Abstract
Genetic studies of the nematode Caenorhabditis elegans have uncovered four genes, egl-1 (BH3 only), ced-9 (Bcl-2 related), ced-4 (apoptosis protease activating factor-1), and ced-3 (caspase), which function in a linear pathway to promote developmental cell death in this organism. While this core pathway functions in many cells, recent studies suggest that additional regulators, acting on or in lieu of these core genes, can promote or inhibit the onset of cell death. Here, we discuss the evidence for these noncanonical mechanisms of C. elegans cell death control. We consider novel modes for regulating the core apoptosis genes, and describe a newly identified cell death pathway independent of all known C. elegans cell death genes. The existence of these noncanonical cell death programs suggests that organisms have evolved multiple ways to ensure appropriate cellular demise during development.
Similar content being viewed by others
Main
Cell death is an important process in metazoan development, and coordinated control of cell division, differentiation, and death governs many aspects of morphogenesis. Often, waves of cell proliferation are followed by cell death, which serves to sculpt and delete structures, adjust cell numbers, and eliminate aberrantly produced cells.1, 2, 3, 4 Animals defective in cell death can display severe developmental abnormalities and may not survive to adulthood.5, 6, 7
Studies of the nematode Caenorhabditis elegans provided initial evidence that developmental cell death has an underlying genetic basis.8 A transparent cuticle and nearly invariant, lineage-restricted development have allowed the construction of cell death maps describing which cells die, when they die, and where they die during C. elegans development. These maps, coupled with genetic tractability, have made C. elegans a powerful system in which to study cell death. In the developing C. elegans hermaphrodite, two waves of somatic cell death occur: 113 cells die during embryonic development, and an additional 18 somatic cells die in early postembryonic stages.9, 10 A third wave of cell death occurs in the adult hermaphrodite germ line, where roughly half of developing germ cells die.11
Unlike dying cells in many other organisms, most somatic cells fated to die in C. elegans are not differentiated and succumb less than 30 min after they are born. Four cells fated to die in the hermaphrodite, the cephalic companion neurons (CEM) neurons, live for nearly 3 h before dying. In addition, three other cells fated to die, MS.pppaaa, the sister cell of the Z1 germ-line precursor cell, the tail-spike cell, and the male-specific linker cell, acquire differentiated features, and survive for six, five, and 30 h, respectively, before they die.9, 10 Although the death of the MS.pppaaa cell has not been examined in detail, the CEM neurons, the tail-spike cell, and the linker cell have been shown to employ noncanonical cell death programs (see below), suggesting that elimination of longer-lived, differentiated cells may require special cell death mechanisms. Here, we review the programs leading to the demise of these, and of other C. elegans cells exhibiting alternative modes of cell death regulation.
The Canonical C. elegans Cell Death Program
Four core genes that mediate cell death during C. elegans development have been described. Loss-of-function (lf) mutations in egl-1, ced-4, and ced-3 result in the survival of many cells destined to die, and overexpression of these genes promotes cell death. These results indicate that egl-1, ced-4, and ced-3 function normally to promote cellular demise.8, 12, 13, 14, 15 By contrast, ced-9(lf) mutations result in excessive cell death and organismal lethality, and a ced-9 gain-of-function (gf) mutation blocks cell death, suggesting that ced-9 functions normally to protect against cell death.16, 17 Epistasis studies between these genes suggest a linear pathway of action, in which egl-1 inhibits ced-9,12 which, in turn, inhibits ced-4, preventing activation of ced-315 (Figure 1).
Noncanonical mechanisms for cell death initiation in C. elegans. Execution of the canonical cell death program (black letters) is thought to be initiated by activation of egl-1, which, in turn, inhibits ced-9, allowing ced-4 to activate ced-3. Interaction of noncanonical regulators with this core cell death pathway has been documented in the following settings: (1) germ cell death can be prevented by pax-2, egl-38, and lin-35 transcriptional regulation of ced-9 or stimulated by dpl-1 transcriptional regulation of ced-4 and ced-3. In addition, RAS/MAPK may promote germ cell death through an unknown mechanism. (2) Tail-spike cell death is initiated by pal-1 transcriptional upregulation of ced-3, and still requires ced-4. (3) The sexually dimorphic survival of the cephalic companion neurons (CEM) neurons is regulated by the sex determination pathway (tra-1) through transcriptional activation of ceh-30. The mechanism of ceh-30's anti-apoptotic activity is unknown
Molecular characterization of the core cell death genes and their subsequent biochemical study revealed that the genetic interactions observed between these genes reflect physical-binding events that ultimately control the activation of CED-3, a cysteine protease of the caspase family.18 In the absence of apoptotic stimuli, CED-9, an anti-apoptotic member of the B-cell lymphoma/leukemia-2 protein family, binds to and sequesters CED-4 (apoptosis protease activating factor-1) to the outer mitochondrial membrane.19, 20, 21 In cells destined to die, EGL-1, a Bcl-2 homology domain 3 (BH3)-only protein, binds to CED-9, disrupting its interaction with CED-4.12, 22, 23 CED-4 is then free to promote CED-3 activation and subsequent cellular destruction.19, 20, 24, 25
The linear nature of the canonical C. elegans cell death pathway has led to the hypothesis that specification of which cells live and which cells die during development is accomplished by controlling the levels of EGL-1. Thus, it has been proposed that in cells destined to live, EGL-1 activity is low, and that in cells destined to die, EGL-1 activity is high. Indeed, studies of a small number of cells fated to die have shown that expression of egl-1 is under the control of known developmental transcription factors.26, 27, 28, 29, 30, 31 However, careful examination of the genetic interactions between C. elegans cell death genes suggests that the story is not so simple. If the linear model is correct, in double mutant animals containing both ced-9(lf) and ced-3(weak lf) alleles, the status of EGL-1 should be irrelevant, because its target, CED-9, is absent. However, many cells still die appropriately in these animals, and few, if any, ectopic cell deaths are observed17 (S Shaham unpublished data). These results suggest either that EGL-1 can target proteins other than CED-9, or, perhaps more likely, that other cell death initiation mechanisms independent of EGL-1 must exist. As we review here, a number of EGL-1-independent regulatory interactions have been described.
