Abstract
Little is known about the cellular mechanisms modulating the shift in balance from a state of survival to cell death by caspase-mediated apoptosis in response to a lethal stress. Here we show that the RNA-binding protein HuR has an important function in mediating this switch. During caspase-mediated apoptosis, HuR is cleaved to generate two cleavage products (CPs). Our data demonstrate that the cleavage of HuR switches its function from being a prosurvival factor under normal conditions to becoming a promoter of apoptosis in response to a lethal stress. In the absence of an apoptotic stimuli, HuR associates with and promotes the expression of caspase-9 and prothymosin α (ProT) mRNAs, and pro- and antiapoptotic factors, respectively, both of which have been characterized as important players in determining cell fate. During the early steps of caspase-mediated apoptosis, however, the level of caspase-9 protein increases, while ProT remains unchanged. Under these conditions, the two HuR-CPs selectively bind to and stabilize caspase-9 mRNA, but do not bind to ProT. Hence, taken together, our data show that by maintaining a threshold of expression of proapoptotic factors such as caspase-9 in response to a lethal stress, the HuR-CPs help a cell to switch from resisting death to undergoing apoptosis.
Similar content being viewed by others
Main
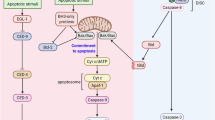
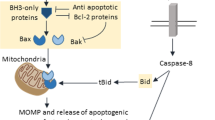
Caspase-mediated apoptotic cell death depends on the balance between numerous pro- and antiapoptotic regulators.1, 2 During stress response, this balance is tilted, initially to activate prosurvival mechanisms, but if the stress becomes unsustainable, apoptosis is engaged.3 This shift of equilibrium depends on multiple factors, including members of the Bcl-2 family, which can either promote apoptosis (e.g. Bad, Bax), by mediating the release of cytochrome c (cyt c) from the mitochondria to the cytoplasm, or inhibit it, by preventing this release (Bcl-2, Bcl-xL).4, 5 The cytoplasmic localization of cyt c enables the formation of an active apoptosome, a complex bringing together Apaf-1 protein and caspase-9.4, 6 Once active, the apoptosome triggers the activation of caspase-9, allowing it to cleave and activate downstream caspases (such as -3 and -7), leading to cell death.4 Not surprisingly, caspase-9 has been characterized as an important regulator of caspase-mediated apoptosis.7, 8, 9 The activity of the apoptosome may be either increased by activators such as pp32/PHAPI or decreased by inhibitors such as prothymosin α (ProT).10 In cancer development, the activities of apoptotic proteins are defective, and a decrease or increase in the respective expression levels of pro- and antiapoptotic factors is also observed.4, 11
It is well documented that the expression of various apoptotic players is regulated at the level of transcription.4, 12, 13, 14 More recent evidence suggests that apoptotic genes are also regulated post-transcriptionally.15, 16 One way by which this occurs is via the interaction of AU-rich elements (AREs) located in the 3′ untranslated region (3′-UTR) of pro- and antiapoptotic mRNAs with ARE-binding proteins. HuR is one such protein that has an important part in stress response.15, 17, 18 Typically, ARE-containing mRNAs are quite labile, as they undergo ARE-mediated decay (AMD).19 Although many ARE-binding proteins destabilize these mRNAs, HuR is best known to increase their half-lives and/or modulate their translation.15, 20 Curiously, it has been shown that UV stress causes HuR to stimulate the translation of both pro- (p53 and cyt c) and anti- (ProT) apoptotic factors.21, 22, 23 However, no explanation has been provided to address how one stress can cause HuR to promote two opposing effects.
We recently showed that during cell response to lethal stress, HuR is exported to the cytoplasm while associated with the apoptosome activator pp32/PHAPI, and is then cleaved in a PKR-dependent pathway by caspases-3 and -7, yielding two cleavage products (HuR-CP1 and HuR-CP2).18, 24 Physiological significance of this cleavage, which occurs for approximately 50% of cytoplasmic HuR, was shown in the process of muscle development.25 The significance of HuR cleavage in apoptosis was demonstrated by the fact that a non-cleavable mutant (HuRD226A) of HuR failed to rescue apoptosis in cells depleted of endogenous HuR.18 In addition, the expression of these HuR-CPs can stimulate apoptosis in the presence of a non-lethal stress (conditions under which apoptosis is not engaged and HuR cleavage does not occur).24 These results highlight a function for these CPs in triggering cell death, although the mechanisms responsible for this effect are unclear.
As HuR-CPs harbor the RRMs of HuR (HuR-CP1, RRM1-2 and HuR-CP2, RRM3),18 in this study we investigated the possibility that they can promote apoptosis by modulating the expression of proapoptotic mRNAs. Here we show that the two HuR-CPs associate and positively affect the expression of caspase-9, but not that of ProT. Our data support a model whereby the cleavage of HuR is a trigger for its role reversal, which explains how and why during cell response to severe stress HuR switches from promoting survival to assisting with apoptotic cell death.
Results
HuR promotes apoptosis by regulating caspase-9 expression in an ARE-dependent manner
As apoptosome activation involves cyt c release, to further establish how HuR influences apoptosis, we asked if the proapoptotic function of HuR occurs downstream of this event. We observed that by knocking down HuR using siRNA (siRNA-HuR) in HeLa cells (Figure 1A), Bax- (a well-established regulator of cyt c release26) induced cell death was prevented (Figures 1B and C). HuR expression was rescued in this system by providing cells with HuR conjugated to the cell-permeable peptide AP (antennepedia) (AP-HuR),18 and this demonstrated a simultaneous rescue of cell death (image c), which did not occur with AP-GST control (image d). These results suggest that HuR promotes apoptosis by acting downstream of Bax, and possibly cyt c release.
