Abstract
Apoptosis inducing factor (AIF) is a mitochondrial oxidoreductase that scavenges reactive oxygen species under normal conditions. Under certain stresses, such as exposure to N -methyl- N′ -nitro- N′ -nitrosoguanidine (MNNG), AIF is truncated and released from the mitochondria and translocated into the nucleus, where the truncated AIF (tAIF) induces caspase-independent cell death. However, it is unknown how cells decide to kill themselves or operate ways to survive when they encounter stresses that induce the release of tAIF. Here, we demonstrated that USP2 and CHIP contribute to the control of tAIF stability. USP2 deubiquitinated and stabilized tAIF, thus promoting AIF-mediated cell death. In contrast, CHIP ubiquitinated and destabilized tAIF, thus preventing the cell death. Consistently, CHIP-deficient cells showed an increased sensitivity to MNNG. On the other hand, knockdown of USP2 attenuated MNNG-induced cell death. Moreover, exposure to MNNG caused a dramatic decrease in CHIP level, but not that of USP2, concurrent with cell shrinkage and chromatin condensation. These findings indicate that CHIP and USP2 show antagonistic functions in the control of AIF-mediated cell death, and implicate the role of the enzymes as a switch for cells to live or die under stresses that cause tAIF release.
Similar content being viewed by others
Main
AIF is a mitochondrial oxidoreductase that normally functions in scavenging reactive oxygen species.1 Harlequin (Hq) mice, which exhibit decreased expression of AIF (∼20% of normal mice), show increased sensitivity to oxidative stress.2 In addition, knockout models of mouse and Drosophila indicate that AIF is essential to mitochondrial oxidative phosphorylation.3, 4 AIF deficiency in conditional knockout mice impairs the activity and expression of mitochondrial respiratory complex I (see ref. 5). Targeted deletion of AIF also protects the animals from obesity and diabetes.6 Therefore, AIF appears to have important roles in the protection against oxidative stress as well as in the energy metabolism.7
Under certain stresses, however, AIF induces caspase-independent cell death, showing the characteristics of large-scale DNA breakage, chromatin condensation, and cell shrinkage.8, 9, 10 For example, some genotoxic agents, such as MNNG, induce the caspase-independent cell death.11 The cell-death process is initiated by the cleavage of AIF that is normally inserted in the inner mitochondrial membrane, the release of tAIF to the cytoplasm, and the translocation into the nucleus.12 The tAIF in the nucleus associates with histone H2AX and induces chromatinolysis for execution of cell death.13 It has been shown that the AIF release process requires the activation of calpain, which in turn activates Bax that functions in permeabilization of the mitochondrial outer membrane.14, 15 It has also been shown that poly (ADP-ribose) (PAR) polymer, the product of PAR polymerase-1 (PARP1), serves as a cell death signal that induces AIF release and translocation to the nucleus, but independently of calpain activation.10, 16 Thus, different mechanisms for AIF release seems to operate depending on specific conditions or signaling pathways involved. Intriguingly, a brain-specific AIF isomer, AIF2, has recently been isolated, implicating its neuron-specific function.17
Ubiquitin-dependent proteolysis has an essential role in the regulation of a variety of cellular processes, including cell proliferation, differentiation, and apoptosis.18, 19 Ubiquitin, a 76-amino acid polypeptide, is covalently attached to target proteins by a cascade enzyme system consisting of ubiquitin activating (E1), conjugating (E2), and ligating (E3) enzymes.18 Reversal of ubiquitination catalyzed by deubiquitinating enzymes also has important roles in the regulation of numerous biological pathways, such as by stabilization of critical regulatory proteins.20, 21
CHIP is a U-box-containing ubiquitin E3 ligase that mediates the degradation of misfolded proteins.22, 23 CHIP also functions as a co-chaperone in assisting Hsp70-dependent refolding of nonnative proteins.24 On the other hand, USP2, also called UBP69, is a deubiquitinating enzyme that has a unique N-terminal extension of 28 kDa in addition to the C-terminal domain, which is identical to its isoform, called UBP41.25, 26 Like USP7, USP2 deubiquitinates and stabilizes Mdm2, an ubiquitin E3 ligase for p53, thus functioning in the control of p53 degradation.27 USP2 also removes ubiquitin from fatty acid synthase (FAS), which is often overexpressed in aggressive human tumors, such as prostate cancer.28 Therefore, USP2 has been implicated in the survival of prostate cancer cells through FAS stabilization. On the other hand, UBP41 has been reported as a pro-apoptotic protein, as its overexpression elicits classical caspase-dependent cell death in human cells.29 However, overexpression of USP2, unlike that of UBP41, does not evoke any sign of caspase-dependent cell death.28
Here, we show that CHIP E3 ligase ubiquitinates tAIF, whereas USP2 removes ubiquitin from tAIF. Moreover, CHIP was found to attenuate tAIF-mediated cell death, in contrast to USP2 that accelerates it. Thus, the antagonistic functions of CHIP and USP2 appear to have a critical role in the control of AIF-mediated, caspase-independent cell death.
Results
USP2 interacts with tAIF
To identify target substrates of USP2, we performed proteomic analysis of the proteins that interacted with the overexpressed, Flag-tagged catalytically inactive mutant of USP2, in which Cys276 was replaced by Ala. Henceforth, the USP2 mutant was referred to as C276A. Among the identified proteins that interact with C276A (Supplementary Table 1), we chose AIF for further investigation to determine the role of USP2 in AIF-mediated cell death. To confirm the interaction between USP2 and AIF, Flag-tagged USP2 or C276A was expressed in HEK293T cells with tAIF-V5-His. Immunoprecipitation analysis revealed that tAIF interacted with both USP2 and C276A (Figure 1a), indicating that the catalytic activity of USP2 is not required for its interaction with tAIF. We next examined the interaction between USP2 and tAIF under in vitro conditions. Purified tAIF was coprecipitated with GST-USP2, but not with GST (Figure 1b), indicating that USP2 directly binds to tAIF. To determine whether endogenous forms of tAIF and USP2 interact with each other, HeLa cells cultured in the absence and presence of MNNG were disrupted by homogenization and centrifused. Immunoprecipitation analysis of the supernatant fraction revealed that endogenous tAIF coprecipitates with USP2 when the cells were treated with MNNG (Figure 1c). These results confirm previous findings that tAIF is cleaved off from AIF and released into the cytosol from the mitochondria under certain genotoxic stresses, such as exposure to MNNG.10, 11 Collectively, these observations indicate that USP2 interacts with tAIF under both in vivo and in vitro conditions.