EGL-1-Independent Cell Death in the C. elegans Germ Line
In adult C. elegans hermaphrodites, roughly 50% of developing germ cells undergo programmed cell death. Characterization of the underlying death program in these cells has revealed noncanonical initiation of the process.11
The distal ends of the hermaphrodite gonad arms are capped by a distal tip cell (DTC) that maintains germ cells in a mitotic state.32, 33 As they move away from the DTCs, germ cells enter meiosis, and arrest at the pachytene stage of meiosis I. Activation of the RAS/mitogen-activated protein kinase (MAPK) signaling pathway is then required for the germ cells to exit pachytene.34 It is also at this stage of development that germ cells undergo programmed cell death. As they die, they are rapidly cellularized away from the germ-line syncytium, and are engulfed by gonadal sheath cells.11 Strong loss-of-function mutations in ced-3 or ced-4 prevent nearly all germ cell deaths, as they do in the soma. Conversely, CED-9 functions to protect germ cells against death. In animals, homozygous for a temperature-sensitive ced-9(lf) allele, the number of cell deaths observed in the germ line is nearly three times than that observed in wild-type animals. Surprisingly, however, neither egl-1(lf) mutations nor a ced-9(gf) mutation, reported to disrupt binding to EGL-1,23, 35 affects germ cell death. Thus, in the germ line, egl-1 cannot serve to integrate signals that control cell death onset.
How is cell death initiation regulated in germ cells? One possibility involves the RAS/MAPK pathway, which regulates meiotic entry of germ cells (Figure 1). Gumienny11 demonstrated that loss-of-function mutations in components of this pathway indeed block the accumulation of dying germ cells. However, gain-of-function mutations in the pathway fail to increase the number of dying cells, suggesting that the RAS/MAPK pathway establishes the competency of germ cells to die, but additional genes must also interact with the cell death machinery.
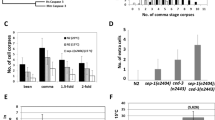
Studies of the Pax2/5/8-related proteins EGL-38 and PAX-2 suggest that these transcriptional regulators may directly control the activity of the cell death machinery in the germ line.36 Genetically reducing the levels of either EGL-38 or PAX-2 proteins, using loss-of-function mutations in these genes, increases the number of dying germ cells, consistent with the idea that egl-38 and pax-2 normally function to protect against cell death. egl-38(lf); pax-2(lf) double mutants display additive accumulation of dying germ cells, suggesting that they may function in parallel. Overexpressing, either egl-38 or pax-2, using a heat-shock promoter reduces the number of dying germ cells and does so only in animals in which ced-9 is intact. These results are consistent with a model in which egl-38 and pax-2 function genetically upstream of ced-9 to promote ced-9 function. Supporting this hypothesis, overexpressing CED-9 using a heat-shock promoter in egl-38(lf); pax-2(lf) animals, suppresses the increase in germ cell death. Using real-time reverse transcriptase (RT)–PCR, Park et al.36 demonstrated that ced-9 transcript levels were decreased in egl-38(lf); pax-2(lf) animals, and increased in response to induced expression of EGL-38 and PAX-2. Furthermore, using chromatin immunoprecipitation, these authors showed that EGL-38 and PAX-2 physically bind to the ced-9 promoter. These results suggest that control of ced-9 transcription may provide an avenue for the noncanonical regulation of caspase-dependent cell death in the C. elegans germ line (Figure 1).
In cultured mammalian cells, Pax5 can interact with the RB protein,37, 38 and a recent study reported that the RB complex controls somatic cell death in C. elegans,39 suggesting that similar interactions might occur in this organism as well. Indeed, Schertel and Conradt40 demonstrated that loss of lin-35/Rb also blocks cell death in the C. elegans germ line. Furthermore, using quantitative real-time PCR, these authors demonstrated that in lin-35 loss-of-function mutants, ced-9 transcript in the germ line increases fivefold. These results suggest perhaps that LIN-35 interacts with EGL-38 and/or PAX-2 to control ced-9 mRNA levels; however, this model has not been directly tested. Additional components of the RB complex were also suggested to regulate mRNA levels of ced-3 and ced-4 independently of lin-35/RB40; however, in mutants of these components, only a twofold reduction in ced-3/4 RNA was detected, suggesting that these components have a more minor role in controlling germ-line cell death.
It is important to note that correlations between the levels of ced-9 transcript and of EGL-38, PAX-2, and LIN-35 proteins in wild-type germ cells have not been examined, leaving open the possibility that these proteins have only permissive roles in regulating cell death initiation.
Transcriptional Control Downstream of egl-1 and ced-9 in the CEM Neurons
The idea that transcriptional control of cell death genes other than egl-1 could be used as a mechanism for regulating cell death has also emerged from studies of the sexually dimorphic survival of the CEM neurons.41 Death of the CEM neurons in wild-type hermaphrodites can be blocked by loss-of-function mutations in egl-1, ced-4, or ced-3, or by a gain-of-function mutation in ced-9, indicating that the canonical apoptotic machinery mediates this specific cell death.41, 42 However, survival of the CEMs in wild-type males may not be mediated by ced-9, since in ced-9(lf); ced-3(weak lf) males most CEM neurons still survive,41 unlike other cells destined to die.
Recent studies have demonstrated that the homeodomain transcription factor CEH-30 and the zinc-finger transcriptional regulator TRA-1A are important in the decision of CEM neurons to live or die. Schwartz and Horvitz41 and Peden et al.42 showed that loss-of-function mutations in ceh-30 promote the death of male CEM neurons, suggesting that ceh-30 normally inhibits CEM death. This idea is further supported by the demonstration that a gain-of-function mutation in ceh-30 causes CEM neurons to inappropriately survive in hermaphrodites.41 The ceh-30(gf) mutation alters an intronic consensus sequence for binding of TRA-1A, a regulator of sexual differentiation expressed only in hermaphrodites.26, 43 Thus, in ceh-30(gf) hermaphrodites, weakened TRA-1A binding mimics the low wild-type levels of TRA-1A in males, thereby allowing CEH-30 to exert its anti-apoptotic influence to promote CEM survival.
In the presence of CEH-30, the CEM neurons survive even in the absence of CED-9,41 indicating that CEH-30 protects male CEM neurons independently of CED-9, suggesting that control of EGL-1 activity in this context cannot mediate cell survival (Figure 1). How CEH-30 functions, and what its targets are is still not known; however, defects in the mammalian protein most related to CEH-30, Barhl1, also promote increased sensory neuron cell death,44, 45 suggesting that the protective effect of this class of transcriptional regulators may be conserved.