HuR is involved in apoptosis downstream of Bax and binds to caspase-9 mRNA. (A–C) HuR is needed for Bax-induced apoptosis. (A) HeLa cells were transfected with siRNA against HuR, or Control (C), or mock transfected, and 24 h later were transfected with HA-Bax plasmid. Lysates of these cells were used for western blot, probing with anti-HuR and anti-Bax antibodies. Shown are representative images from two independent experiments. (B and C) HeLa cells transfected as in (A) were given a 50 nM dose of AP-GST (D) or AP-HuR-GST (C) for 16 h before being photographed. The percentage of dead cells was assessed by subtracting from 100% the dividend of the number of adherent cells following HA-Bax transfection over the total number of cells present in a non-transfected sample. Shown are representative panels of cells from five different fields of view, each from two independent experiments, with the mean values±the standard error of the mean (S.E.M.). These values are graphed in (C). (D) HuR associates with caspase-9 mRNA. Total mRNA from HeLa cells was incubated with anti-HuR antibody (3A2), anti-Sam68 antibody or IgG control, and the IPed mRNA was used for RT-PCR. Shown is a representative image of three independent experiments. (E) HuR associates with caspase-9 mRNA by binding its AREs. (Top panel) Cartoon representing full-length caspase-9, both the coding region (CDR) and UTRs, and indicating the location of the two AREs. (Bottom panel) Radiolabeled RNA probes representing ARE1 and ARE2 (see cartoon) were incubated with 500 ng of recombinant GST-HuR or GST-alone proteins. Samples were then resolved by a non-denaturing gel. Asterisks denote complexes, and shown is a representative autoradiograph of five independent experiments. (F–I) Knockdown of HuR decreases caspase-9 mRNA and protein levels. HeLa cells were treated with siRNA against HuR or control siRNA for 48h, and their total mRNA (F and G) was then collected and analyzed by northern blotting. Probes against HuR, caspase-9, ProT and 18S (loading control) mRNA were used. In parallel, cells that were transfected as described were provided with 50 nM AP-GST or AP-HuR-GST 16 h before being collected, lysed and analyzed by western blot (H, I), using antibodies against caspase-9, HuR and G3BP (loading control). Quantification of band intensities was performed using the ImageQuant software. Asterisks indicate significant differences in mean values, with P-values stated, and not significant differences are also indicated (NS). Shown are representative blots of at least three independent experiments
To address how HuR, by acting at this level, can shift from promoting survival to activating cell death, we performed RIP-CHIP (RNP-Immunoprecipitation-microchip)27, 28, 29 experiments to identify the HuR mRNA targets that could affect apoptosome activity. To do this, an immunoprecipitation (IP) was carried out on HeLa cells using anti-HuR antibody (3A2),30 and the mRNA associating with HuR were hybridized on an apoptosis-specific cDNA microarray. Numerous messages were identified (Supplementary Figure 1), but of these, caspase-9 stood out as an mRNA encoding for a component of the apoptotic response that acts downstream of Bax.12 We validated this result by an IP/RT-PCR experiment, where it was confirmed that HuR associates with caspase-9 mRNA. As a positive control, we observed that HuR also binds to its mRNA target ProT22 (Figure 1D), which was not included in the array that was used. The mRNA of heterogenous nuclear ribonucleoprotein A1 (hnRNP A1), a negative control, did not associate with HuR.31 By scanning the 3′-UTR of caspase-9 mRNA, we identified two AREs (ARE1: 1841–1870; ARE2: 1944–1988) (Supplementary Figure 2). Gel-shift experiments using radioactive-labeled probes showed that both AREs associate with GST-HuR but not GST alone (Figure 1E). In addition, knockdown and rescue experiments (Figures 1F–I and Supplementary Figure 3) confirmed that HuR is required for the expression of both caspase-9 mRNA and protein.
Next, we determined the importance of these AREs in regulating the expression of caspase-9. To do so, we acquired murine embryonic fibroblasts (MEFs) isolated from caspase-9−/− mice,6 in which we expressed full-length caspase-9 mRNA with and without functional AREs. Overexpressing HA-Bax in these cells promoted the cleavage of caspase-3 and poly (ADP-ribose) polymerase (PARP), well-known indicators of caspase-mediated apoptosis activation,12 in wild-type (wt) but not in caspase-9−/− MEFs (Figures 2a and b). To assess the interplay between HuR, Bax and caspase-9, we depleted HuR expression in wtMEFs and caspase-9−/− MEFs while overexpressing Bax (Figures 2c and d), and observed that both HuR and caspase-9 are required for Bax-induced apoptosis. These observations clearly indicate that Bax triggers caspase-dependent cell death via HuR, which acts upstream of caspase-9.
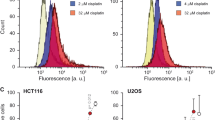
Cells lacking caspase-9 are resistant to caspase-mediated apoptosis. (a, b) Bax-induced caspase-dependent apoptosis is reduced in the absence of caspase-9. wtMEFs and caspase-9−/− MEFs were transfected twice with HA-tagged Bax plasmid or empty pcDNA3 plasmid, and were lysed to be analyzed by western blot. Western blots (a) were probed with antibodies against HA tag, caspase-3 CP (casp-3 CP), PARP CP (PARP CP) and α-tubulin (loading control). The asterisk indicates a nonspecific band seen using the caspase-3 antibody. A representative blot of three independent experiments is shown. Quantification of caspase-3 and PARP CPs was performed using the ImageQuant software and relative average amounts are graphed in (b), with error bars representing the S.E.M. of three independent experiments, and the asterisk indicating that the observed difference is significant. (c, d) The induction of caspase-dependent apoptosis by Bax overexpression involves HuR and caspase-9. wtMEFs and caspase-9−/− MEFs were transfected with siRNA against HuR or control siRNA, as well as with HA-Bax. Cells were lysed and analyzed by western blot, a representative sample of which is shown in (c), using antibodies against caspase-9 (casp-9), caspase-3 CP (asterisk indicates a nonspecific band), PARP CP, HuR, HA-tag and α-tubulin (loading control). Quantified levels of cleaved PARP and caspase-3 were determined as described above (b), and average values, relative to loading control, are graphed in (d), with error bars representing the S.E.M. of two independent experiments. (e–h) Caspase-9-deficient cells are resistant to STS-induced apoptosis. wtMEFs and caspase-9−/− MEFs were treated with 1 μM STS for 6 h (e–h). Lysates from these cells were analyzed by western blot (e) and probed with antibodies against PARP CP, caspase-3 CP, caspase-9, HuR and α-tubulin (loading control). Quantified levels of cleaved PARP and caspase-3 were determined as described above (b), and average values, relative to loading control, are graphed in (f), with error bars representing the S.E.M. of three independent experiments. The asterisks indicate that the observed differences in the means are significant. Alternatively, subsequent to these treatments, cells were stained with Annexin V-FITC and analyzed by flow cytometry (g, h). The relative fluorescent intensity, measured in the FL-1 channel, is graphed against the number of counts (g) and average relative values from three independent experiments are graphed in (h), with error bars representing the S.E.M. and the asterisk indicating that the observed difference in the means is significant
We then evaluated the ability of caspase-9-deficient cells to undergo apoptosis in response to the apoptosis-inducing agent staurosporine (STS).24 Caspase-9−/− and wtMEFs were treated with 1 μM STS for 6h.24 Not surprisingly, we observed a significant reduction in STS-induced apoptosis in caspase-9−/− MEFs when compared with their wt counterparts (Figures 2e–h). Collectively, the data in Figure 2 confirm that caspase-9 has an important function in Bax-mediated caspase-dependent apoptosis.
To assess the importance of AREs in caspase-9 expression, we generated plasmids expressing either full-length caspase-9 or full-length caspase-9 in which the AREs were mutated (replacing the Us by Cs) (Figure 3a). Transfecting these plasmids into caspase-9−/− MEFs showed that mutating the AREs caused >80% decrease in the expression of caspase-9 mRNA (Figures 3b and c). Using these cells in an actinomycin D (ActD) pulse-chase experiment,32 we observed that mutated caspase-9 mRNA had a reduced half-life (of ∼4.1 h) compared with full-length caspase-9 mRNA (∼6.6 h; Figures 3d and e and Supplementary Figure 4). Although AREs are generally known to destabilize their host messages, our data clearly suggest that in the case of caspase-9 mRNA, they are in fact required for its stabilization. Our results also show that this ARE-mediated stabilization of caspase-9 mRNA is required for the expression of its protein under normal conditions (Figures 3f and g). We then investigated the effect of ARE mutations on the ability for caspase-9 to rescue apoptotic cell death in caspase-9−/− MEFs. We observed that caspase-9 lacking functional AREs was less effective in rescuing apoptosis compared with the transfection of full-length caspase-9 (Figures 3h–k).