USP2 interacts with tAIF. (a) tAIF-V5-His was expressed in HEK293T cells with Flag-USP2 or Flag-C276A. After incubation with 10 μM MG132 for 4 h, cell lysates were subjected to immunoprecipitation with anti-Flag antibody followed by immunoblot with anti-V5 and anti-Flag antibodies. Cell lysates were also directly probed with the same antibodies. (b) Purified tAIF was incubated with GST or GST-USP2 at 4°C for 12 h and then with GSH-Sepharose for the next 1 h. Proteins bound to the beads were subjected to SDS-PAGE followed by immunoblot with anti-AIF antibody. Proteins in the gels were also stained with Coomassie blue R-250. (c) HeLa cells were incubated with 0.5 mM MNNG for 15 min, washed with PBS, and further incubated for the next for 14 h. After treatment with 10 μM MG132 for 4 h, cells were disrupted by using Dounce homogenizer followed by centrifugation. The supernatants were subjected to immunoprecipitation with anti-IgG, anti-USP2, and anti-AIF antibodies followed by immunoblot with the same antibodies. Both the supernatant and pellet fractions were also directly probed with the antibodies against α-tubulin and lamin A/C, which are the marker proteins for the cytoplasm and the nucleus, respectively. (d) Deletions of tAIF (Δ1–Δ6) were generated and tagged with V5-His to their C-termini. (e) The tAIF deletions were expressed in HEK293T cells with and without Flag-USP2. Cell lysates were subjected to pull-down with anti-Flag M2 antibody-conjugated agarose (Flag-M2-bead) followed by immunoblot with anti-V5 antibody. Cell lysates were also directly probed with anti-V5 and anti-Flag antibodies. The asterisk indicates the IgG heavy chain and the double asterisk shows the IgG light chain. (f) Deletions of tAIF (Δ7 and Δ8) were generated and subcloned into pEBG vector for the expression of GST-fused deletions in mammalian cells. (g) The tAIF deletions were expressed in HEK293T cells with and without Flag-USP2. Cell lysates were subjected to pull-down with Flag-M2-bead followed by immunoblot with anti-GST antibody. Cell lysates were also directly probed with anti-GST and anti-Flag antibodies. (h) Deletions of USP2 were generated and tagged with Flag. (i) USP2 deletions (UΔ1 and UΔ2) were expressed in HEK293T cells with and without tAIF-V5-His. Cell lysates were subjected to pull-down with Flag-M2-bead followed by immunoblot with anti-V5 and anti-Flag antibodies. Cell lysates were also directly probed with the same antibodies
To locate the USP2-binding region within AIF, we generated various deletions of tAIF and tagged with V5-His to their C-termini (Figure 1d). Immunoprecipitation analysis revealed that both the deletion fragments Δ4 (amino acids 221–363) and Δ5 (342–492) could interact with USP2 (Figure 1e). In addition, the fragment Δ3 that lacks the overlapping sequence (342–363) in Δ4 and Δ5 was also capable of interacting with USP2. However, the fragment Δ6 that lacks the sequence (221–492) comprising both Δ4 and Δ5 could not interact with USP2, suggesting that tAIF has at least two USP2-binding sites within the sequence 221–492. To confirm this finding, GST-fused fragments Δ7 (221–341) and Δ8 (364–492) were generated (Figure 1f). Immunoprecipitation analysis showed that both Δ7 and Δ8 could interact with USP2 (Figure 1g). Taken together, these results indicate that tAIF has two USP2-binding sites: one in the sequence of 221–341 and the other in that of 364–492.
To determine the tAIF-binding region within USP2, deletions of USP2 were generated (Figure 1h). Immunoprecipitation analysis showed that tAIF interacted with the C-terminal catalytic domain of USP2, but not to its N-terminal extension (Figure 1i).
USP2 deubiquitinates tAIF
To determine whether USP2 is capable of deubiquitinating tAIF, HA-ubiquitin (HA-Ub) and tAIF-V5-His were expressed in HEK293T cells with Flag-USP2 or Flag-C276A. NTA pull-down analysis revealed that tAIF was ubiquitinated and this modification could be reversed by USP2, but not by C276A or USP3 (Figure 2a). In addition, the extents of ubiquitination in whole-cell lysates were not much affected by overexpression of USP2 or USP3. These results suggest that tAIF is a specific substrate of USP2. To confirm this finding, we examined the effect of knockdown of endogenous USP2 by using two USP2-specific shRNAs (shUSP2-1 and shUSP2-2). These shRNAs effectively reduced the mRNA level of USP2, and their transfection (Figure 2b), but not that of nonspecific shRNA (siNS), led to a significant increase in tAIF ubiquitination (Figure 2c), indicating that USP2 is responsible for deubiquitnation of tAIF. To determine whether USP2-mediated deubiquitination affects the stability of tAIF, tAIF-V5-His was expressed in HEK293T cells transfected with shUSP2s or shNS. Cells were then incubated with DMSO or MG132. The level of tAIF was significantly decreased by USP2 knockdown in cells treated with DMSO, and this decrease was prevented upon treatment of cells with MG132 (Figure 2d), indicating that USP2 can stabilize tAIF by preventing its degradation by the proteasome.
USP2 deubiquitinates tAIF. (a) HA-Ub and tAIF-V5-His were expressed in HEK293T cells with Flag-tagged USP2, C276A, or USP3. After incubation with 10 μM MG132 for 4 h, cell lysates were subjected to pull-down with NTA resins followed by immunoblot with anti-HA or anti-V5 antibody. Cell lysates were also directly probed with anti-Flag, anti-V5, and anti-HA antibodies. (b) Cells were transfected with shUSP2-1, shUSP2-2, or shNS. Total RNAs were prepared and subjected to RT-PCR using a primer specific to USP2. (c) shUSP2-1, shUSP2-2, or shNS was transfected to cells with pcDNA4-tAIF-V5-His and pcDNA3-HA-Ub. After incubation with 10 μM MG132 for 4 h, cell lysates were treated as in (a). β-Actin was used as a loading control. (d) Cells prepared as in (c) were cultured for 24 h and treated with DMSO or 10 μM MG132 for 4 h. Cell lysates were subjected to immunoblot with respective antibodies
CHIP interacts with tAIF
As both CHIP and AIF interact with Hsp70 and CHIP acts as an ubiquitin E3 ligase for Hsp70 (see refs. 30,31), we hypothesized that CHIP might directly or indirectly interact with AIF. To test this hypothesis, tAIF-V5-His was expressed in HEK293T cells with and without Myc-CHIP. Immunoprecipitation analysis revealed that tAIF interacted with CHIP (Figure 3a). As this interaction under in vivo conditions could be mediated by Hsp70, we examined the interaction between CHIP and tAIF under in vitro conditions. Purified tAIF was coprecipitated with GST-CHIP, but not with GST (Figure 3b), indicating CHIP directly binds to tAIF. To determine whether endogenous forms of tAIF and CHIP interact with each other, HeLa cells cultured in the absence and presence of MNNG were homogenized and centrifused. Immunoprecipitation analysis of the supernatant fraction revealed that endogenous tAIF coprecipitates with CHIP when the cells were treated with MNNG (Figure 3c). Taken together, these results indicate that CHIP can interact with tAIF under both in vivo and in vitro conditions.