EGL-1/CED-9-Independent Cell Death of the C. elegans Tail-Spike Cell
Studies of the death of the C. elegans tail-spike cell have also revealed that cell death initiation can be mediated by noncanonical transcriptional regulation. These studies also reveal that tail-spike cell death deviates even further from the canonical cell death pathway.46
During embryogenesis, the tail-spike cell dies 5 h after its birth, displaying differentiated features including an extended filamentous process that may serve to guide morphogenesis of neighboring hypodermal cells.10 Tail-spike cell death is only partially dependent on egl-1: whereas strong loss-of-function mutations in ced-3 or ced-4 completely block tail-spike cell death, strong loss-of-function mutations in egl-1 only block tail-spike cell death in 30% of animals.46 Surprisingly, tail-spike cell death is also partially independent of ced-9: gain-of-function mutations in ced-9, which prevent most other somatic cell deaths, have virtually no effect on tail-spike cell death. Furthermore, a previously described pro-apoptotic function of ced-917 is entirely absent in the tail-spike cell. Taken together, these results suggest that a novel mechanism regulates caspase activity in the tail-spike cell to promote its death. This mechanism must display remarkable temporal accuracy, since inappropriate early activation of the cell death program would be predicted to lead to defects in hypodermal cell morphogenesis.
Although the nature of the temporal regulator is unknown, Maurer et al.46 demonstrated that this regulator acts, at least in part, through transcriptional control of the ced-3 caspase. In animals carrying ced-3 promoter::green fluorescent protein (GFP) reporters, GFP expression was observed in the tail-spike cell only 20–30 min before death ensued. The same promoter sequence driving a ced-3 cDNA was able to rescue the tail-spike cell death defect in ced-3(lf) mutants. These results indicate that expression of ced-3 just prior to tail-spike cell death is sufficient to promote its death. Maurer et al.46 identified mutants in which transcriptional induction of ced-3 failed to occur, and which led to inappropriate survival of the tail-spike cell. Two such mutants carried lesions in the homeodomain transcription factor gene pal-1. The authors went on to show that PAL-1 protein can directly bind to redundant conserved elements within the ced-3 promoter, and that these elements were required specifically for tail-spike cell death but not for other cell deaths (Figure 1). Thus, PAL-1 either directly influences the onset of cell death, or acts permissively to set a point beyond which this event can occur.
The transcription of caspases has not been extensively studied in any organism, and the results of the tail-spike cell studies suggest that caspase transcription may be important in regulating developmental cell death. The mammalian homolog of PAL-1, caudal-related homeobox 2 (Cdx2), is an intestinal tumor suppressor, and mutations in Cdx2 promote intestinal and colon tumor formation.47, 48, 49 Strikingly, in the intestinal epithelium, Cdx2 protein expression is highest in the outer, most differentiated, cell population, which normally undergoes apoptotic cell death.50 It is tempting to speculate therefore that as with PAL-1 in C. elegans, Cdx2 might function to regulate caspase transcription in the intestine.
A Novel Morphologically Conserved Program Mediates Linker Cell Death
Recent work examining the death of the C. elegans male-specific linker cell has provided strong evidence for a bona fide non-apoptotic developmental cell death program in this animal.51 Strikingly, all four core apoptotic genes as well as all known engulfment mediators appear irrelevant for this cell's demise.51
The linker cell is born in the second larval stage in the central region of the animal, and leads the migration of the male gonad behind it52, 53 (Figure 2a). The path of migration terminates at the posterior end of the animal, placing the vas deferens in close apposition to the cloaca. Upon its death during the fourth larval stage (L4)-to-adult transition, the linker cell is engulfed and removed by either the U.lp or U.rp epithelial cells (henceforth abbreviated as U.l/rp cells).53 It is thought that linker cell death occurs in order for the gonad to fuse to the cloaca, thereby creating an exit channel for sperm. Indeed, at least some mutants that block linker cell death accumulate sperm within the male gonad, presumably leading to sterility (ES Blum and S Shaham, unpublished data).
Linker cell death in C. elegans. (a) Linker cell migration in adult males. The linker cell is born in second larval stage (L2) animals and begins migrating anteriorly (i). By early L4, the linker cell is migrating posteriorly on the dorsal side (ii). By late L4, the linker cell has reached the cloaca, begins to die, and is engulfed by the U.l/rp cells (iii). In adult males, the linker cell has died and has been removed; the vas deferens is now connected to the cloaca (iv). (b) A dying and fully engulfed linker cell (arrow) in a ced-3(n717lf) male expressing a lin-48 promoter::GFP reporter in the U.l/rp engulfing cell. Arrowheads, nuclei of engulfing cell. DIC (top) and fluorescence (bottom) images. Scale bar, 5 μm. (c) Electron micrograph of a dying linker cell. Note the crenellated (indented) nucleus (white arrow), large membrane-bound vesicles (white arrowhead), and white ‘empty’ structures (red arrows), which may represent swollen endoplasmic reticulum. (d) Electron micrograph of an engulfed apoptotic cell (arrow) in C. elegans. Note the compaction of darkened cytoplasm around the nucleus
Unlike most cells that die during C. elegans development, the linker cell lives for over 30 h after it is born, and becomes highly differentiated. Anecdotal reports had suggested that linker cell death may depend on engulfment by the U.l/rp cells.53 However, when Abraham et al.51 ablated the U.l/rp grandparental precursor cell, U, the linker cell still died normally, indicating that engulfment by the U.l/rp cells is not required for linker cell death. These and other results suggest that local cues from the cloacal region may only be required to fine tune efficient destruction of the linker cell.
Temporal, rather than spatial cues, however, profoundly influence the onset of linker cell death. C. elegans possesses a developmental timing program that communicates the developmental stage of the organism to individual cells in the animal. Thus, specific cells in mutants of this timing program will execute early or delayed cell fates, depending on the nature of the timing mutant examined.54, 55 The zinc-finger transcription factor lin-29 is the most downstream known gene in the developmental timing pathway, and acts at the transition from the L4 to the adult.54, 56, 57 Strikingly, the linker cell inappropriately survives in 53% of lin-29(lf) males. Mutations in the let-7 microRNA gene, which activates lin-29 function through the lin-41 gene, also inhibit linker cell death.51, 58, 59 lin-29 is expressed in the linker cell beginning at the L3/L4 stage, and mosaic studies demonstrated that this gene functions cell autonomously within the linker cell to promote its demise. These results, together with the cell ablation and migration studies suggest that the linker cell employs a cell-autonomous program for its death. Because lin-29 is expressed in other cells not destined to die, as well as in the migrating linker cell, it is unlikely that lin-29 is sufficient to promote linker cell death. Thus, lin-29 might act together with other genes to transcriptionally control death-promoting genes.