The AREs of caspase-9 mRNA regulate its stability. (a) Caspase-9 mutated AREs. Cartoon illustrating the caspase-9 plasmids produced, highlighting the mutations made in the AREs. Asterisks indicate nucleotides that were mutated. (b, c) Mutating caspase-9 AREs decreases RNA levels. Caspase-9−/− MEFs were transfected with GFP and caspase-9 mutants (full-length, or with mutated AREs). At 24 h following this transfection, total RNA was isolated and analyzed by northern blot, using radiolabeled probes against GFP (transfection control), caspase-9 and 18S (loading control). Caspase-9 and 18S levels were quantified using the ImageQuant software, and average caspase-9 values were graphed, with error bars representing the S.E.M. of four independent experiments. The asterisk indicates that the observed difference in means is significant. (d, e) Mutating caspase-9 AREs decreases mRNA stability. Caspase-9−/− MEFs were transfected as in (b), and 24 h after transfection, cells were treated with 5 μg/ml Act D for the indicated periods of time. Total RNA was collected at each of these time points and analyzed by northern blot, using radiolabeled probes against GFP, caspase-9 and 18S (loading control). Quantification using the ImageQuant software was performed, determining relative caspase-9 levels compared with 18S. Stability curves were generated using Microsoft Excel’s exponential trendline feature, from which half-lives could be determined for each experiment. The average half-life of each mRNA is shown with S.E.M. from four independent experiments, and the asterisk indicates that the difference in half-lives is significant. The average % remaining RNA values for each time point have also been graphed, with error bars representing the S.E.M. of these four experiments. (f, g) Mutating caspase-9 AREs decreases protein levels. Cells transfected as in (b) were lysed and analyzed by western blot, using antibodies against caspase-9, GFP (transfection control) and α-tubulin (loading control). Quantification was carried out as in (b), graphing average values, with error bars representing the S.E.M. of four independent experiments. The asterisk indicates that the difference in the means is significant. (h–k) Mutating AREs of caspase-9 inhibits apoptosis. Full-length caspase-9, or caspase-9 with mutated AREs, was transfected into caspase-9-deficient MEFs, and 24 h later, cells were treated or not with 1 μM STS for 6 h. Total protein extracts from cells were obtained and were analyzed by western blot (h, i) with antibodies against caspase-9, PARP CP, caspase-3 CP, GFP and α-tubulin (loading control). Quantified levels of cleaved PARP and caspase-3 were determined using the ImageQuant software, and average values are graphed in (i), with error bars representing the S.E.M. of four independent experiments. The asterisks indicate that the observed differences in the means are significant. Alternatively, following STS treatment, cells were stained with Annexin V-FITC and analyzed by flow cytometry (j, k). The FL-1 channel measured the relative fluorescent intensity of FITC, and is graphed against the number of counts (j). Average relative values from two independent experiments are graphed in (k), with error bars representing the S.E.M. All blots shown (b, d, f and h) are representative of the indicated number of experiments
Taken together, these results support the idea that by associating with HuR, the AREs of caspase-9 mRNA may protect the message from AMD and that this interaction has an important part in promoting apoptosis.
STS treatment causes an increase in caspase-9 protein levels
It has been shown that under non-lethal conditions, HuR supports survival by promoting the translation of ProT.22 This, with the results described above, raises the possibility that under normal conditions HuR promotes survival, but in response to severe stress, executes functions on mRNA targets to advance apoptosis. If this is the case, under lethal conditions HuR should favor the expression of caspase-9 over ProT. We found that while a substantial decrease in the levels of caspase-9 and ProT mRNAs was observed only after 2.5 h of STS treatment, the combined protein levels of caspase-9 and its p35- and p37-active CPs increased by >50% at 3 h after STS treatment. The caspase-9 protein level was not further enhanced by co-treating cells with MG132, an established chemical inhibitor of the proteosome (Figures 4a–d and Supplementary Figure 5), indicating that the mechanism responsible for this increase in caspase-9 expression does not involve increasing its protein stability. As several technical challenges make it difficult to detect endogenous ProT by western blot,22 a plasmid encoding full-length ProT mRNA (GFP-ProT) was transfected into HeLa cells, which were treated with STS as in Figure 4a (Figures 4e and f). Unlike caspase-9, no substantial change in the protein level of ProT was seen. Collectively these observations suggest that during apoptosis a specific post-transcriptional regulatory mechanism (mRNA stability or translation) ensures the high expression levels of caspase-9, but not ProT, during apoptosis. Polysome fractionation experiments (Figures 4g–i) showed that STS treatment does not affect the rate of recruitment of endogenous caspase-9 and ProT mRNAs to heavy polysomes, suggesting that it is not at the level of translation that these messages are differentially regulated.
STS treatment causes an increase in protein levels of caspase-9, but not of ProT. (a, b) The mRNA levels of ProT and caspase-9 decrease during 3 h of STS treatment. Total mRNA was isolated from HeLa cells treated for 0, 0.5, 1, 1.5, 2, 2.5 or 3 h STS, which were then analyzed by northern blot, probing for caspase-9, ProT and 18S (loading control). (a) Shown is a representative blot from four independent experiments. (b) The signal from the total levels of mRNA shown in (a) were quantified using the ImageQuant software. Average values of RNA, relative to 18S, were graphed, with error bars representing the S.E.M. of four independent experiments. (c, d) Treatment with STS, but not with the proteasome inhibitor MG132, enhances the expression of caspase-9 protein. HeLa cells were treated with MG132, a well-established proteosome inhibitor, 1 μM STS (as described in a), or a combination of both for 0, 0.5, 1, 1.5, 2, 2.5 or 3 h, after which protein extracts were prepared and analyzed by western blot (c). Antibodies against caspase-9, HuR and G3BP (loading control) were used and representative blots from three independent experiments are shown. Caspase-9 protein levels were quantified and normalized against G3BP, and average values are graphed in (d), with error bars representing the S.E.M. of three independent experiments. The asterisks indicate that the difference between these values and their corresponding 0 h sample is significant. (e, f) The expression level of ProT protein does not change during STS treatment. HeLa cells were transfected with a plasmid encoding GFP-tagged ProT, and 48 h later were treated as in (a), and western blots using the lysates of these cells were probed with GFP and G3BP (loading control; e). Band intensities were quantified and relative averages graphed (f), with error bars again representing the S.E.M. of three independent experiments. (g–i) STS treatment has no effect on the recruitment to polysomes of caspase-9 and ProT mRNAs. HeLa cells were treated with STS for 3 h, or not, and their total RNA was collected and fractionated on a sucrose gradient to generate fractions containing different numbers of ribosomes (polysomes). The RNA from each sample was loaded onto a slot blot, which was incubated with probes for casp-9, ProT and 18S (loading control). The profile of these fractions is shown in (g), and a representative slot blot from two independent experiments is shown in (h) and quantified in (i). Quantification was carried out by dividing the quantity of mRNA (casp-9 or ProT) in the fractions representing polysomes by the total ProT mRNA present in all fractions, using the ImageQuant software, and mean values are graphed, with error bars representing the S.E.M.
