supplemented with 16 μl NaCl 5 M and incubated overnight at 65 °C to reverse protein–DNA crosslinks. An equivalent quantity of input chromatin was treated the same way: ∼100 μg DNA was diluted in 400 μl elution buffer, supplemented with 16 μl NaCl 5 M and incubated overnight at 65 °C. RNA and proteins were removed from the samples by the addition of 8 μl EDTA 0.5 M, 16 μl Tris-HCl (pH 6.5) 1 M, 4 μl proteinase K (10 mg/ml) and 3.3 μl RNAse (24 mg/ml), and incubation for 1 h at 45 °C. ChIP samples and input chromatin were then phenol–chloroform extracted. Purified DNA was precipitated with NaHCO3 and ethanol, washed with 70% ethanol, dissolved in water and subjected to quantitative PCR as described above. To detect binding of NFATc3 and c-Jun to the potential AP-1 and NFAT-AP-1 binding sites, in the mouse Trim17 promoter around the transcription start site, the following primer pair was used: forward primer: 5′-CGTGAGTTGACGCCACAAT-3′; reverse primer: 5′-CGGTGGTTACTCACAAGGAAG-3′. The same primer pair was used to detect H3K4me3, a marker of active chromatin. A standard curve of chromatin was used to quantify the results and the amount of Trim17 promoter DNA precipitated by each antibody was expressed as the percentage of the amount of input chromatin used for each immunoprecipitation.
Statistics
Statistical analyses were performed using GraphPad InStat version 3.0 for Mac (GraphPad Software, San Diego, CA, USA; http: http://www.graphpad.com/).
Accession codes
Abbreviations
- NFAT:
-
nuclear factor of activated T cells
- AP-1:
-
activator protein 1
- SUMO:
-
small ubiquitin-like modifier
- NEM:
-
N-ethylmaleimide
- GFP:
-
green fluorescent protein
- HA:
-
hemagglutinin tag
- SIM:
-
SUMO-interacting motif
- WT:
-
wild type
- BHK:
-
baby hamster kidney
- PMA:
-
phorbol 12-myristate 13-acetate
- BDNF:
-
brain-derived neurotrophic factor
- shRNA:
-
short hairpin RNA
- DIV:
-
days in vitro
- CGN:
-
cerebellar granule neurons
- K5:
-
serum-free medium containing 5 mM KCl
- PI3K:
-
phosphatidyl inositol-3-OH kinase
- JNK:
-
c-Jun N-terminal kinase
- GSK3:
-
glycogen synthase kinase 3
- DRG:
-
dorsal root ganglion
- ChIP:
-
chromatin immunoprecipitation
References
Hyman BT, Yuan J . Apoptotic and non-apoptotic roles of caspases in neuronal physiology and pathophysiology. Nat Rev Neurosci 2012; 13: 395–406.
Yuan J, Yankner BA . Apoptosis in the nervous system. Nature 2000; 407: 802–809.
Ham J, Eilers A, Whitfield J, Neame SJ, Shah B . c-Jun and the transcriptional control of neuronal apoptosis. Biochem Pharmacol 2000; 60: 1015–1021.
D'Mello SR, Galli C, Ciotti T, Calissano P . Induction of apoptosis in cerebellar granule neurons by low potassium: inhibition of death by insulin-like growth factor I and cAMP. Proc Natl Acad Sci USA 1993; 90: 10989–10993.
Martin DP, Schmidt RE, DiStefano PS, Lowry OH, Carter JG, Johnson EMJ . Inhibitors of protein synthesis and RNA synthesis prevent neuronal death caused by nerve growth factor deprivation. J Cell Biol 1988; 106: 829–844.
Schulz JB, Weller M, Klockgether T . Potassium deprivation-induced apoptosis of cerebellar granule neurons: a sequential requirement for new mRNA and protein synthesis, ICE-like protease activity, and reactive oxygen species. J Neurosci 1996; 16: 4696–4706.