Attempts to characterize possible targets for lin-29 among known C. elegans cell death genes revealed that linker cell death is independent of all known cell death genes, including the core apoptotic genes egl-1, ced-9, ced-4, and ced-3.51 These studies extend initial observations suggesting that linker cell death may only partially depend on ced-3 and ced-4.8 Further studies suggest that linker cell death does not require any caspase activity, because loss-of-function mutations in the other known C. elegans caspase-related genes, csp-1, -2, and -3, or mutations in both ced-3 and csp-1 did not affect linker cell death.60
Although caspase-independent death has been claimed to occur in a number of settings in vertebrate systems,61, 62, 63, 64 all such studies rely on examination of only selected caspase-deficient mutants, or on inhibitor studies. Because vertebrates contain roughly a dozen caspases, with differing substrate and inhibitor specificities, caspase dependence is very difficult to rule out, and in all these studies it is possible that caspases that have not been examined, or combinations of caspases, are relevant for the process at hand. In C. elegans, however, only four caspase-related genes exist,18, 60 and only two, ced-3 and csp-1, have enzymatic function.60 Thus, the studies of Abraham et al.51 make a strong case for the argument that caspases are not essential for linker cell death.
To examine linker cell death, Abraham et al.51 used a GFP reporter to follow linker cell fate, and it has been suggested that these reporters may in some way influence the death process. However, careful examination of animals carrying a GFP reporter transgene in the U.l/rp engulfing cell, but not in the linker cell, showed that in 33/36 ced-3(lf) mutant males and 39/39 ced-4(lf) mutant males, the linker cell died and was engulfed during the time window examined (0–2 h after the L4-to-adult transition; Figure 2b), alleviating any concerns regarding reporter interference (M Abraham and S Shaham, unpublished data).
Linker cell death is also independent of calpains and aspartyl proteases that have been shown to mediate necrotic cell death (see below),65 and it is also unlikely that the linker cell undergoes autophagic cell death. The autophagosome-tethered reporter LGG-1::GFP is not expressed at higher levels in the dying linker cell, and mutations in the autophagy genes bec-1 and unc-51, homologs of beclin and APG1, respectively, have no effect on linker cell death. Furthermore, the morphology of dying linker cells does not appear similar to autophagic cell death (Figure 2c).
Phagocytosis of the linker cell also appears to have unique genetic and cell-biological features, since engulfment of the cell by the U.l/rp cells is independent of all known engulfment genes in C. elegans.51 Specifically, neither single nor double combinations of mutations in genes of the two known cell death engulfment pathways blocks linker cell engulfment. Consistent with these genetic results, the CED-1 protein, a transmembrane protein that accumulates on engulfing cell membranes surrounding dying cells,66 was never observed around the linker cell. Although engulfment by the U.l/rp cells is not necessary for linker cell death, the role of engulfment in wild-type linker cell death has not been directly assessed, since no known mutations block its engulfment. While it appears that in wild-type animals, linker cell engulfment is mediated, at least in part, by an unknown mechanism, in him-4(lf) mutant animals, CED-1 clustering is seen around the dying linker cell. The observation that this clustering begins to occur after a linker cell corpse becomes visible suggests that the linker cell may initiate cell death independently of engulfment.
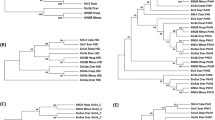
The unique genetic features of linker cell death suggest that the process is unlikely to proceed by apoptosis. Consistent with this hypothesis, electron microscopy of dying linker cells revealed non-apoptotic features, including nuclear crenellation (indentation), absence of chromatin condensation, organelle swelling, and accumulation of cytoplasmic membrane-bound structures51 (Figures 2c and d). Remarkably similar features are seen during the normal developmental death of chick ciliary ganglion cells67 and chick spinal cord motor neurons,68 a type of cell death that has been coined ‘type III’ cell death.69, 70 These observations suggest that linker cell death is morphologically conserved. Identification of genes required for linker cell death and examination of their mammalian counterparts for roles in type III cell death may help to determine whether the cell death program required to kill the linker cell is evolutionarily conserved (Figure 3).
C. elegans Mutants Displaying Noncanonical Cell Death Programs
A number of C. elegans mutant strains carrying defined genetic lesions display inappropriate cell death patterns in which cells normally destined to live instead die. In several of these mutants, cell death is mediated either by a modified version of the canonical developmental cell death pathway, or by a completely different mechanism. Although these alternate cell death programs have no known parallels during normal C. elegans development, their existence suggests that they might reflect underlying endogenous programs. We briefly describe these programs below.
Knockdown by RNA interference of the inhibitor of cell death (icd-1) gene promotes the death of many cells in the C. elegans embryo in a ced-4-dependent, yet ced-3-independent manner, and overexpression of ICD-1 can block some normally occurring embryonic cell deaths.71 ICD-1 localizes to mitochondria and is homologous to the β-subunit of the nascent-polypeptide-associated complex, which is thought to regulate protein localization during translation.71 Electron microscopy of dying cells in an icd-1(RNAi); ced-3(lf) double mutant revealed features of apoptosis, including condensed chromatin, suggesting that such features can be generated in the absence of caspases, or that a different C. elegans caspase60 substitutes for ced-3 in these dying cells.
Loss-of-function mutations in the pvl-5 gene cause incompletely penetrant embryonic lethality, and loss of sensory structures in the male tail.72 Furthermore, in animals carrying the pvl-5(ga87) mutation, some Pn.p cells, a group of precursor cells that generate hypodermal and vulval cells,9 undergo inappropriate cell death.72 Mutant animals also exhibit semi-penetrant developmental defects including embryonic lethality and gonadal migration defects. The Pn.p cell deaths can be suppressed by ced-3(lf) and ced-9(gf) mutations but not by ced-4(lf) or egl-1(lf) mutations. Similarly, a small number of embryonic deaths in pvl-5(ga87) mutants are suppressed by ced-3(lf) but not ced-4(lf) mutations. Interestingly, mutations in both known cell-corpse engulfment pathways73 also suppress Pn.p cell death.72 These results suggest that pvl-5 may normally be required to prevent ced-3-dependent cell death in a select group of cells. The molecular identity of the pvl-5 gene is not yet known.