CHIP interacts with tAIF. (a) The tAIF-V5-His, Myc-CHIP, or both were expressed in HEK293T cells. After incubation with 10 μM MG132 for 4 h, cell lysates were subjected to immunoprecipitation with anti-Myc antibody followed by immunoblot with anti-V5 or anti-Myc antibody. Cell lyastes were also directly probed with the same antibodies. (b) Purified tAIF was incubated with GST or GST-CHIP at 4°C for 12 h and then with GSH-Sepharose for the next 1 h. Bound proteins were subjected to SDS-PAGE followed by immunoblot with anti-AIF antibody. Proteins in the gels were also stained with Coomassie blue R-250. (c) HeLa cells were incubated with 0.5 mM MNNG for 15 min, washed with PBS, and further incubated for the next 14 h. After treatment with 10 μM MG132 for 4 h, cells were disrupted by using Dounce homogenizer followed by centrifugation. The supernatants were subjected to immunoprecipitation with anti-IgG, anti-CHIP, and anti-AIF antibodies followed by immunoblot with the same antibodies. Both the supernatant and pellet fractions were also directly probed with anti-α-tubulin and lamin A/C antibodies. (d) Deletions of tAIF (Δ1–Δ6) were prepared as in Figure 1d. (e) The tAIF deletions were expressed in HEK293T cells with and without Myc-CHIP. Cell lysates were subjected to immunoprecipitation with anti-Myc antibody followed by immunoblot with anti-V5 antibody. Cell lysates were also directly probed with anti-V5 and anti-Myc antibodies. The asterisk indicates the IgG heavy chain and the double asterisk shows the IgG light chain. (f) Deletions of tAIF (Δ7 and Δ8) were prepared as in Figure 1f. (g) The tAIF deletions were expressed in HEK293T cells with and without Myc-CHIP. Cell lysates were subjected to immunoprecipitation with anti-Myc antibody followed by immunoblot with GST antibody. Cell lysates were also directly probed with anti-GST and anti-Myc antibodies. (h) Deletions of CHIP were generated and tagged with Myc. (i) The CHIP deletions (CΔ1 and CΔ2) were expressed in HEK293T cells with and without tAIF-V5-His. Cell lysates were subjected to immunoprecipitation with anti-Myc antibody followed by immunoblot with anti-V5 or anti-Myc antibody. Cell lysates were also directly probed with the same antibodies
To map the CHIP-binding region within AIF, we used the same tAIF deletions, which were generated for locating USP2-binding regions (Figure 3d). Immunoprecipitation analysis revealed that CHIP interacted with the same regions where USP2 binds; that is, both the fragments Δ4 and Δ5 as well as the fragment Δ3 that lacks the overlapping sequence in Δ4 and Δ5 (Figure 3e). These results suggest that CHIP binds to at least two regions within the sequence 221–492 of tAIF. Immunoprecipitation analysis showed that both the GST-fused fragments Δ7 and Δ8 could interact with CHIP (Figures 3f and g), indicating that like USP2, CHIP binds to two different regions in tAIF: one in the sequence of 221–341 and the other in that of 364–492. However, CHIP and USP2 did not compete with each other for binding to the identified regions in tAIF (data not shown).
To determine the tAIF-binding region within CHIP, deletions of CHIP were generated (Figure 3h). Immunoprecipitation analysis showed that tAIF interacted with the N-terminal tetratricopeptide-repeat domain of CHIP, but not to its C-terminal U-box domain (Figure 3i).
CHIP ubiquitinates tAIF
To determine whether CHIP could act as an ubiquitin E3 ligase for tAIF, HA-Ub and tAIF-V5-His were expressed in HEK293T cells with Myc-CHIP or its mutant form lacking the U-box (CHIPΔU). NTA pull-down analysis revealed that the level of ubiquitinated tAIF was significantly increased by the expression of CHIP, but not by that of CHIPΔU (Figure 4a). CHIP also ubiquitinated tAIF under in vitro conditions, and this ubiquitination could be reversed by USP2 (Figure 4b), showing that tAIF is a substrate of CHIP as well as of USP2. To confirm the role of CHIP as an E3 ligase for tAIF, tAIF-V5-His and HA-Ub were expressed with and without Myc-CHIP or Myc-CHIPΔU in CHIP−/− MEF cells, which were obtained from CHIP-deficient mice that we had generated.32 CHIP−/− mice showed a decrease in viability and body weight (data not shown), as reported previously.32 NTA pull-down analysis revealed that tAIF ubiquitination could be observed when CHIP−/− cells were complemented with CHIP, but not without it or with CHIPΔU (Figure 4c). These results confirm that CHIP serves as an ubiquitin E3 ligase for tAIF.