HuR CPs preferentially bind caspase-9 mRNA via AREs in the 3′-UTR
The data described above raise the possibility that HuR could be responsible for the preferential expression of caspase-9 over ProT during cell response to a lethal stress such as STS. To address this, we first assessed whether the differential effects on caspase-9 and ProT expression could be due to changes in the association of these messages with HuR. Surprisingly, an IP experiment showed that HuR continued to associate with ProT message during STS treatment (0–3 h), although association with caspase-9 mRNA decreased (Figures 5a–c). Given that STS induces the cleavage of HuR to generate HuR-CP1 and HuR-CP2 after 1.5–2 h of treatment18 and that STS significantly increases caspase-9 expression (Figure 4c), we hypothesized that this differential effect may be due to the presence of these CPs, and not full-length HuR. To assess this, an IP/RT-quantitative PCR (qPCR) experiment was performed on cells expressing GFP-HuR, or GFP-HuR-CPs. Intriguingly, while GFP-HuR associated as expected with both caspase-9 and ProT mRNAs, the CPs preferentially associated with caspase-9 mRNA over ProT (Figures 5d and e). A northern blot analysis indicated that this differential binding is not due to an effect of the HuR-CPs on the expression levels of caspase-9 and ProT mRNAs (Supplementary Figure 6). This difference in binding of the HuR-CPs with ProT mRNA compared with wtHuR may explain in part why, in response to STS treatment, although caspase-9 expression was increased, no effect on ProT was observed.
The CPs of HuR associate with caspase-9 mRNA. (a–c) The association of HuR with caspase-9 mRNA, but not with ProT mRNA, decreases following STS treatment. HeLa cells were treated with 1 μM STS for 0, 1.5 or 3 h, and lysates from these samples were incubated with IgG control, or anti-HuR (3 A2) antibodies. Immunoprecipitated mRNA was isolated and used for RT-PCR or RT-qPCR for each time point, performing PCRs for caspase-9 and ProT messages. Shown is a representative agarose gel/RT-PCR of three independent experiments (a), or average values following two qPCR experiments (b, c), where caspase-9 and ProT message levels were normalized against IgG signals, and are relative to the corresponding IgG sample at each time point. (d, e) HuR-CPs associate with caspase-9 mRNA, but not with that of ProT. HeLa cells transfected with GFP plasmid alone, or fused with HuR, HuR-CP1 or HuR-CP2 were lysed and incubated with anti-GFP antibody. mRNA was isolated from the immunoprecipitate, and RT-PCR and qPCR was performed as in (a–c). Shown is a representative agarose gel from three independent experiments, with an interfering lane removed by cropping (d), and average values of caspase-9 mRNA (e) from three independent experiments found to associate with GFP-HuR, GFP-HuR-CP1 and GFP-HuR-CP2, relative to GFP. (f, g) HuR-CPs associate with the AREs of caspase-9 mRNA. Gel-shift experiments were performed as described in Figure 1d, using recombinant GST, GST-HuR, GST-HuR-CP1 or GST-HuR-CP2 proteins, and radiolabeled RNA probes representing ARE1, ARE2 or mutants of these AREs where all uracil ribonucleotides were replaced with cytosines. Asterisks denote complexes, and shown are representative autoradiographs of four independent experiments. (h, i) HuR-CP1 binds ARE2 with a higher affinity than HuR. Gel-shift experiments were performed as described above using varying concentrations of GST-HuR or GST-HuR-CP1, and 10 000 c.p.m. of radiolabeled ARE2 probe. Quantification of bound and unbound signal allowed dissociation constants to be determined (Kd) for both GST-HuR and GST-HuR-CP1, using Microsoft Excel’s trendline feature on a graph of bound/unbound signal against concentration of protein. The average Kd values of three independent experiments, ±the S.E.M. of the experiments, are presented in (i), and a representative blot of these experiments in shown in (h)
To further characterize how these CPs regulate caspase-9 mRNA, we repeated our gel-shift experiment from Figure 1e in the presence of each of the two CPs, and saw that both are capable of associating with the AREs of the caspase-9 mRNA 3′-UTR (Figures 5f and g). Mutation of the U nucleotides in these AREs abolished binding of both CPs and full-length HuR, demonstrating that in all cases, the binding seen is specific. Also of particular note, HuR-CP1 appeared to have a stronger association with the message than even full-length HuR. This observation may explain why HuR loses association with caspase-9 mRNA after 1.5 h, when HuR-CP1 is generated, possibly competing for association. By using various concentrations of GST-HuR and GST-HuR-CP1 in gel-shift experiments, we observed that HuR-CP1 binds the ARE2 probe with a significantly greater affinity (Kd) than HuR (Figures 5h and i). Thus, these observations show that HuR-CPs interact with the same proapoptotic target of HuR, although with different affinities, while suggesting that they may not associate with its antiapoptotic mRNA ligand ProT. As the generation of these CPs occurs at a time point when the association between HuR and caspase-9 mRNA decreases, yet the expression of caspase-9 protein increases, it is possible that these CPs are responsible for the upregulation of caspase-9.
Caspase-9 mRNA stability is promoted by HuR CPs in an ARE-dependent manner
As mutating the AREs of caspase-9 mRNA, which serve as HuR/CPs-binding sites, decreased its mRNA stability, we next tested if addition of either wtHuR or its two CPs could also affect the stability of this message. Unfortunately, a combined treatment of Act D and STS is fatal for cells, so instead we transiently transfected the GFP-tagged wtHuR or HuR-CP1 or HuR-CP2 or both CPs together (to mimic cleavage conditions where both are present), and caspase-9 mRNA stability were then verified (Figures 6a–e). HuR-CP2 had no stabilizing effect on caspase-9 mRNA (Figure 6d), whereas HuR-CP1 and wtHuR increased the half-life of this message (Figures 6b and c). Interestingly, the expression of both HuR-CPs together significantly increased the half-life of endogenous caspase-9 mRNA (from ∼4.8 to over 9 h) and the expression levels of its protein (Figures 6e–g). This increase in caspase-9 protein correlates with the increase seen during STS-induced stress, when HuR-CP1 and HuR-CP2 are endogenously generated (Figure 4c).
HuR-CPs stabilize and promote the expression of caspase-9 protein. (a–e) The effect of HuR and its CPs on the stability of caspase-9 mRNA. HeLa cells transfected with GFP, GFP-HuR, GFP-HuR-CP1, GFP-HuR-CP2 or both GFP-HuR-CPs (in a 1:1 ratio, but with half the amount of each) were treated with Act D, and the total mRNA was collected and run on a northern blot, which was then probed for caspase-9 and 18S (loading control) RNA. Quantification of three independent experiments (a representative blot of which is shown) for each condition was performed using the ImageQuant software and is shown in (b–e), with mean values±S.E.M. Graphs and half-lives were generated as described for Figure 3e, with the asterisk (e) indicating that the difference in half-lives is significant. (f, g) HuR-CPs increase the expression of caspase-9 protein. Following the same transfections described above, protein extracts of HeLa cells were analyzed by western blot, incubating with antibodies against GFP, caspase-9, HuR (endogenous HuR; end. HuR) and G3BP (loading control). The blot shown is representative of five independent experiments. Protein levels of the caspase-9 band were quantified relative to G3BP (g), with mean values graphed, with error bars representing the S.E.M. The asterisk denotes the significant difference between GFP and GFP-HuR-CP1+GFP-HuR-CP2-treated cells
These results suggest that it is when both HuR-CP1 and HuR-CP2 bind caspase-9 mRNA that a strong stabilizing effect takes place. To gain better mechanistic insight into the HuR-mediated post-transcriptional regulation of caspase-9 expression, we investigated if the effects of HuR-CPs on the stability of full-length caspase-9 mRNA depends on the presence of ARE-binding sites. Caspase-9−/− MEFs were co-transfected with either full-length caspase-9 or caspase-9 in which the AREs were mutated (Figure 3a), and with either GFP or a combination of GFP-HuR-CP1 and GFP-HuR-CP2. Addition of HuR-CPs had no effect on mRNA steady-state levels (Figures 7a and b), although mutating the AREs caused a decrease in caspase-9 mRNA levels, as expected (Figures 3b and c). However, the HuR-CPs did increase the half-life of wt caspase-9 mRNA (from ∼6.1 to ∼9.9 h), but not for its mutated counterpart (Figures 7c–e). By transfecting caspase-9-deficient MEFs as in Figure 7a, we saw that similarly to HeLa cells (Figures 6f and g), co-transfection of HuR-CP1 and HuR-CP2 caused an increase in caspase-9 protein (Figures 7f and g). When the AREs were mutated, however, HuR-CPs failed to produce this increase. This supports that HuR-CPs regulate caspase-9 through ARE binding, and explains why the AREs of caspase-9 mRNA are important in regulating caspase-9 expression (Figure 3). Interestingly, while the HuR-CPs were previously shown to trigger apoptosis in HeLa cells exposed to non-lethal stress24 and successfully rescued cell death following HuR depletion (Supplementary Figure 7), they failed to rescue apoptosis in caspase-9−/− MEFs (Figures 7h and i). This result confirms the importance of caspase-9 in the induction of cell death by HuR CPs (Figures 7h and i).