The semi-dominant alleles of the lin-24 and lin-33 genes have also been shown to affect the survival of the Pn.p cells.74 These alleles promote ced-3- and ced-4-independent cell death, and dying cells display an abnormal morphology when viewed using Nomarski optics. Mutations in some C. elegans engulfment genes protect against lin-24 or lin-33-induced death.75
Perhaps the best characterized mutation-induced non-apoptotic cell death type in C. elegans is a necrotic-like neuronal death induced by dominant (d) gain-of-function mutations that hyperactivate the ion-channel subunits mec-476, 77 or deg-178 (degeneration inducing) of the degenerin and epithelial Na+ channel (DEG/EnaC) family.
In this necrotic death type, the cell bodies of dying neurons swell and can be readily viewed in living animals (Figure 4a). At the ultrastructural level (Figure 4b), mec-4(d)-induced neurodegeneration involves initial formation of electron-dense multilamellar figures near the plasma membrane that appear to coalesce, later intracellular vacuolation, nuclear distortion and chromatin clumping, and final degradation of intracellular contents.79 Necrotic neurons may lyse, but more often appear to be removed by phagocytosis before membrane disruption.79, 80 Interestingly, the C. elegans genes involved in phagocytotic removal of apoptotic corpses are also needed for efficient elimination of necrotic corpses, suggesting common recognition and removal mechanisms of apoptotic and necrotic cells.80
Morphological features of necrotic-like cell death in C. elegans. (a) Necrotic touch receptor neurons become dramatically swollen as neurons die. Indicated by white arrows are the two posterior PLM neurons that have become necrotic in a first larval (L1) stage animal. Note that necrosis is readily apparent in transparent living animals using low power magnification. Photo courtesy of Yury Nunez. (b) Electron micrograph showing ultrastructural features of a necrotic PVM neuron in a mec-4(d) mutant. Blue trace parallels the plasma membrane of the PLM. White arrow indicates electron-dense whorls that are prominent in dying neurons. The nucleus is outlined in red and features vacuoles and clumped chromatin. Reprinted with permission from Hall et al.79
Toxic mec-4(d) mutations encode amino-acid substitutions that increase channel conductance.77, 81, 82, 83 Genetic perturbations of other C. elegans channels, including those that reduce desensitization of the DEG-3 acetylcholine receptor84, 85 and expression of constitutively active Gαs86, 87 (thought to induce glutamate excitotoxicity), can also initiate necrosis. Mutant ion channels that are all neurotoxic can conduct calcium,83 and thus elevated calcium influx through plasma membrane ion channels appears to be a common necrosis-triggering event. In mammals, analogous glutamate excitotoxicity88 and DEG/ENaC channel hyperactivation89 are important in neuronal death consequent to stroke and ischemia.
Genetic studies suggest a general model for ion-channel-initiated necrosis in which catastrophic ER calcium release figures prominently. Null alleles of the ER Ca2+-binding chaperone calreticulin, which are important in maintenance of intracellular Ca2+ stores,90 strongly suppress both mec-4(d)-induced necrosis91 and glutamate-dependent excitotoxicity (Driscoll and Mano, unpublished). Mutations in ER Ca2+-release channels IP3-receptor ITR-1 and ryanodine receptor UNC-68 also significantly suppress death, as can dantrolene treatment, which blocks ER Ca2+ release. These studies suggest that the intracellular Ca2+ concentration must rise to critical levels for progression through necrosis and that the ER is an important source of the extreme Ca2+ elevations required for necrotic neuronal death.92 How excess ion influx through plasma membrane ion channels provokes the deleterious release of ER calcium stores remains to be elaborated.
C. elegans calcium-activated proteases (calpains TRA-3 and CLP-1) are required for efficient progression through necrosis65 and may be activated by ER calcium release. Roles for the lysosome in necrosis are underscored by the requirement for vacuolar-ATPase-mediated lysosomal acidification93 and by the existence of the intracellular anti-necrosis serine protease inhibitor serpin SRP-6, which defends against lysosomal protease release during general tissue necrosis.94 Finally, disruption of autophagy genes that impact lysosomal function can also influence the efficacy of channel-induced necrosis.95, 96
Concluding Remarks
Studies of normally occurring developmental cell death in C. elegans have been instrumental in uncovering a molecular pathway driving this process. The studies reviewed here suggest that not all components of this pathway are required to execute cell death in all cells, and that regulation of developmental cell death can occur not only by regulating the upstream BH3-only-encoding gene egl-1, but also by impinging on other, and perhaps all, components of the core apoptotic machinery. Studies of the C. elegans linker cell demonstrate in vivo roles for a non-apoptotic cell death program that can function independently of all known cell death genes. Given the conservation of the apoptotic cell death machinery across metazoans, studies of noncanonical regulation of this machinery, as well as the novel cell death pathways leading to linker cell death and necrotic death, may provide insight into the regulation of cell death in some, and perhaps all other animals.
Abbreviations
- BH3:
-
Bcl-2 Homology domain 3
- CEM neurons:
-
cephalic companion neurons
- Cdx2:
-
caudal-related homeobox 2
- MAPK:
-
mitogen-activated protein kinase
- GFP:
-
green fluorescent protein
- RT–PCR:
-
reverse transcriptase-PCR
- DEG/ENaC:
-
degenerin and epithelial Na+ channel
References
Baehrecke EH . How death shapes life during development. Nat Rev Mol Cell Biol 2002; 3: 779–787.
Hipfner DR, Cohen SM . Connecting proliferation and apoptosis in development and disease. Nat Rev Mol Cell Biol 2004; 5: 805–815.
Jacobson MD, Weil M, Raff MC . Programmed cell death in animal development. Cell 1997; 88: 347–354.
Meier P, Finch A, Evan G . Apoptosis in development. Nature 2000; 407: 796–801.
Kuida K, Zheng TS, Na S, Kuan C, Yang D, Karasuyama H et al. Decreased apoptosis in the brain and premature lethality in CPP32-deficient mice. Nature 1996; 384: 368–372.