CHIP ubiquitinates tAIF. (a) HA-Ub and tAIF-V5-His were expressed in HEK293T cells with Myc-CHIP or Myc-CHIPΔU. After treatment with 10 μM MG132 for 4 h, cell lysates were subjected to pull-down with NTA resins followed by immunoblot with anti-HA or anti-V5 antibody. Cell lysates were also directly probed with anti-V5, anti-Myc, and anti-HA antibodies. (b) Purified tAIF, His-Uba1 (E1), UbcH5A (E2), and ubiquitin (Ub) were incubated with purified CHIP, CHIPΔU, or/and USP2 at 37°C for 1 h. The samples were subjected to immunoblot with anti-AIF antibody. Proteins in the gel were also stained by Coomassie blue R-250. (c) HA-Ub and tAIF-V5-His were expressed in CHIP−/− cells with Myc-CHIP or Myc-CHIPΔU. Cell lysates were subjected to pull-down with NTA resins followed by immunoblot with anti-HA antibody. Cell lysates were also probed with anti-V5, anti-Myc, and anti-HA antibodies. (d) CHIP+/+ and CHIP−/− cells were incubated with 0.25 mM MNNG for 15 min and washed with PBS. After incubation with 5 μM MG132 for 6 h, cells were disrupted by using Dounce homogenizer followed by centrifugation. Both supernatant and pellet fractions were subjected to immunoprecipitation with anti-AIF antibody followed by immunoblot with anti-ubiquitin antibody. They were also directly probed with anti-AIF, anti-CHIP, anti-αtubulin, anti-lamin A/C, and anti-ubiquitin antibodies. (e) The tAIF-V5-His was expressed in HEK293T cells with Myc-CHIP or Myc-CHIPΔU. After culturing for 24 h, cells were treated with DMSO or 10 μM MG132 for 4 h. Cell lysates were then subjected to immunoblot with respective antibodies. (f) Experiments were performed as in (e), except that EGFP-AIF was expressed in place of tAIF-V5-His
It has been shown that CHIP can generate both Lys48- and Lys63-linked polyubiquitin chains and USP2 can hydrolyze both chains.33 To determine whether CHIP can ubiquitinate tAIF through Lys48- and/or Lys63-linked isopeptide bonds, we made HA-tagged ubiquitin variants, in which all Lys residues, except either Lys48 or Lys63, were replaced by Arg (termed Ub/K48 and Ub/K63, respectively). HA-Ub and its variants were then expressed in HEK293T cells with tAIF-V5-His and Myc-CHIP. Immunoprecipitation analysis showed that CHIP could generate both Lys48- and Lys63-linked polyubiquitin conjugates of tAIF (Supplementary Figure 1). In addition, it is likely that USP2 hydrolyzes both types of polyubiquitin chains, as its overexpression led to almost complete elimination of ubiquitin chains from tAIF (see Figures 2a and 4b).
To determine whether endogenous CHIP could ubiquitinate endogenous tAIF, CHIP+/+ and CHIP−/− cells incubated with and without MNNG were disrupted by homogenization and centrifused. Immunoprecipitation analysis showed that tAIF ubiquitination could be seen only in the supernatant fraction of MNNG-treated CHIP+/+ cells (Figure 4d). In addition, MNNG treatment led to the generation of tAIF in both CHIP+/+ and CHIP−/− cells. These results indicate that CHIP is capable of ubiquitinating tAIF that was released from the mitochondria in response to MNNG treatment. To determine whether the CHIP-mediated ubiquitination leads to destabilization of tAIF, Myc-CHIP or Myc-CHIPΔU was expressed in HEK293T cells with tAIF-V5-His followed by incubation with DMSO or MG132. The level of tAIF was significantly decreased by the expression of CHIP, but not by that of CHIPΔU (Figure 4e). This decrease in tAIF level was prevented by MG132, indicating that the proteasome is responsible for tAIF degradation. On the other hand, the level of EGFP-tagged intact AIF remained unchanged whether CHIP or CHIPΔU was overexpressed and whether MG132 was treated or not (Figure 4f), consistent with our finding that CHIP localizes exclusively in the cytosol (see Figure 1c) and cannot ubiquitinate AIF that resides in the inner mitochondrial membrane.
To confirm whether both CHIP and USP2 co-localize in the cytosol with tAIF for catalyzing reversible ubiquitination of tAIF, HeLa cells treated with MNNG were incubated for increasing periods followed by digitonin treatment and centrifugation. Although the intact uncleaved AIF remained in the pellet (i.e., the mixture of the nuclear and the mitochondrial fraction), tAIF was released into the supernatant (the cytosolic fraction) and colocalized with both CHIP and USP2 (Supplementary Figure 2). Moreover, the isolated mitochondria contained little or no CHIP or USP2. These results indicate that both CHIP and USP2 act on tAIF in the cytosol.
USP2 promotes MNNG-induced cell death
As USP2 stabilizes tAIF, we examined whether USP2 might be involved in the control of tAIF-mediated cell death. HeLa cells expressing Flag-tagged USP2 or C276A were treated with MNNG for 15 min, washed, and incubated for increasing periods. In cells expressing USP2, cell shrinkage and chromatin condensation could be seen from about 4 h after MNNG treatment (Figure 5b). In contrast, the morphological changes could be observed at least 12 h after MNNG treatment in cells transfected with an empty vector (Figure 5a) or a vector expressing C276A (Figure 5c). Supplementary Figure 3 shows the statistical data for the changes in the number of dead cells and the immunoblot data for the changes in the level tAIF during the culture periods. Moreover, knockdown of USP2 led to a marked decrease in MNNG-induced cell shrinkage and chromatin condensation (Supplementary Figure 4). In addition, FACS analysis showed that USP2 knockdown led to the prevention of MNNG-mediated increase in cell number in sub Go/G1 stage (Supplementary Figure 5a). Collectively, these results indicate that USP2 accelerates MNNG-induced cell death, likely by deubiquitination and stabilization of tAIF.
USP2 promotes MNNG-induced cell death. HeLa cells were transfected with (a) an empty vector (pFlag-CMV-2), (b) pFlag-USP2, or (c) pFlag-C267A. After incubation for 24 h, they were treated with 0.5 mM MNNG for 15 min, washed, and incubated for the indicated periods. Cells were fixed, stained with anti-USP2 (red) or anti-AIF antibody (green) and DAPI (blue), and observed under a confocal microscope
CHIP blocks tAIF-mediated cell death
Overexpression of tAIF alone has been shown to induce cell death.9, 15 As CHIP could mediate the degradation of tAIF by the proteasome, we examined whether overexpression of CHIP could block tAIF-induced cell death. As expected, like MNNG treatment, expression of tAIF-V5-His alone in HeLa cells resulted in cell shrinkage and chromatin condensation (Figure 6a, II and III). However, coexpression of Myc-CHIP, but not Myc-CHIPΔU, prevented tAIF-mediated cell death (Figure 6a, VI and VII). On the other hand, no sign of cell death could be observed in cells expressing Myc-CHIP or Myc-CHIPΔU alone (Figure 6a, IV and V) as in control cells (Figure 6a, I). Supplementary Figure 6 shows the statistical data for the changes in the number of dead cells and the immunoblot data for the expression levels of overproduced CHIP and AIF and endogenous tAIF. These results indicate that CHIP is capable of preventing tAIF-mediated cell death, likely by ubiquitination and degradation of tAIF.