Nguyen T, Di Giovanni S . NFAT signaling in neural development and axon growth. Int J Dev Neurosci 2008; 26: 141–145.
Graef IA, Wang F, Charron F, Chen L, Neilson J, Tessier-Lavigne M et al. Neurotrophins and netrins require calcineurin/NFAT signaling to stimulate outgrowth of embryonic axons. Cell 2003; 113: 657–670.
Benedito AB, Lehtinen M, Massol R, Lopes UG, Kirchhausen T, Rao A et al. The transcription factor NFAT3 mediates neuronal survival. J Biol Chem 2005; 280: 2818–2825.
Vashishta A, Habas A, Pruunsild P, Zheng JJ, Timmusk T, Hetman M . Nuclear factor of activated T-cells isoform c4 (NFATc4/NFAT3) as a mediator of antiapoptotic transcription in NMDA receptor-stimulated cortical neurons. J Neurosci 2009; 29: 15331–15340.
Quadrato G, Benevento M, Alber S, Jacob C, Floriddia EM, Nguyen T et al. Nuclear factor of activated T cells (NFATc4) is required for BDNF-dependent survival of adult-born neurons and spatial memory formation in the hippocampus. Proc Natl Acad Sci USA 2012; 109: E1499–E1508.
Gomez-Sintes R, Lucas JJ . NFAT/Fas signaling mediates the neuronal apoptosis and motor side effects of GSK-3 inhibition in a mouse model of lithium therapy. J Clin Invest 2010; 120: 2432–2445.
Luoma JI, Zirpel L . Deafferentation-induced activation of NFAT (nuclear factor of activated T-cells) in cochlear nucleus neurons during a developmental critical period: a role for NFATc4-dependent apoptosis in the CNS. J Neurosci 2008; 28: 3159–3169.
Jayanthi S, Deng X, Ladenheim B, McCoy MT, Cluster A, Cai NS et al. Calcineurin/NFAT-induced up-regulation of the Fas ligand/Fas death pathway is involved in methamphetamine-induced neuronal apoptosis. Proc Natl Acad Sci USA 2005; 102: 868–873.
Hogan PG, Chen L, Nardone J, Rao A . Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev 2003; 17: 2205–2232.
Macian F . NFAT proteins: key regulators of T-cell development and function. Nat Rev Immunol 2005; 5: 472–484.
Muller MR, Rao A . NFAT, immunity and cancer: a transcription factor comes of age. Nat Rev Immunol 2010; 10: 645–656.
Wu H, Peisley A, Graef IA, Crabtree GR . NFAT signaling and the invention of vertebrates. Trends Cell Biol 2007; 17: 251–260.
Kim MS, Usachev YM . Mitochondrial Ca2+ cycling facilitates activation of the transcription factor NFAT in sensory neurons. J Neurosci 2009; 29: 12101–12114.
Glud SZ, Sorensen AB, Andrulis M, Wang B, Kondo E, Jessen R et al. A tumor-suppressor function for NFATc3 in T-cell lymphomagenesis by murine leukemia virus. Blood 2005; 106: 3546–3552.
Cante-Barrett K, Winslow MM, Crabtree GR . Selective role of NFATc3 in positive selection of thymocytes. J Immunol 2007; 179: 103–110.
Oukka M, Ho IC, de la Brousse FC, Hoey T, Grusby MJ, Glimcher LH . The transcription factor NFAT4 is involved in the generation and survival of T cells. Immunity 1998; 9: 295–304.
Nayak A, Glockner-Pagel J, Vaeth M, Schumann JE, Buttmann M, Bopp T et al. Sumoylation of the transcription factor NFATc1 leads to its subnuclear relocalization and interleukin-2 repression by histone deacetylase. J Biol Chem 2009; 284: 10935–10946.