Kuida K, Haydar TF, Kuan CY, Gu Y, Taya C, Karasuyama H et al. Reduced apoptosis and cytochrome c-mediated caspase activation in mice lacking caspase 9. Cell 1998; 94: 325–337.
White K, Grether ME, Abrams JM, Young L, Farrell K, Steller H . Genetic control of programmed cell death in Drosophila. Science 1994; 264: 677–683.
Ellis HM, Horvitz HR . Genetic control of programmed cell death in the nematode C. elegans. Cell 1986; 44: 817–829.
Sulston JE, Horvitz HR . Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev Biol 1977; 56: 110–156.
Sulston JE, Schierenberg E, White JG, Thomson JN . The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol 1983; 100: 64–119.
Gumienny TL . Genetic control of programmed cell death in the Caenorhabditis elegans hermaphrodite germline. Development 1999; 126: 1011–1022.
Conradt B, Horvitz HR . The C. elegans protein EGL-1 is required for programmed cell death and interacts with the Bcl-2-like protein CED-9. Cell 1998; 93: 519–529.
Metzstein MM, Stanfield GM, Horvitz HR . Genetics of programmed cell death in C. elegans past, present and future. Trends Genet 1998; 14: 410–416.
Lettre G, Hengartner MO . Developmental apoptosis in C. elegans: a complex CEDnario. Nat Rev Mol Cell Biol 2006; 7: 97–108.
Shaham S, Horvitz HR . Developing Caenorhabditis elegans neurons may contain both cell-death protective and killer activities. Genes Dev 1996; 10: 578–591.
Hengartner MO, Ellis RE, Horvitz HR . Caenorhabditis elegans gene ced-9 protects cells from programmed cell death. Nature 1992; 356: 494–499.
Hengartner MO, Horvitz HR . Activation of C. elegans cell death protein CED-9 by an amino-acid substitution in a domain conserved in Bcl-2. Nature 1994; 369: 318–320.
Yuan J, Shaham S, Ledoux S, Ellis HM, Horvitz HR . The C. elegans cell death gene ced-3 encodes a protein similar to mammalian interleukin-1 beta-converting enzyme. Cell 1993; 75: 641–652.
Chen F, Hersh BM, Conradt B, Zhou Z, Riemer D, Gruenbaum Y et al. Translocation of C. elegans CED-4 to nuclear membranes during programmed cell death. Science 2000; 287: 1485–1489.
Yan N, Chai J, Lee ES, Gu L, Liu Q, He J et al. Structure of the CED-4-CED-9 complex provides insights into programmed cell death in Caenorhabditis elegans. Nature 2005; 437: 831–837.
Spector MS, Desnoyers S, Hoeppner DJ, Hengartner MO . Interaction between the C. elegans cell-death regulators CED-9 and CED-4. Nature 1997; 385: 653–656.
del Peso L, González VM, Núñez G . Caenorhabditis elegans EGL-1 disrupts the interaction of CED-9 with CED-4 and promotes CED-3 activation. J Biol Chem 1998; 273: 33495–33500.
Yan N, Gu L, Kokel D, Chai J, Li W, Han A et al. Structural, biochemical, and functional analyses of CED-9 recognition by the proapoptotic proteins EGL-1 and CED-4. Mol Cell 2004; 15: 999–1006.
Yang X, Chang HY, Baltimore D . Essential role of CED-4 oligomerization in CED-3 activation and apoptosis. Science 1998; 281: 1355–1357.
Chinnaiyan AM, O′Rourke K, Lane BR, Dixit VM . Interaction of CED-4 with CED-3 and CED-9: a molecular framework for cell death. Science 1997; 275: 1122–1126.
Conradt B, Horvitz HR . The TRA-1A sex determination protein of C. elegans regulates sexually dimorphic cell deaths by repressing the egl-1 cell death activator gene. Cell 1999; 98: 317–327.
Metzstein MM, Hengartner MO, Tsung N, Ellis RE, Horvitz HR . Transcriptional regulator of programmed cell death encoded by Caenorhabditis elegans gene ces-2. Nature 1996; 382: 545–547.
Metzstein MM, Horvitz HR . The C. elegans cell death specification gene ces-1 encodes a snail family zinc finger protein. Mol Cell 1999; 4: 309–319.
Ellis RE, Horvitz HR . Two C. elegans genes control the programmed deaths of specific cells in the pharynx. Development 1991; 112: 591–603.
Thellmann M, Hatzold J, Conradt B . The Snail-like CES-1 protein of C. elegans can block the expression of the BH3-only cell-death activator gene egl-1 by antagonizing the function of bHLH proteins. Development 2003; 130: 4057–4071.
Liu H, Strauss TJ, Potts MB, Cameron S . Direct regulation of egl-1 and of programmed cell death by the Hox protein MAB-5 and by CEH-20, a C. elegans homolog of Pbx1. Development 2006; 133: 641–650.
Kimble JE, White JG . On the control of germ cell development in Caenorhabditis elegans. Dev Biol 1981; 81: 208–219.
Kimble J, Crittenden SL . Controls of germline stem cells, entry into meiosis, and the sperm/oocyte decision in Caenorhabditis elegans. Annu Rev Cell Dev Biol 2007; 23: 405–433.
Church DL, Guan KL, Lambie EJ . Three genes of the MAP kinase cascade, mek-2, mpk-1/sur-1 and let-60 ras, are required for meiotic cell cycle progression in Caenorhabditis elegans. Development 1995; 121: 2525–2535.
Parrish J, Metters H, Chen L, Xue D . Demonstration of the in vivo interaction of key cell death regulators by structure-based design of second-site suppressors. Proc Natl Acad Sci USA 2000; 97: 11916–11921.
Park D, Jia H, Rajakumar V, Chamberlin HM . Pax2/5/8 proteins promote cell survival in C. elegans. Development 2006; 133: 4193–4202.
Sato H, Wang D, Kudo A . Dissociation of Pax-5 from KI and KII sites during kappa-chain gene rearrangement correlates with its association with the underphosphorylated form of retinoblastoma. J Immunol 2001; 166: 6704–6710.
Eberhard D, Busslinger M . The partial homeodomain of the transcription factor Pax-5 (BSAP) is an interaction motif for the retinoblastoma and TATA-binding proteins. Cancer Res 1999; 59: 1716s–1724s;discussion 1724s–1725s.