CHIP prevents tAIF-mediated cell death. (a) HeLa cells transfected with an empty vector were incubated in the absence (I) or presence of MNNG (II) for 15 min. After incubation, cells were washed, cultured for 24 h, and stained with anti-CHIP (red) and anti-AIF antibodies (green) and DAPI (blue). Cells were also transfected with pcDNA4-tAIF-V5-His (III), pcDNA3-Myc-CHIP (IV), or pcDNA3-Myc-CHIPΔU (V). The pcDNA4-tAIF-V5-His was transfected to cells with pcDNA3-Myc-CHIP (VI) or pcDNA3-Myc-CHIPΔU (VII). After incubation for 48 h, III-V was stained as above and VI and VII were stained with anti-Myc (red) and anti-V5 antibodies (green). They were then observed under a confocal microscope. (b) CHIP+/+ and CHIP−/− cells were treated with MNNG for 15 min followed by washing with PBS. After incubation for the indicated times, they were fixed, stained with anti-AIF antibody (green) and DAPI (blue), and observed under a fluorescent microscope
To verify whether endogenous CHIP is involved in the control of AIF-mediated cell death, both CHIP−/− and CHIP+/+ cells were treated with MNNG. In CHIP−/− cells, cell shrinkage and chromatin condensation could be observed as early as 1 h after MNNG treatment (Figure 6b). However, the morphological changes could be seen 8 h after MNNG treatment in CHIP+/+ cells. Supplementary Figure 7 shows the statistical data for the changes in the number of dead cells and the immunoblot data for the changes in the level of tAIF during the culture periods. In addition, FACS analysis revealed that upon MNNG treatment, the number of CHIP−/− cells in sub Go/G1 stage was much higher than that of CHIP+/+ cells (Supplementary Figure 5b). These results indicate that CHIP attenuates MNNG-induced cell death, likely by ubiquitination and destabilization of tAIF. To confirm whether MNNG induces caspase-independent cell death, CHIP+/+ and CHIP−/− cells were incubated with and without MNNG or staurosporine. Immunoblot analysis revealed that staurosporine, but not MNNG, caused a marked increase in the levels of cleaved caspase-3, which is a typical phenotype of caspase-dependent cell death (Supplementary Figure 8). These results confirm that MNNG induces caspase-independent cell death.
MNNG induces downregulation of CHIP level
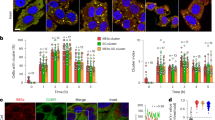
As USP2 accelerates tAIF-mediated cell death and CHIP attenuates it by their antagonistic effects on tAIF stability, cells must operate a mechanism that shifts the balance toward tAIF stabilization for executing cell death when exposed to stresses that induce the release of tAIF. In an attempt to explore the mechanism, we examined whether the level of endogenous CHIP and/or USP2 in CHIP+/+ cells might change upon MNNG treatment. Remarkably, the level of CHIP was sharply declined from about 6 h after MNNG treatment, whereas that of USP2 remained relatively constant (Figure 7a). Nearly the same results were obtained in three independent experiments (Figure 7b). Furthermore, the dramatic decrease in CHIP level occurred at the same time (i.e., at 8 h after MNNG treatment) when cell shrinkage and chromatin condensation were evident in CHIP+/+ cells (see Figure 6b). These results suggest that MNNG downregulates the cellular level of CHIP, although through unknown mechanism(s). Figure 7c shows a model for the antagonistic functions of CHIP and USP2 in the control of AIF-mediated cell death.
MNNG induces downregulation of CHIP level. (a) CHIP+/+ cells were treated with and without MNNG for 15 min, washed, and incubated for the indicated periods. Cell lysates were subjected to immunoblot with anti-CHIP, USP2, or β-actin antibody. (b) Experiments were performed as in (a), and the band intensities seen at the 0-time points were expressed as 1.0 and the others were as its relative values. The error bars indicate the mean±S.D. of three independent experiments. (c) Schematic diagram shows a model for antagonistic functions of CHIP and USP2 in the control of tAIF-mediated cell death
Discussion
Lines of evidence presented in this study indicate that the antagonistic actions of CHIP and USP2 toward tAIF have a critical role in the control of AIF-mediated cell death. First, USP2 could deubiquitinate tAIF and thus prevented its degradation by the proteasome. In contrast, CHIP served as an ubiquitin E3 ligase for tAIF and thereby promoted its degradation. Second, overexpression of USP2, but not C276A, promoted MNNG-induced cell death, as evidenced by a dramatic increase in cell shrinkage and chromatin condensation. On the other hand, knockdown of USP2 attenuated MNNG-induced cell death. Third, tAIF-mediated cell death could be prevented by coexpression of CHIP, but not by that of CHIPΔU. Finally, CHIP−/− cells were much more sensitive to MNNG-induced cell death than CHIP+/+ cells. Furthermore, MNNG-induced cell shrinkage and chromatin condensation occurred about the same time when CHIP level in CHIP+/+ cells was markedly decreased. These findings implicate critical roles of USP2 and CHIP in the regulation of MNNG-induced cell death by controlling tAIF stability. Unfortunately, however, we were unable to determine the precise half-life of endogenous or ectopically expressed tAIF, because treatment with cycloheximide to cells that had been exposed to MNNG or overexpressed with tAIF dramatically decreased the cell number because of their combined toxicity.
When cells encounter genotoxic stresses, it is important for cells to decide whether to kill themselves or operate ways to survive. On the basis of our present findings with previous reports,9, 10 we propose a mechanism on how cells make a decision (Figure 7c). Under mild stress conditions, tAIF released from the mitochondria would be ubiquitinated and degraded by CHIP, thus allowing cells to repair DNA damage for their survival. However, when a genotoxic stress prolongs or exceeds the ability of cells to repair the damage, USP2 would deubiquitinate and stabilize tAIF as CHIP level falls, thus allowing tAIF to accumulate and translocate into the nucleus for the execution of caspase-independent cell death.
HSP70 binds to AIF in the cytosol and inhibits the translocation of AIF to the nucleus.30 As HSP70 and CHIP are known to interact with each other and cooperate in the degradation of unfolded proteins as well as for their refolding,34, 35 it is possible that HSP70 facilitates CHIP-mediated degradation of tAIF by securing tAIF in the cytosol. However, CHIP is capable of binding to and ubiquitinating tAIF in the absence of HSP70 under in vitro conditions. Moreover, CHIP−/− cells are much more sensitive to MNNG for the induction of cell death than CHIP+/+ cells. Thus, it seems likely that HSP70 has an auxiliary role in the control of tAIF-mediated cell death.