HuR CPs require AREs to stabilize caspase-9 mRNA. (a, b) HuR-CPs do not affect total RNA levels of caspase-9 mutants. GFP or GFP-HuR-CP1+HuR-CP2 (in a 1:1 ratio, but with half as much of each compared with GFP) were co-transfected with either full-length or mutated AREs containing caspase-9 into caspase-9−/− MEFs. Total RNA was isolated from these cells 24 h after transfection and was analyzed by northern blot, using probes against caspase-9 or 18S (loading control). Quantified caspase-9 values were taken relative to 18S intensity using the ImageQuant software, and were graphed in (b), with error bars representing the S.E.M. of six independent experiments. Not significant differences are indicated (NS) (c–e) HuR-CPs stabilize caspase-9 mRNA in the presence of AREs. Caspase-9−/− MEFs were transfected as described in (a) and were treated with Act D as described above. RNA was collected at the indicated time points, and probes against caspase-9 and 18S (loading control) were analyzed using northern blots (c). Average values were determined for full-length (d) and mut ARE (e) caspase-9 samples and graphed as a stability curve as in Figure 3e, with error bars representing the S.E.M. of three independent experiments. Half-lives were determined using Microsoft Excel as described above (Figures 3d and e), and averages are shown with S.E.M.s from three independent experiments. The asterisk indicates a significant difference. (f, g) HuR-CPs increase the protein level of caspase-9 in an ARE-dependent manner. (f) Protein lysates were taken from cells treated as described in (a) and were analyzed by western blot, probing with antibodies against caspase-9, GFP, HuR and α-tubulin (loading control). (g) Quantified caspase-9 levels were averaged, and are graphed, with error bars representing the S.E.M. of five independent experiments. The asterisk shows that the difference in means was significant. (h, i) HuR-CPs require caspase-9 to induce caspase-dependent apoptosis. wtMEFs and caspase-9−/− MEFs were treated with 1 μM STS for 6h, and caspase-9-deficient MEFs were first provided with either 150 nM AP-GST, 150 nM AP-HuR-GST or 75 nM each of AP-HuR-CP1-GST and AP-HuR-CP2-GST. Cells were lysed and analyzed by western blot using antibodies against PARP and caspase-3 cleavage products, caspase-9, HuR and α-tubulin (loading control). Quantified levels of caspase-3 and PARP cleavage products were averaged, and are graphed in (i), with error bars representing the S.E.M. of two independent experiments. All blots shown (a, c, f and h) are representative of the indicated number of experiments
Discussion
HuR is one of the best-characterized post-transcriptional regulators of gene expression. By affecting the fate of a variety of mRNA targets at different post-transcriptional levels, HuR modulates opposing cellular processes such as survival and apoptosis.15, 18, 24 The molecular mechanisms that mediate this functional dichotomy of HuR have remained unknown. In this study, we show that caspase-mediated cleavage switches HuR from promoting cell survival to activating apoptosis (Figure 8). We observed that full-length HuR associates with both pro- (caspase-9) and anti- (ProT) apoptotic mRNAs. Although ProT was previously established as a target of HuR,22 we identified the AREs of caspase-9 to which HuR binds, and found that these elements contribute to caspase-9 expression and mRNA stability (Figures 1 and 3). In normal conditions, HuR regulates the expression of both these factors; however, in response to lethal stress, HuR is cleaved, yielding two CPs that selectively promote caspase-9 expression in an ARE-dependent manner, via mRNA stabilization (Figure 7). Our data also show that the HuR-CP-mediated regulation of caspase-9 expression is required for it to contribute to caspase-mediated apoptosis.
Model illustrating how the cleavage of HuR alters its mRNA-based regulation of apoptosis. HuR has previously been shown to promote ProT expression by regulating the translation of this target.22 In this study, we found that HuR also associates with the mRNA of the proapoptotic caspase-9. Furthermore, we showed that in response to severe stress, when HuR is cleaved to generate two cleavage fragments (HuR-CP1 and HuR-CP2), changes in the expression of caspase-9, but not ProT, occur. This is explained by the fact that these CPs do not associate with ProT mRNA, but do associate, and stabilize, caspase-9 mRNA. Optimal caspase-9 stabilization depends on the presence of both cleavage products and their ARE-binding sites. This preferential effect of supporting only this proapoptotic mRNA target underscores the importance of HuR cleavage in promoting the HuR-mediated post-transcriptional regulation of apoptotic cell death
Unlike post-translational modifications such as phosphorylation, this caspase-mediated cleavage of HuR serves as a non-reversible switch for cells to commit to a process such as apoptosis. Caspases are known to cleave several target proteins to generate proapoptotic products.12, 33 Our previous work18 identified HuR as one such protein, as the consequence of its cleavage is the release of its ligand pp32/PHAPI, an enhancer of apoptosome activity.10 Here we report a novel function for the CPs of HuR and show that they mediate apoptosis by selectively promoting the expression of caspase-9. As far as we know, this is the first example showing that by cleaving a target protein, caspases promote the post-transcriptional regulation of a proapoptotic factor. Although caspase-9 was used as a model here, the extent to which HuR-CPs can regulate the expression of other apoptotic messages remains to be seen. Several of the messages that bound HuR by our RIP-ChIP approach are involved at different levels in mediating apoptosis, such as Bad, a promoter of cyt c release.34 As the HuR-CPs seem to influence the progression of apoptosis at different steps,18, 35 it would be valuable to determine if their multiple-target approach may rescue the ability of cancer cells to respond to chemotherapeutic treatments.
Our earlier work has indicated that only a fraction of HuR (∼10%) is cleaved.18, 24, 25 This raised the question of how cleaving only a portion of this highly expressed protein could have such a dramatic impact on cell fate. HuR shuttles between the nucleus and the cytoplasm, and while only a fraction of it (∼10–20%) localizes to the cytoplasm,36, 37 it is in this compartment where HuR exercises its main biological effects.15, 16, 17, 30, 38, 39, 40 Our recent papers18, 24, 25, 41 strengthen this idea and show that the HuR-CPs, generated from ∼50% of cytoplasmic HuR, are both sufficient and necessary for HuR-mediated effects.18 However, as not all cytoplasmic HuR is cleaved, wt HuR and both of its CPs are all present simultaneously, and thus may be affected by one another when exercising their effects. Indeed, HuR-CP1 promotes muscle development in murine myoblasts by enabling the cytoplasmic accumulation of wt HuR.25 HuR-CP1 causes a similar effect in response to lethal stress, by binding to the import factor transportin-2 and augmenting cytoplasmic localization of HuR.41 Here we show that together the two CPs are capable of stabilizing caspase-9 mRNA, even to a greater extent than wtHuR, and our previous data also indicate that the two CPs of HuR are most potent for inducing apoptosis when present simultaneously.24 Therefore, these data illustrate that the coordination between HuR and its CPs is an important aspect of the effects they have.