Reddien PW, Andersen EC, Huang MC, Horvitz HR . DPL-1 DP, LIN-35 Rb and EFL-1 E2F act with the MCD-1 zinc-finger protein to promote programmed cell death in Caenorhabditis elegans. Genetics 2007; 175: 1719–1733.
Schertel C, Conradt B . C. elegans orthologs of components of the RB tumor suppressor complex have distinct pro-apoptotic functions. Development 2007; 134: 3691–3701.
Schwartz HT, Horvitz HR . The C. elegans protein CEH-30 protects male-specific neurons from apoptosis independently of the Bcl-2 homolog CED-9. Genes Dev 2007; 21: 3181–3194.
Peden E, Kimberly E, Gengyo-Ando K, Mitani S, Xue D . Control of sex-specific apoptosis in C. elegans by the BarH homeodomain protein CEH-30 and the transcriptional repressor UNC-37/Groucho. Genes Dev 2007; 21: 3195–3207.
Zarkower D, Hodgkin J . Zinc fingers in sex determination: only one of the two C. elegans Tra-1 proteins binds DNA in vitro. Nucleic Acids Res 1993; 21: 3691–3698.
Li S, Price SM, Cahill H, Ryugo DK, Shen MM, Xiang M . Hearing loss caused by progressive degeneration of cochlear hair cells in mice deficient for the Barhl1 homeobox gene. Development 2002; 129: 3523–3532.
Li S, Xiang M . Barhl1 is required for maintenance of a large population of neurons in the zonal layer of the superior colliculus. Dev Dyn 2006; 235: 2260–2265.
Maurer CW, Chiorazzi M, Shaham S . Timing of the onset of a developmental cell death is controlled by transcriptional induction of the C. elegans ced-3 caspase-encoding gene. Development 2007; 134: 1357–1368.
Bonhomme C, Duluc I, Martin E, Chawengsaksophak K, Chenard MP, Kedinger M et al. The Cdx2 homeobox gene has a tumour suppressor function in the distal colon in addition to a homeotic role during gut development. Gut 2003; 52: 1465–1471.
Aoki K, Tamai Y, Horiike S, Oshima M, Taketo MM . Colonic polyposis caused by mTOR-mediated chromosomal instability in Apc+/Delta716 Cdx2+/− compound mutant mice. Nat Genet 2003; 35: 323–330.
Chawengsaksophak K, James R, Hammond VE, Kontgen F, Beck F . Homeosis and intestinal tumours in Cdx2 mutant mice. Nature 1997; 386: 84–87.
Silberg DG, Swain GP, Suh ER, Traber PG . Cdx1 and cdx2 expression during intestinal development. Gastroenterology 2000; 119: 961–971.
Abraham MC, Lu Y, Shaham S . A morphologically conserved nonapoptotic program promotes linker cell death in Caenorhabditis elegans. Dev Cell 2007; 12: 73–86.
Kimble J, Hirsh D . The postembryonic cell lineages of the hermaphrodite and male gonads in Caenorhabditis elegans. Dev Biol 1979; 70: 396–417.
Sulston JE, Albertson DG, Thomson JN . The Caenorhabditis elegans male: postembryonic development of nongonadal structures. Dev Biol 1980; 78: 542–576.
Ambros V, Horvitz HR . Heterochronic mutants of the nematode Caenorhabditis elegans. Science 1984; 226: 409–416.
Slack F, Ruvkun G . Temporal pattern formation by heterochronic genes. Annu Rev Genet 1997; 31: 611–634.
Rougvie AE, Ambros V . The heterochronic gene lin-29 encodes a zinc finger protein that controls a terminal differentiation event in Caenorhabditis elegans. Development 1995; 121: 2491–2500.
Liu Z, Kirch S, Ambros V . The Caenorhabditis elegans heterochronic gene pathway controls stage-specific transcription of collagen genes. Development 1995; 121: 2471–2478.
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000; 403: 901–906.
Slack FJ, Basson M, Liu Z, Ambros V, Horvitz HR, Ruvkun G . The lin-41 RBCC gene acts in the C. elegans heterochronic pathway between the let-7 regulatory RNA and the LIN-29 transcription factor. Mol Cell 2000; 5: 659–669.
Shaham S . Identification of multiple Caenorhabditis elegans caspases and their potential roles in proteolytic cascades. J Biol Chem 1998; 273: 35109–35117.
Chautan M, Chazal G, Cecconi F, Gruss P, Golstein P . Interdigital cell death can occur through a necrotic and caspase-independent pathway. Curr Biol 1999; 9: 967–970.
Susin SA, Lorenzo HK, Zamzami N, Marzo I, Snow BE, Brothers GM et al. Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 1999; 397: 441–446.
Cheng EH, Wei MC, Weiler S, Flavell RA, Mak TW, Lindsten T et al. BCL-2, BCL-X(L) sequester BH3 domain-only molecules preventing BAX- and BAK-mediated mitochondrial apoptosis. Mol Cell 2001; 8: 705–711.
Holler N, Zaru R, Micheau O, Thome M, Attinger A, Valitutti S et al. Fas triggers an alternative, caspase-8-independent cell death pathway using the kinase RIP as effector molecule. Nat Immunol 2000; 1: 489–495.
Syntichaki P, Xu K, Driscoll M, Tavernarakis N . Specific aspartyl and calpain proteases are required for neurodegeneration in C. elegans. Nature 2002; 419: 939–944.
Zhou Z, Hartwieg E, Horvitz HR . CED-1 is a transmembrane receptor that mediates cell corpse engulfment in C. elegans. Cell 2001; 104: 43–56.
Pilar G, Landmesser L . Ultrastructural differences during embryonic cell death in normal and peripherally deprived ciliary ganglia. J Cell Biol 1976; 68: 339–356.
Chu-Wang IW, Oppenheim RW . Cell death of motoneurons in the chick embryo spinal cord. I. A light and electron microscopic study of naturally occurring and induced cell loss during development. J Comp Neurol 1978; 177: 33–57.
Clarke PG . Developmental cell death: morphological diversity and multiple mechanisms. Anat Embryol 1990; 181: 195–213.