AIF has been regarded as a main mediator of caspase-independent cell death in cerebral and cardiac ischemic injuries as well as in degenerative disorders, including amyotrophic lateral sclerosis and Alzheimer’s and Parkinson’s diseases.36 AIF-mediated cell death has also been implicated in the control of unregulated cell-proliferation disorders, such as cancer. Therefore, the factors that directly or indirectly induce the release of tAIF from the mitochondria (e.g., PARP1, calpain, and cathepsin B) have been targeted for the development of therapeutic drugs. For example, BZL101, a plant drug from Scutellaria barbate, which specifically provokes AIF translocation from the mitochondria to the nucleus and thereby induces cell death, has been shown to exert an efficient anti-tumor activity in patients with advanced breast cancer in phase I trial.37 On the other hand, treatment with calpain-specific inhibitors, which block the release of tAIF from the mitochondria, has been shown to significantly reduce the infarct size in rat ischemia models.38 In these regards, USP2 and CHIP could also serve as specific targets for the development of therapeutic drugs: while the inhibitors that block the USP2 activity can preclude excessive cell death observed in ischemic or degenerative processes, CHIP-specific inhibitors may attenuate cancer development by accelerating cell death.
Materials and Methods
Plasmids, antibodies, and recombinant proteins
Human USP2 cDNA was amplified from its coding sequence by PCR using a 5′ primer containing KpnI site and a 3′ primer containing SalI site. The PCR fragment was ligated into the same restriction sites of pFlag-CMV-2 (Sigma-Aldrich, St. Louis, MO, USA). To generate an inactive USP2 mutant, the active site residue Cys276 was substituted with Ala by site-directed mutagenesis. The pcDNA3-AIF encoding rat AIF was obtained from Dr. G. Kroemer (INSERM, Paris, France). The cDNA sequence that corresponds to tAIF was amplified and inserted into pcDNA4.1-V5-His/TOPO (Invitrogen, Carlsbad, CA, USA). The cDNAs for CHIP and CHIPΔU were inserted into pcDNA3-Myc. For bacterial expression, the coding sequences for tAIF, CHIP, and CHIPΔU were inserted to pGEX4T1.
Antibodies used were as follows: mouse monoclonal anti-Flag M2 antibody (Sigma), mouse monoclonal anti-HA antibody (Roche, Mannheim, Germany), mouse monoclonal anti-Xpress antibody (Invitrogen), mouse monoclonal anti-AIF E-1, rabbit polyclonal anti-AIF H-300, rabbit polyclonal anti-GAPDH antibody (Santa Cruz, Santa Cruz, CA, USA), rabbit polyclonal anti-USP2 (Abcam, Cambridge, MA, USA), anti-cleaved caspase-3 antibody (Cell Signaling, Danvers, MA, USA), horseradish peroxidase-conjugated anti-rabbit IgG, anti-mouse IgG, anti-mouse conjugated FITC, and anti-rabbit conjugated TRITC (Jackson ImmunoResearch, Baltimore, MD, USA).
GST-tAIF, GST-CHIP, and GST-USP2 were overproduced in BL21-Codon Plus Bacteria (Stratagene, La Jolla, CA, USA). Each protein was then purified by using glutathione (GSH)-Sepharose column. To also obtain the proteins without GST tag, each of GST-fused-proteins bound to GSH-Sepharose was incubated overnight with 200 units/ml of thrombin at 4°C. The cleaved proteins were then eluted with PBS containing 0.1 mM PMSF. His-UbcH5A and His-Uba1 were overproduced in BL21 cells and Sf9 insect cells using baculovirus system, respectively. They were then purified by using Ni2+-NTA agarose columns (Qiagen, Hilden, Germany).
Mass spectrometry
The USP2-associated complexes were digested with Achromobacter protease I and the resulting peptides were analyzed using a nanoscale LC–MS/MS system as described previously.39 The peptide mixture was applied to a Mightysil-PR-18 (1 μm particle, Kanto Chemical, Tokyo, Japan) frit-less column (45 mm × 0.150 mm ID) and separated using a 0–40% gradient of acetonitrile containing 0.1% formic acid over 30 min at a flow rate of 50 nl/min. Eluted peptides were sprayed directly into a quadropole time-of-flight hybrid mass spectrometer (Q-Tof Ultima, Micromass, Manchester, UK). MS and MS/MS spectra were obtained in a data-dependent mode. Up to four precursor ions above an intensity threshold of 10 counts/s were selected for MS/MS analyses from each survey scan. All MS/MS spectra were searched against protein sequences of Swiss Prot and RefSeq (NCBI) using batch processes of Mascot software package (Matrix Science, London, UK). The criteria for match acceptance were the following: (1) When the match score was 10 over each threshold, identification was accepted without further consideration, (2) when the difference of score and threshold was lower than 10, or when proteins were identified on the basis of a single matched MS/MS spectrum, we manually confirmed the raw data before acceptance, (3) peptides assigned by less than three y series ions and peptides with +4 charge state were all eliminated regardless of their scores.
Cell culture, transfection, and immunoprecipitation
HEK293T and HeLa cells were cultured at 37°C in a 5% CO2 atmosphere in DMEM supplemented with 10% FBS, 2 mM L-glutamine, and 25 U/ml of penicillin and streptomycin. Transfections of plasmids were performed using Metafectene reagent (Biontex, Planegg-Martinsried, Germany). MEF cells were transfected with plasmids by electroporation at 220 V, 950 μF. After transfection, cells were further cultured for 24–48 h. Total amounts of transfected DNA were kept equal by supplementing empty vectors. To induce genotoxic stress, cells were treated 0.5 mM MNNG, washed with PBS, and incubated in DMEM containing 10% FBS. For immunoprecipitation, cell lysates were prepared in 50 mM Tris-HCl (pH 7.4) containing 150 mM NaCl, 1 mM EDTA, 1 mM NEM, 1 mM sodium vanadate, 1 mM NaF, 1 mM PMSF, and 1 × protease inhibitor cocktail (Roche) in the presence or absence of 0.5% (v/v) Triton X-100. Cell lysates were incubated with appropriate antibodies for 1 h at 4°C and then with protein A-conjugated Sepharose for the next 1 h.
Subcellular fractionation
HeLa cells were disrupted by using Dounce homogenizer40 or by treatment with 0.025% digitonin,15 followed by centrifugation. The resulting supernatants were used as the cytosolic fraction, and the pellets were as the mixture of the mitochondrial and nuclear fractions.