By generating two unique CPs, the ability for each to act differently also exists. This is comparable to observations made by Talwar et al.35 that HuR-CP1, and not wtHuR, inhibits the translation of c-myc mRNA. As previous observations indicated that HuR-CPs exercise their function by associating with various protein partners,18, 25 it would be interesting to determine if their capacity to post-transcriptionally regulate gene expression also depends on other ligands. HuR-mediated effects can involve microRNAs,42, 43 which are established inhibitors of gene expression involved in a variety of processes including apoptosis.44 This raises the possibility that HuR-CPs can also use these non-coding RNAs to modulate various cellular processes including death. Here, both of the HuR-CPs showed to have poor binding with ProT, but as HuR has been shown to repress translation,46 it is not unreasonable to imagine that perhaps one or both of the HuR-CPs could do the same to other antiapoptotic targets, differently from wtHuR. Investigating this may also reveal how HuR-CPs can distinguish between ARE-binding sites to allow them to have different affinities, and even prefer one message over another. As advances are made at delineating the preferred binding sites of HuR,46, 47 the importance of particular ARE sequences may be revealed. The nature of the caspase-9 AREs are particularly interesting, given that they demonstrated a clear stabilizing effect, despite the fact that AREs have broadly been demonstrated as destabilizing factors.
The balance between survival and death factors is tightly regulated to maintain cell life, and end it when necessary. Once a stimulus obligates a cell to commit to apoptosis, this balance becomes tilted and death ensues. Several studies have reported on apoptotic factors that undergo a role reversal following cleavage-based processing.48, 49, 50 HuR, clearly implicated in survival and death, is unlike these other apoptotic factors, as it is an upstream regulator of a multitude of survival and apoptosis-related genes.16 HuR was recently shown to be important for enabling apoptotic cell death,18 but this study demonstrates that through its cleavage, HuR can actually modify the post-transcriptional regulatory function it has, thus having a significant influence on cell physiology. This suggests that HuR is not just part of the apoptotic cell death execution process, but rather that through its cleavage, HuR may seal the fate of the cell.
Materials and Methods
Cell culture and transfection
HeLa CCL-2 cells, GFPμ-1 293 cells (ATCC, Manassas, VA, USA), as well as caspase-9−/− (generous donation from M. Saleh) and wtMEFs (kind donation from D. Ron) were grown and maintained in DME medium containing L-glutamine (Sigma-Aldrich, Oakville, ON, Canada) with fetal bovine serum (10%) and penicillin/streptomycin (Sigma-Aldrich). DNA plasmids (HA-Bax (kind gift from G. Shore), GFP, GFP-HuR,18 GFP-HuR-CP1, GFP-HuR-CP2,24 GFP-ProT (generous gift from M. Gorospe),22 FLAG-caspase-9 full-length and FLAG-caspase-9 mut AREs (see Supplementary Methods)) and siRNA duplexes (Control and siRNA-HuR51) were transfected as described previously24 when HeLa cells were approximately 70% (for DNA) or 50% (for siRNA) confluent, or when MEFs were 50% (for DNA) confluent, using Lipofectamine and Plus Reagents (Invitrogen, Burlington, ON, Canada) according to the manufacturer’s instructions. Samples were harvested 48h following the transfection for HeLa cells, and after 24 h for MEFs.
Preparation of cell extracts and immunoblotting
Cell extracts were prepared as described previously.24 Western blotting was carried out as described previously37 using antibodies against HuR,30 Ras-GAP SH3 domain-binding protein (G3BP),52 Bax (Santa Cruz , Santa Cruz, CA, USA), caspase-9, caspase-3 CP, PARP CP (Cell Signaling, Boston, MA, USA), GFP (Clontech, Mountan View, CA, USA), α-tubulin and HA (Sigma-Aldrich). Of note, the antibodies against PARP and caspase-3 have been shown by many laboratories to recognize only the cleavage forms of these proteins.6, 53
Immunoprecipitation and RT-PCR/qPCR
Cells were collected and immunoprecipitation was performed as described previously29 to isolate mRNA that was precipitated using antibodies against Sam68 (generous gift from S. Richard), HuR, GFP or an IgG control. mRNA was reverse-transcribed using Thermoscript RT-PCR Kit (Invitrogen) according to the manufacturer’s specifications. PCRs were then performed using primers for the amplification of caspase-9, hnRNP A1 and ProT (see Supplementary Methods), and PCR products were run on agarose or polyacrylamide gels. qPCR was performed using primers for amplification of caspase-9 and ProT (see Supplementary Methods) using SsoFast EvaGreen reagent (Bio-Rad, Mississauga, ON, Canada) according to the manufacturer’s details. Relative expression levels were determined by calculating 2−ΔΔCt, where ΔΔCt is the difference in Ct between the target gene in experimental samples and control sample(s). Ct is the number of cycles at which the amount of amplified target exceeds a fixed threshold.
Gel shift and determination of Kd values
See Supplementary Methods for more details.
mRNA isolation, northern blot analysis and stability assay
Cells were collected and resuspended in Trizol Reagent (Invitrogen) to isolate mRNA. Samples were analyzed by northern blot as described previously,54 using probes against caspase-9, HuR, ProT, GFP, GAPDH and 18S (see Supplementary Methods for detailed description of probe synthesis). For stability experiments, cells were treated with Act D (Sigma-Aldrich) at a concentration of 5 μg/ml for indicated amounts of time, after which mRNA was collected as described. Quantification was carried out using the ImageQuant software, normalizing band intensities against those seen in the loading control, and stability curves were generated by comparing the mRNA levels after each time treatment to the starting quantity before Act D treatment. Average points were graphed using Microsoft Excel, and half-lives were determined by generating trendlines for each experiment. Statistical analysis to determine significance on quantification values was performed using online GraphPad software. Two-tailed, unpaired t-test analyses were used when comparing two experimental means, and a one-sample t-test was performed when comparing one mean to a normalized control.
Polysome fractionation and slot blot
See Supplementary Methods for more details.
Cell treatments and assessing cell death
To induce apoptosis, cells were treated with STS (Sigma-Aldrich) at a 1 μM concentration. MG132 proteosome inhibitor (Sigma-Aldrich) was used at a 20 μM concentration. AP-conjugated proteins were also given to cells to either rescue expression or in an attempt to sensitize cells to death. Rescue of expression involved providing cells with a 50 nM dose of HuR, or 25 nM of each HuR-CP, 16 h before collection. For sensitization, cells were given 75 or 150 nM AP-conjugated proteins 1 h before a 6 h STS treatment. AP-GST, AP-HuR-GST, AP-HuR-CP1-GST and AP-HuR-CP2-GST proteins were produced and purified as described previously.24
Cell death was assessed by cell number, flow cytometry and by western blot. Using a Zeiss Axiovert 25 light microscope, cells were photographed with a Sony Cybershot DSC-S75 camera and the number of cells in different fields of view were counted. For flow cytometry, cells were washed with phosphate-buffered saline and then stained with Annexin V-FITC (BioVision, Milpitas, CA, USA) as described in the manufacturer’s protocol, and then counted. Western blot analysis was performed as described above and caspase-3 and PARP-CPs were quantified.