Oppenheim RW, Flavell RA, Vinsant S, Prevette D, Kuan CY, Rakic P . Programmed cell death of developing mammalian neurons after genetic deletion of caspases. J Neurosci 2001; 21: 4752–4760.
Bloss TA, Witze ES, Rothman JH . Suppression of CED-3-independent apoptosis by mitochondrial betaNAC in Caenorhabditis elegans. Nature 2003; 424: 1066–1071.
Joshi P, Eisenmann DM . The Caenorhabditis elegans pvl-5 gene protects hypodermal cells from ced-3-dependent, ced-4-independent cell death. Genetics 2004; 167: 673–685.
Mangahas PM, Zhou Z . Clearance of apoptotic cells in Caenorhabditis elegans. Semin Cell Dev Biol 2005; 16: 295–306.
Ferguson EL, Horvitz HR . Identification and characterization of 22 genes that affect the vulval cell lineages of the nematode Caenorhabditis elegans. Genetics 1985; 110: 17–72.
Driscoll M . Molecular genetics of cell death in the nematode Caenorhabditis elegans. J Neurobiol 1992; 23: 1327–1351.
Chalfie M, Sulston J . Developmental genetics of the mechanosensory neurons of Caenorhabditis elegans. Dev Biol 1981; 82: 358–370.
Driscoll M, Chalfie M . The mec-4 gene is a member of a family of Caenorhabditis elegans genes that can mutate to induce neuronal degeneration. Nature 1991; 349: 588–593.
Chalfie M, Wolinsky E . The identification and suppression of inherited neurodegeneration in Caenorhabditis elegans. Nature 1990; 345: 410–416.
Hall DH, Gu G, García-Añoveros J, Gong L, Chalfie M, Driscoll M . Neuropathology of degenerative cell death in Caenorhabditis elegans. J Neurosci 1997; 17: 1033–1045.
Chung S, Gumienny TL, Hengartner MO, Driscoll M . A common set of engulfment genes mediates removal of both apoptotic and necrotic cell corpses in C. elegans. Nat Cell Biol 2000; 2: 931–937.
Hong K, Driscoll M . A transmembrane domain of the putative channel subunit MEC-4 influences mechanotransduction and neurodegeneration in C. elegans. Nature 1994; 367: 470–473.
Goodman MB, Ernstrom GG, Chelur DS, O'Hagan R, Yao CA, Chalfie M . MEC-2 regulates C. elegans DEG/ENaC channels needed for mechanosensation. Nature 2002; 415: 1039–1042.
Bianchi L, Gerstbrein B, Frokjaer-Jensen C, Royal DC, Mukherjee G, Royal MA et al. The neurotoxic MEC-4(d) DEG/ENaC sodium channel conducts calcium: implications for necrosis initiation. Nat Neurosci 2004; 7: 1337–1344.
Treinin M, Chalfie M . A mutated acetylcholine receptor subunit causes neuronal degeneration in C. elegans. Neuron 1995; 14: 871–877.
Yassin L, Gillo B, Kahan T, Halevi S, Eshel M, Treinin M . Characterization of the deg-3/des-2 receptor: a nicotinic acetylcholine receptor that mutates to cause neuronal degeneration. Mol Cell Neurosci 2001; 17: 589–599.
Korswagen HC, Park JH, Ohshima Y, Plasterk RH . An activating mutation in a Caenorhabditis elegans Gs protein induces neural degeneration. Genes Dev 1997; 11: 1493–1503.
Berger AJ, Hart AC, Kaplan JM . G alphas-induced neurodegeneration in Caenorhabditis elegans. J Neurosci 1998; 18: 2871–2880.
Mongin AA . Disruption of ionic and cell volume homeostasis in cerebral ischemia: the perfect storm. Pathophysiology 2007; 14: 183–193.
Xiong ZG, Zhu XM, Chu XP, Minami M, Hey J, Wei WL et al. Neuroprotection in ischemia: blocking calcium-permeable acid-sensing ion channels. Cell 2004; 118: 687–698.
Michalak M, Corbett EF, Mesaeli N, Nakamura K, Opas M . Calreticulin: one protein, one gene, many functions. Biochem J 1999; 344 (Part 2): 281–292.
Xu K, Tavernarakis N, Driscoll M . Necrotic cell death in C. elegans requires the function of calreticulin and regulators of Ca(2+) release from the endoplasmic reticulum. Neuron 2001; 31: 957–971.
Mattson MP, LaFerla FM, Chan SL, Leissring MA, Shepel PN, Geiger JD . Calcium signaling in the ER: its role in neuronal plasticity and neurodegenerative disorders. Trends Neurosci 2000; 23: 222–229.
Syntichaki P, Samara C, Tavernarakis N . The vacuolar H+ -ATPase mediates intracellular acidification required for neurodegeneration in C. elegans. Curr Biol 2005; 15: 1249–1254.
Luke CJ, Pak SC, Askew YS, Naviglia TL, Askew DJ, Nobar SM et al. An intracellular serpin regulates necrosis by inhibiting the induction and sequelae of lysosomal injury. Cell 2007; 130: 1108–1119.
Toth ML, Simon P, Kovacs AL, Vellai T . Influence of autophagy genes on ion-channel-dependent neuronal degeneration in Caenorhabditis elegans. J Cell Sci 2007; 120: 1134–1141.
Samara C, Syntichaki P, Tavernarakis N . Autophagy is required for necrotic cell death in Caenorhabditis elegans. Cell Death Differ 2008; 15: 105–112.
Acknowledgements
We thank Michael Chiorazzi for comments on the manuscript. ESB is supported in part by The Rockefeller University's Women & Science Fellowship Program and by a grant from the NIH. MD is supported by NIH grant R01NS034435 and by the NJ Commission on Spinal Cord Research. SS is a Klingenstein fellow in the neurosciences and a Monique Weill-Caulier Scholar. This work was supported in part by a grant from the NIH to SS. We take this opportunity to offer our best wishes to Richard Lockshin, a key figure in developing the concept of programmed cell death, on the occasion of his 70th birthday.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Blum, E., Driscoll, M. & Shaham, S. Noncanonical cell death programs in the nematode Caenorhabditis elegans. Cell Death Differ 15, 1124–1131 (2008). https://doi.org/10.1038/cdd.2008.56
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.56
Keywords
This article is cited by
-
Non-caspase proteases: triggers or amplifiers of apoptosis?
Cellular and Molecular Life Sciences (2010)