Ubiquitination assays
For in vivo assay, HA-Ub was expressed in HEK293T cells with tAIF-V5-His. Cell lysates were prepared in 50 mM NaH2PO4/Na2HPO4 buffer (pH 8.0) containing 150 mM NaCl, 0.2% Tween-20, 1% SDS, 1 mM EDTA, 1 mM NEM, 1 mM sodium vanadate, 1 mM NaF, 1 mM PMSF, and 1 × protease inhibitor cocktail. After incubating them with NTA resins at 4°C for 2 h, precipitates were washed with the same buffer and subjected to immunoblot analysis. For in vitro assay, His-Uba1, His-UbcH5A, and tAIF were incubated with CHIP or CHIPΔU in 50 mM Tris-HCl (pH 7.4), 5 mM ATP, 1 mM DTT, and 5 mM MgCl2. After incubation at 37°C for 1 h, the samples were subjected to immunoblot analysis.
Immunocytochemistry
HeLa cells grown on cover-slips were transfected with appropriate plasmids. After culturing appropriate periods, cells were fixed by 3.7% paraformaldehyde in PBS for 10 min. They were washed three times with 0.1% Triton X-100, and permeabilized with 0.5% Triton X-100 for 5 min. They were blocked with 0.1% Triton, 10% goat serum, 1% BSA, and 1% gelatin. Cells were incubated for 1 h with appropriate antibodies in 0.1% Triton X-100 and 3% BSA. After washing, cells were incubated for 1 h with FITC-labeled goat anti-mouse IgG and TRITC-labeled goat anti-rabbit IgG in 0.1% Triton X-100 and 3% BSA. Cells were then observed under a confocal laser scanning microscope (Carl Zeiss-LSM510, Jena, Germany). Images were processed using Photoshop (Adobe). Cells were also observed under a fluorescence microscope (Axioplan II, Zeiss). Fluorescence images were captured using a cooled charged coupled-devised (CCD) digital camera (Axio-Cam, Zeiss).
Abbreviations
- AIF:
-
apoptosis inducing factor
- CHIP:
-
C-terminus of Hsp70-interacting protein
- FITC:
-
fluorescein isothiocyanate
- GSH:
-
glutathione
- GST:
-
glutathione S-transferase
- MNNG:
-
N-methyl-N′-nitro-N′-nitrosoguanidine
- NEM:
-
N-ethylmaleimide
- NTA:
-
nitrilotriacetic acid
- PARP:
-
poly(ADP-ribose) polymerase-1
- PMSF:
-
phenylmethylsulfonyl fluoride
- TRITC:
-
tetramethylrhodamine isothiocyanate
- USP2:
-
ubiquitin-specific protease-2
References
Daugas E, Nochy D, Ravagnan L, Loeffler M, Susin SA, Zamzami N et al. Apoptosis-inducing factor (AIF): a ubiquitous mitochondrial oxidoreductase involved in apoptosis. FEBS Lett 2000; 476: 118–123.
Klein JA, Longo-Guess CM, Rossmann MP, Seburn KL, Hurd RE, Frankel WN et al. The harlequin mouse mutation downregulates apoptosis-inducing factor. Nature 2002; 419: 367–374.
Brown D, Yu BD, Joza N, Benit P, Meneses J, Firpo M et al. Loss of Aif function causes cell death in the mouse embryo, but the temporal progression of patterning is normal. Proc Natl Acad Sci USA 2006; 103: 9918–9923.
Joza N, Galindo K, Pospisilik JA, Benit P, Rangachari M, Kanitz EE et al. The molecular archaeology of a mitochondrial death effector: AIF in Drosophila. Cell Death Differ 2008; 15: 1009–1018.
Joza N, Oudit GY, Brown D, Benit P, Kassiri Z, Vahsen N et al. Muscle-specific loss of apoptosis-inducing factor leads to mitochondrial dysfunction, skeletal muscle atrophy, and dilated cardiomyopathy. Mol Cell Biol 2005; 25: 10261–10272.
Pospisilik JA, Knauf C, Joza N, Benit P, Orthofer M, Cani PD et al. Targeted deletion of AIF decreases mitochondrial oxidative phosphorylation and protects from obesity and diabetes. Cell 2007; 131: 476–491.
Hangen E, Blomgren K, Benit P, Kroemer G, Modjtahedi N . Life with or without AIF. Trends Biochem Sci 2010; 35: 278–287.
Boujrad H, Gubkina O, Robert N, Krantic S, Susin SA . AIF-mediated programmed necrosis: a highly regulated way to die. Cell Cycle 2007; 6: 2612–2619.
Susin SA, Lorenzo HK, Zamzami N, Marzo I, Snow BE, Brothers GM et al. Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 1999; 397: 441–446.
Yu S-W, Wang H, Poitras MF, Coombs C, Bowers WJ, Federoff HJ et al. Mediation of poly(ADP-ribose) polymerase-1-dependent cell death by apoptosis-inducing factor. Science 2002; 297: 259–263.
Wesierska-Gadek J, Gueorguieva M, Schloffer D, Uhl M, Wojciechowski J . Non-apoptogenic killing of hela cervical carcinoma cells after short exposure to the alkylating agent N-methyl-N’-nitro-N-nitrosoguanidine (MNNG). J Cell Biochem 2003; 89: 1222–1234.
Otera H, Ohsakaya S, Nagaura ZI, Ishihara N, Mihara K . Export of mitochondrial AIF in response to proapoptotic stimuli depends on processing at the intermembrane space. EMBO J 2005; 24: 1375–1386.
Artus C, Boujrad H, Bouharrour A, Brunelle MN, Hoos S, Yuste VJ et al. AIF promotes chromatinolysis and caspase-independent programmed necrosis by interacting with histone H2AX. EMBO J 2010; 29: 1585–1599.
Yuste VJ, Moubarak RS, Delettre C, Bras M, Sancho P, Robert N et al. Cysteine protease inhibition prevents mitochondrial apoptosis-inducing factor (AIF) release. Cell Death Differ 2005; 12: 1445–1448.
Moubarak RS, Yuste VJ, Artus C, Bouharrour A, Greer PA, Menissier-de Murcia J et al. Sequential activation of poly(ADP-ribose) polymerase 1, calpains, and Bax is essential in apoptosis-inducing factor-mediated programmed necrosis. Mol Cell Biol 2007; 27: 4844–4862.
Wang Y, Kim NS, Li X, Greer PA, Koehler RC, Dawson VL et al. Calpain activation is not required for AIF translocation in PARP-1-dependent cell death (parthanatos). J Neurochem 2009; 110: 687–696.
Hangen E, De Zio D, Bordi M, Zhu C, Dessen P, Caffin F et al. A brain-specific isoform of mitochondrial apoptosis-inducing factor: AIF2. Cell Death Differ 2010; 17: 1155–1166.
Hershko A, Ciechanover A . The ubiquitin system. Annu Rev Biochem 1998; 67: 425–479.