Terui Y, Saad N, Jia S, McKeon F, Yuan J . Dual role of sumoylation in the nuclear localization and transcriptional activation of NFAT1. J Biol Chem 2004; 279: 28257–28265.
Lassot I, Robbins I, Kristiansen M, Rahmeh R, Jaudon F, Magiera MM et al. Trim17, a novel E3 ubiquitin-ligase, initiates neuronal apoptosis. Cell Death Differ 2010; 17: 1928–1941.
Gareau JR, Lima CD . The SUMO pathway: emerging mechanisms that shape specificity, conjugation and recognition. Nat Rev Mol Cell Biol 2010; 11: 861–871.
Kerscher O . SUMO junction-what's your function? New insights through SUMO-interacting motifs. EMBO Rep 2007; 8: 550–555.
Hecker CM, Rabiller M, Haglund K, Bayer P, Dikic I . Specification of SUMO1- and SUMO2-interacting motifs. J Biol Chem 2006; 281: 16117–16127.
Magiera MM, Mora S, Mojsa B, Robbins I, Lassot I, Desagher S . Trim17-mediated ubiquitination and degradation of Mcl-1 initiate apoptosis in neurons. Cell Death Differ 2013; 20: 281–292.
Urano T, Usui T, Takeda S, Ikeda K, Okada A, Ishida Y et al. TRIM44 interacts with and stabilizes terf, a TRIM ubiquitin E3 ligase. Biochem Biophys Res Commun 2009; 383: 263–268.
Contestabile A . Cerebellar granule cells as a model to study mechanisms of neuronal apoptosis or survival in vivo and in vitro. Cerebellum 2002; 1: 41–55.
Ikonomidou C, Bosch F, Miksa M, Bittigau P, Vöckler J, Dikranian K et al. Blockade of NMDA receptors and apoptotic neurodegeneration in the developing brain. Science 1999; 283: 70–74.
Wood KA, Dipasquale B, Youle RJ . In situ labelling of granule cells for apoptosis-associated DNA fragmentation reveals different mechanisms of cell loss in developing cerebellum. Neuron 1993; 11: 621–632.
Desagher S, Severac D, Lipkin A, Bernis C, Ritchie W, Le Digarcher A et al. Genes regulated in neurons undergoing transcription-dependent apoptosis belong to signaling pathways rather than the apoptotic machinery. J Biol Chem 2005; 280: 5693–5702.
Chen L, Glover JN, Hogan PG, Rao A, Harrison SC . Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature 1998; 392: 42–48.
Watson A, Eilers A, Lallemand D, Kyriakis J, Rubin LL, Ham J . Phosphorylation of c-Jun is necessary for apoptosis induced by survival signal withdrawal in cerebellar granule neurons. J Neurosci 1998; 18: 751–762.
Meroni G, Diez-Roux G . TRIM/RBCC, a novel class of 'single protein RING finger' E3 ubiquitin ligases. Bioessays 2005; 27: 1147–1157.
Napolitano LM, Meroni G . TRIM family: Pleiotropy and diversification through homomultimer and heteromultimer formation. IUBMB Life 2012; 64: 64–71.
Harada J, Sugimoto M . An inhibitor of p38 and JNK MAP kinases prevents activation of caspase and apoptosis of cultured cerebellar granule neurons. Jpn J Pharmacol 1999; 79: 369–378.
Chin PC, Majdzadeh N, D'Mello SR . Inhibition of GSK3beta is a common event in neuroprotection by different survival factors. Brain Res Mol Brain Res 2005; 137: 193–201.
Morrison BE, Majdzadeh N, Zhang X, Lyles A, Bassel-Duby R, Olson EN et al. Neuroprotection by histone deacetylase-related protein. Mol Cell Biol 2006; 26: 3550–3564.
Ulrich JD, Kim MS, Houlihan PR, Shutov LP, Mohapatra DP, Strack S et al. Distinct activation properties of the nuclear factor of activated T-cells (NFAT) isoforms NFATc3 and NFATc4 in neurons. J Biol Chem 2012; 287: 37594–37609.