Abbreviations
- CP:
-
cleavage product
- ProT:
-
prothymosin α
- cyt c:
-
cytochrome c
- ARE:
-
AU-rich element
- UTR:
-
untranslated region
- AMD:
-
ARE-mediated decay
- AP:
-
antennepedia
- RIP-CHIP:
-
RNP-Immunoprecipitation-microchip
- hnRNP A1:
-
heterogenous nuclear ribonucleoprotein A1
- PARP:
-
poly (ADP-ribose) polymerase
- MEF:
-
murine embryonic fibroblast
- STS:
-
staurosporine
- ActD:
-
actinomycin D
References
Bree RT, Stenson-Cox C, Grealy M, Byrnes L, Gorman AM, Samali A . Cellular longevity: role of apoptosis and replicative senescence. Biogerontology 2002; 3: 195–206.
Lowe SW, Lin AW . Apoptosis in cancer. Carcinogenesis 2000; 21: 485–495.
Beere HM . ‘The stress of dying’: the role of heat shock proteins in the regulation of apoptosis. J Cell Sci 2004; 117 (Part 13): 2641–2651.
Fadeel B, Ottosson A, Pervaiz S . Big wheel keeps on turning: apoptosome regulation and its role in chemoresistance. Cell Death Differ 2008; 15: 443–452.
Youle RJ, Strasser A . The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol 2008; 9: 47–59.
Ho AT, Li QH, Hakem R, Mak TW, Zacksenhaus E . Coupling of caspase-9 to Apaf1 in response to loss of pRb or cytotoxic drugs is cell-type-specific. EMBO J 2004; 23: 460–472.
Pan G, O’Rourke K, Dixit VM . Caspase-9, Bcl-XL, and Apaf-1 form a ternary complex. J Biol Chem 1998; 273: 5841–5845.
Schmitt CA, Fridman JS, Yang M, Baranov E, Hoffman RM, Lowe SW . Dissecting p53 tumor suppressor functions in vivo. Cancer Cell 2002; 1: 289–298.
Tait SW, Green DR . Mitochondria and cell death: outer membrane permeabilization and beyond. Nat Mol Cell Biol 2010; 11: 621–632.
Jiang X, Kim HE, Shu H, Zhao Y, Zhang H, Kofron J et al. Distinctive roles of PHAP proteins and prothymosin-alpha in a death regulatory pathway. Science 2003; 299: 223–226.
Schafer ZT, Parrish AB, Wright KM, Margolis SS, Marks JR, Deshmukh M et al. Enhanced sensitivity to cytochrome c-induced apoptosis mediated by PHAPI in breast cancer cells. Cancer Res 2006; 66: 2210–2218.
Schafer ZT, Kornbluth S . The apoptosome: physiological, developmental, and pathological modes of regulation. Dev Cell 2006; 10: 549–561.
Hofseth LJ, Robles AI, Yang Q, Wang XW, Hussain SP, Harris C . P53: at the crossroads of molecular carcinogenesis and molecular epidemiology. Chest 2004; 125 (Suppl): 83S–85S.
Rozenfeld-Granot G, Krishnamurthy J, Kannan K, Toren A, Amariglio N, Givol D et al. A positive feedback mechanism in the transcriptional activation of Apaf-1 by p53 and the coactivator Zac-1. Oncogene 2002; 21: 1469–1476.
Abdelmohsen K, Kuwano Y, Kim HH, Gorospe M . Posttranscriptional gene regulation by RNA-binding proteins during oxidative stress: implications for cellular senescence. Biol Chem 2008; 389: 243–255.
Abdelmohsen K, Lal A, Kim HH, Gorospe M . Posttranscriptional orchestration of an anti-apoptotic program by HuR. Cell Cycle 2007; 6: 1288–1292.
Gallouzi IE, Steitz JA . Delineation of mRNA export pathways by the use of cell-permeable peptides. Science 2001; 294: 1895–1901.
Mazroui R, Di Marco S, Clair E, von Roretz C, Tenenbaum SA, Keene JD et al. Caspase-mediated cleavage of HuR in the cytoplasm contributes to pp32/PHAP-I regulation of apoptosis. J Cell Biol 2008; 180: 113–127.
von Roretz C, Gallouzi IE . Decoding ARE-mediated decay: is microRNA part of the equation? J Cell Biol 2008; 181: 189–194.
Fan XC, Steitz JA . Overexpression of HuR, a nuclear-cytoplasmic shuttling protein, increases the in vivo stability of ARE-containing mRNAs. EMBO J 1998; 17: 3448–3460.
Kawai T, Lal A, Yang X, Galban S, Mazan-Mamczarz K, Gorospe M . Translational control of cytochrome c by RNA-binding proteins TIA-1 and HuR. Mol Cell Biol 2006; 26: 3295–3307.
Lal A, Kawai T, Yang X, Mazan-Mamczarz K, Gorospe M . Antiapoptotic function of RNA-binding protein HuR effected through prothymosin alpha. EMBO J 2005; 24: 1852–1862.
Mazan-Mamczarz K, Galban S, De Silanes IL, Martindale JL, Atasoy U, Keene JD et al. RNA-binding protein HuR enhances p53 translation in response to ultraviolet light irradiation. Proc Natl Acad Sci USA 2003; 100: 8354–8359.
von Roretz C, Gallouzi IE . Protein kinase RNA/FADD/caspase-8 pathway mediates the proapoptotic activity of the RNA-binding protein human antigen R (HuR). J Biol Chem 2010; 285: 16806–16813.
Beauchamp P, Nassif C, Hillock S, van der Giessen K, von Roretz C, Jasmin BJ et al. The cleavage of HuR interferes with its transportin-2-mediated nuclear import and promotes muscle fiber formation. Cell Death Differ 2010; 17: 1588–1599.
Shore GC, Nguyen M . Bcl-2 proteins and apoptosis: choose your partner. Cell 2008; 135: 1004–1006.
Keene JD, Komisarow JM, Friedersdorf MB . RIP-Chip: the isolation and identification of mRNAs, microRNAs and protein components of ribonucleoprotein complexes from cell extracts. Nat Protoc 2006; 1: 302–307.
Tenenbaum SA, Carson CC, Lager PJ, Keene JD . Identifying mRNA subsets in messenger ribonucleoprotein complexes by using cDNA arrays. Proc Natl Acad Sci USA 2000; 97: 14085–14090.
Tenenbaum SA, Lager PJ, Carson CC, Keene JD . Ribonomics: identifying mRNA subsets in mRNP complexes using antibodies to RNA-binding proteins and genomic arrays. Methods 2002; 26: 191–198.
Gallouzi IE, Brennan CM, Stenberg MG, Swanson MS, Eversole A, Maizels N et al. HuR binding to cytoplasmic mRNA is perturbed by heat shock. Proc Nat Acad Sci USA 2000; 97: 3073–3078.
Itoh M, Haga I, Li QH, Fujisawa J . Identification of cellular mRNA targets for RNA-binding protein Sam68. Nucleic Acids Res 2002; 30: 5452–5464.