Hochstrasser M . Ubiquitin-dependent protein degradation. Annu Rev Genet 1996; 30: 405–439.
Kim JH, Park KC, Chung SS, Bang O, Chung CH . Deubiquitinating enzymes as cellular regulators. J Biochem 2003; 134: 9–18.
Wilkinson KD . Regulation of ubiquitin-dependent processes by deubiquitinating enzymes. FASEB J 1997; 11: 1245–1256.
Murata S, Minami Y, Minami M, Chiba T, Tanaka K . CHIP is a chaperone-dependent E3 ligase that ubiquitylates unfolded protein. EMBO Rep 2001; 2: 1133–1138.
Cyr DM, Höhfeld J, Patterson C . Protein quality control: U-box-containing E3 ubiquitin ligases join the fold. Trends Biochem Sci 2002; 27: 368–375.
Kampinga HH, Kanon B, Salomons FA, Kabakov AE, Patterson C . Overexpression of the cochaperone CHIP enhances Hsp70-dependent folding activity in mammalian cells. Mol Cell Biol 2003; 23: 4948–4958.
Park KC, Kim JH, Choi EJ, Min SW, Rhee S, Baek SH et al. Antagonistic regulation of myogenesis by two deubiquitinating enzymes, UBP45 and UBP69. Proc Natl Acad Sci USA 2002; 99: 9733–9738.
Baek SH, Choi KS, Yoo YJ, Cho JM, Baker RT, Tanaka K et al. Molecular cloning of a novel ubiquitin-specific protease, UBP41, with isopeptidase activity in chick skeletal muscle. J Biol Chem 1997; 272: 25560–25565.
Stevenson LF, Sparks A, Allende-Vega N, Xirodimas DP, Lane DP, Saville MK . The deubiquitinating enzyme USP2a regulates the p53 pathway by targeting Mdm2. EMBO J 2007; 26: 976–986.
Graner E, Tang D, Rossi S, Baron A, Migita T, Weinstein LJ et al. The isopeptidase USP2a regulates the stability of fatty acid synthase in prostate cancer. Cancer Cell 2004; 5: 253–261.
Gewies A, Grimm S . UBP41 is a proapoptotic ubiquitin-specific protease. Cancer Res 2003; 63: 682–688.
Ravagnan L, Gurbuxani S, Susin SA, Maisse C, Daugas E, Zamzami N et al. Heat-shock protein 70 antagonizes apoptosis-inducing factor. Nat Cell Biol 2001; 3: 839–843.
Jiang J, Ballinger CA, Wu Y, Dai Q, Cyr DM, Hohfeld J et al. CHIP is a U-box-dependent E3 ubiquitin ligase: identification of Hsc70 as a target for ubiquitylation. J Biol Chem 2001; 276: 42938–42944.
Sahara N, Murayama M, Mizoroki T, Urushitani M, Imai Y, Takahashi R et al. In vivo evidence of CHIP up-regulation attenuating tau aggregation. J Neurochem 2005; 94: 1254–1263.
Zhang M, Windheim M, Roe SM, Peggie M, Cohen P, Prodromou C et al. Chaperoned ubiquitylation—crystal structures of the CHIP U box E3 ubiquitin ligase and a CHIP-Ubc13-Uev1a complex. Mol Cell 2005; 20: 525–538.
Qian SB, McDonough H, Boellmann F, Cyr DM, Patterson C . CHIP-mediated stress recovery by sequential ubiquitination of substrates and Hsp70. Nature 2006; 440: 551–555.
Shin Y, Klucken J, Patterson C, Hyman BT, McLean PJ . The co-chaperone carboxyl terminus of Hsp70-interacting protein (CHIP) mediates alpha-synuclein degradation decisions between proteasomal and lysosomal pathways. J Biol Chem 2005; 280: 23727–23734.
Lorenzo HK, Susin SA . Therapeutic potential of AIF-mediated caspase-independent programmed cell death. Drug Resist Updat 2007; 10: 235–255.
Rugo H, Shtivelman E, Perez A, Vogel C, Franco S, Tan Chiu E et al. Phase I trial and antitumor effects of BZL101 for patients with advanced breast cancer. Breast Cancer Res Treat 2007; 105: 17–28.
Hong SC, Goto Y, Lanzino G, Soleau S, Kassell NF, Lee KS . Neuroprotection with a calpain inhibitor in a model of focal cerebral ischemia. Stroke 1994; 25: 663–669.
Natsume T, Yamauchi Y, Nakayama H, Shinkawa T, Yanagida M, Takahashi N et al. A direct nanoflow liquid chromatography—tandem mass spectrometry system for interaction proteomics. Anal Chem 2002; 74: 4725–4733.
Robertson JD, Enoksson M, Suomela M, Zhivotovsky B, Orrenius S . Caspase-2 acts upstream of mitochondria to promote cytochrome c release during etoposide-induced apoptois. J Biol Chem 2002; 277: 29803–29809.
Acknowledgements
We thank Dr. G Kroemer for providing pcDNA3-AIF. We also thank Ms. SH Ka for her excellent technical assistance. This work was supported by grants from KRF (KRF-2005-084-C00025) and KOSEF (M10533010001-05N3301). KH Oh was the recipient of TJ Park postdoctoral fellowship. SW Yang and JM Park were the recipients of BK21 fellowship.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by P Vandenabeele
Supplementary Information accompanies the paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Oh, K., Yang, S., Park, J. et al. Control of AIF-mediated cell death by antagonistic functions of CHIP ubiquitin E3 ligase and USP2 deubiquitinating enzyme. Cell Death Differ 18, 1326–1336 (2011). https://doi.org/10.1038/cdd.2011.3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2011.3
Keywords
This article is cited by
-
Ubiquitin ligase CHIP regulates OTUD3 stability and suppresses tumour metastasis in lung cancer
Cell Death & Differentiation (2020)
-
Observation of Parthanatos Involvement in Diminished Ovarian Reserve Patients and Melatonin’s Protective Function Through Inhibiting ADP-Ribose (PAR) Expression and Preventing AIF Translocation into the Nucleus
Reproductive Sciences (2020)
-
Multifaceted C-terminus of HSP70-interacting protein regulates tumorigenesis via protein quality control
Archives of Pharmacal Research (2019)
-
A novel function of cIAP1 as a mediator of CHIP-driven eIF4E regulation
Scientific Reports (2017)
-
Phosphorylation of CHIP at Ser20 by Cdk5 promotes tAIF-mediated neuronal death
Cell Death & Differentiation (2016)