Hongisto V, Smeds N, Brecht S, Herdegen T, Courtney MJ, Coffey ET . Lithium blocks the c-Jun stress response and protects neurons via its action on glycogen synthase kinase 3. Mol Cell Biol 2003; 23: 6027–6036.
Reddy CE, Albanito L, De Marco P, Aiello D, Maggiolini M, Napoli A et al. Multisite phosphorylation of c-Jun at threonine 91/93/95 triggers the onset of c-Jun pro-apoptotic activity in cerebellar granule neurons. Cell Death Dis 2013; 4: e852.
Vojtek AB, Hollenberg SM . Ras–Raf interaction: two-hybrid analysis. Methods Enzymol 1995; 255: 331–342.
Bartel PL, Chien C-T, Sternglanz R, Fields S . Using the two-hybrid system to detect protein-protein interactions In: Hartley DA (ed). Cellular Interactions in Development: A Practical Approach. Oxford University Press: Oxford, UK, 1993 pp 153–179.
Fromont-Racine M, Rain JC, Legrain P . Toward a functional analysis of the yeast genome through exhaustive two-hybrid screens. Nat Genet 1997; 16: 277–282.
Formstecher E, Aresta S, Collura V, Hamburger A, Meil A, Trehin A et al. Protein interaction mapping: a Drosophila case study. Genome Res 2005; 15: 376–384.
Bossis G, Chmielarska K, Gartner U, Pichler A, Stieger E, Melchior F . A fluorescence resonance energy transfer-based assay to study SUMO modification in solution. Methods Enzymol 2005; 398: 20–32.
Swainson L, Mongellaz C, Adjali O, Vicente R, Taylor N . Lentiviral transduction of immune cells. Methods Mol Biol 2008; 415: 301–320.
Acknowledgements
We thank the staff of the Animal facilities of the IGMM for the breeding of mice and the Montpellier RIO imaging platform and their financing bodies. We are grateful to Dr Guillaume Bossis for providing recombinant GST-SENP1, His6-ubiquitin plasmid and all in vitro SUMOylation reagents. We also thank Dr Michal Hetman for plasmids encoding NFATc3 and NFATc4. We are also grateful to Drs Jonathan Ham and Ian Robbins for critical reading of the manuscript and for help in promoter sequence analysis. We thank Drs Olivier Coux, Philippe Marin and Jacques Ghysdael for interesting discussions. This work was supported by the Centre National de la Recherche Scientifique (CNRS), the Institut National de la Santé et de la Recherche Médicale (INSERM), Université Montpellier 2, la Fondation de l’Association pour la Recherche contre le Cancer (ARC) and La Ligue contre le Cancer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by C Borner
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Rights and permissions
About this article
Cite this article
Mojsa, B., Mora, S., Bossowski, J. et al. Control of neuronal apoptosis by reciprocal regulation of NFATc3 and Trim17. Cell Death Differ 22, 274–286 (2015). https://doi.org/10.1038/cdd.2014.141
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2014.141
This article is cited by
-
The E3 ubiquitin ligase TRIM17 promotes gastric cancer survival and progression via controlling BAX stability and antagonizing apoptosis
Cell Death & Differentiation (2023)
-
Tripartite Motif Protein Family in Central Nervous System Diseases
Cellular and Molecular Neurobiology (2023)
-
Arsenic sulfide induces RAG1-dependent DNA damage for cell killing by inhibiting NFATc3 in gastric cancer cells
Journal of Experimental & Clinical Cancer Research (2019)
-
TRIM17 and TRIM28 antagonistically regulate the ubiquitination and anti-apoptotic activity of BCL2A1
Cell Death & Differentiation (2019)
-
A miRNA-101-3p/Bim axis as a determinant of serum deprivation-induced endothelial cell apoptosis
Cell Death & Disease (2017)