Dormoy-Raclet V, Menard I, Clair E, Kurban G, Mazroui R, Di Marco S et al. The RNA-binding protein HuR promotes cell migration and cell invasion by stabilizing the beta-actin mRNA in a U-rich-element-dependent manner. Mol Cell Biol 2007; 27: 5365–5380.
Cain K, Bratton SB, Cohen GM . The Apaf-1 apoptosome: a large caspase-activating complex. Biochimie 2002; 84: 203–214.
Shangary S, Johnson DE . Peptides derived from BH3 domains of Bcl-2 family members: a comparative analysis of inhibition of Bcl-2, Bcl-x(L) and Bax oligomerization, induction of cytochrome c release, and activation of cell death. Biochemistry 2002; 41: 9485–9495.
Talwar S, Jin J, Carroll B, Liu A, Gillespie MB, Palanisamy V . Caspase-mediated cleavage of RNA-binding protein HuR regulates c-Myc protein expression after hypoxic stress. J Biol Chem 2011; 286: 32333–32343.
van der Giessen K, Gallouzi IE . Involvement of transportin 2-mediated HuR import in muscle cell differentiation. Mol Biol Cell 2007; 18: 2619–2629.
Brennan CM, Gallouzi IE, Steitz JA . Protein ligands to HuR modulate its interaction with target mRNAs in vivo. J Cell Biol 2000; 151: 1–14.
Abdelmohsen K, Pullmann R, Lal A, Kim HH, Galban S, Yang X et al. Phosphorylation of HuR by Chk2 regulates SIRT1 expression. Mol Cell 2007; 25: 543–557.
Doller A, Akool el S, Huwiler A, Muller R, Radeke HH, Pfeilschifter J et al. Posttranslational modification of the AU-rich element binding protein HuR by protein kinase Cdelta elicits angiotensin II-induced stabilization and nuclear export of cyclooxygenase 2 mRNA. Mol Cell Biol 2008; 28: 2608–2625.
Doller A, Pfeilschifter J, Eberhardt W . Signalling pathways regulating nucleo-cytoplasmic shuttling of the mRNA-binding protein HuR. Cell Signal 2008; 20: 2165–2173.
von Roretz C, Macri AM, Gallouzi IE . Transportin 2 regulates apoptosis through the RNA-binding protein HuR. J Biol Chem 2011; 286: 25983–25991.
Kim HH, Kuwano Y, Srikantan S, Lee EK, Martindale JL, Gorospe M . HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev 2009; 23: 1743–1748.
Bhattacharyya SN, Habermacher R, Martine U, Closs EI, Filipowicz W . Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 2006; 125: 1111–1124.
Lima RT, Busacca S, Almeida GM, Gaudino G, Fennell DA, Vasconcelos MH . MicroRNA regulation of core apoptosis pathways in cancer. Eur J Cancer 2011; 47: 163–174.
Meng Z, King PH, Nabors LB, Jackson NL, Chen CY, Emanuel PD et al. The ELAV RNA-stability factor HuR binds the 5'-untranslated region of the human IGF-IR transcript and differentially represses cap-dependent and IRES-mediated translation. Nucleic Acids Res 2005; 33: 2962–2979.
Lebedeva S, Jens M, Theil K, Schwanhausser B, Selbach M, Landthaler M et al. Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol Cell 2011; 43: 340–352.
Mukherjee N, Corcoran DL, Nusbaum JD, Reid DW, Georgiev S, Hafner M et al. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol Cell 2011; 43: 327–339.
Cheng EH, Kirsch DG, Clem RJ, Ravi R, Kastan MB, Bedi A et al. Conversion of Bcl-2 to a Bax-like death effector by caspases. Science 1997; 278: 1966–1968.
Clem RJ, Cheng EH, Karp CL, Kirsch DG, Ueno K, Takahashi A et al. Modulation of cell death by Bcl-XL through caspase interaction. Proc Natl Acad Sci USA 1998; 95: 554–559.
Breckenridge DG, Stojanovic M, Marcellus RC, Shore GC . Caspase cleavage product of BAP31 induces mitochondrial fission through endoplasmic reticulum calcium signals, enhancing cytochrome c release to the cytosol. J Cell Biol 2003; 160: 1115–1127.
van der Giessen K, Di-Marco S, Clair E, Gallouzi IE . RNAi-mediated HuR depletion leads to the inhibition of muscle cell differentiation. J Biol Chem 2003; 278: 47119–47128.
Mazroui R, Sukarieh R, Bordeleau ME, Kaufman RJ, Northcote P, Tanaka J et al. Inhibition of ribosome recruitment induces stress granule formation independently of eukaryotic initiation factor 2alpha phosphorylation. Mol Biol Cell 2006; 17: 4212–4219.
Chang NC, Nguyen M, Germain M, Shore GC . Antagonism of Beclin 1-dependent autophagy by BCL-2 at the endoplasmic reticulum requires NAF-1. EMBO J 2010; 29: 606–618.
Di Marco S, Mazroui R, Dallaire P, Chittur S, Tenenbaum SA, Radzioch D et al. NF-(kappa)B-mediated MyoD decay during muscle wasting requires nitric oxide synthase mRNA stabilization, HuR protein, and nitric oxide release. Mol Cell Biol 2005; 25: 6533–6545.
Acknowledgements
We offer our sincere thanks to Dr. Sergio Di Marco for critical review of this manuscript and insightful discussion, Drs. Douglas Green and Judy Lieberman for sharing their insight on apoptotic pathways and to Drs. Gordon Shore and Mai Nguyen for technical assistance with assessing apoptosis. CvR was the recipient of a Canadian Institutes of Health Research (CIHR) Doctoral Research Award, AMM was the recipient of a Natural Sciences and Engineering Research Council of Canada summer undergraduate student award and is supported by the Maysie MacSporran Graduate Studentship and the CIHR/FRSQ (Fonds de Recherche en Sante du Quebec) training grant in cancer research (FRN53888) of the McGill Integrated Cancer Research Training Program (MICRTP), OD received a CIHR Master’s Training Award, VDR was a Research Fellow of the Terry Fox Foundation through an award from the National Cancer Institute of Canada (NCIC) and JFM is supported by an MICRTP fellowship. This work was also supported by an NCIC operating grant (016247) and a CIHR operating grant (MOP-89798). IEG is a recipient of a TierII Canada Research Chair.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by RA Knight
Supplementary Information accompanies the paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
von Roretz, C., Jin Lian, X., Macri, A. et al. Apoptotic-induced cleavage shifts HuR from being a promoter of survival to an activator of caspase-mediated apoptosis. Cell Death Differ 20, 154–168 (2013). https://doi.org/10.1038/cdd.2012.111
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2012.111
Keywords
This article is cited by
-
HuR drives lung fibroblast differentiation but not metabolic reprogramming in response to TGF-β and hypoxia
Respiratory Research (2021)
-
A macrophage-specific lncRNA regulates apoptosis and atherosclerosis by tethering HuR in the nucleus
Nature Communications (2020)
-
PARP1 promotes gene expression at the post-transcriptional level by modulating the RNA-binding protein HuR
Nature Communications (2017)
-
Repression of caspase-3 and RNA-binding protein HuR cleavage by cyclooxygenase-2 promotes drug resistance in oral squamous cell carcinoma
Oncogene (2017)
-
Physiological regulation of the heat shock response by glutamine: implications for chronic low-grade inflammatory diseases in age-related conditions
Nutrire (2016)