Abstract
The enzymatic activity of phospholipase D (PLD) is known to be essential for cell survival and protection from apoptosis. However, the mechanisms regulating PLD activity during apoptosis remain unknown. Here we report that cleavage of PLD1 by caspases facilitates p53-mediated apoptosis. Cleavage of PLD1 into an N-terminal fragment (NF-PLD1) and a C-terminal fragment at the amino-acid sequence, DDVD545, led to a reduction in PLD1 activity. However, a caspase-resistant mutant form of PLD1 retained significant levels of enzymatic activity and apoptotic function as compared to wild-type PLD1. Exogenous NF-PLD1 expression induced apoptosis through a dominant-negative effect on the activity of endogenous PLD1. During apoptosis, a small fraction of PLD1 is cleaved by caspases in a p53-independent manner and NF-PLD1 amplifies apoptotic signaling through inhibition of the remaining PLD1 activity. As PLD1 suppresses the ATM–Chk2–p53 pathway, elimination of PLD1 activity through NF-PLD1 or si-RNA against PLD1 increases apoptosis in a p53-dependent manner. Taken together, our results reveal that cleavage of PLD1 by caspases promotes apoptosis via modulation of the p53-dependent cell death pathway.
Similar content being viewed by others
Main
Phospholipase D (PLD) catalyzes the hydrolysis of phosphatidylcholine (PC) to generate the lipid second messenger, phosphatidic acid (PA), and choline. PLD and PA have been implicated in a wide variety of cellular processes including cell proliferation, survival, vesicle transport, cytoskeletal organization and receptor signaling.1, 2 Members of the PLD superfamily share a highly conserved HKD motif, which contains the sequence HxxxxKxD, and is essential for catalysis.3 Two copies of the HKD motif are present in most PLD superfamily members. Studies of the crystal structure of bacterial PLD have revealed that the two HKD motifs juxtapose to form a single active site.4 To date, two mammalian PLD isozymes (PC-specific PLD1 and PLD2), both with splice variants, have been reported.3, 5 PLD1, but not PLD2, possesses a 116–amino acid loop region between the two catalytic motifs, suggesting that it may act as a negative regulatory element leading to the low basal activity levels of PLD1 as compared to PLD2.6
Apoptosis is a strong barrier against tumorigenesis that cancer cells must overcome through apoptotic inhibition or through the activation of survival signaling.7 PLD is elevated in several human cancers8 and it has been reported to contribute to cancer progression through the suppression of p53-mediated apoptosis.9, 10, 11 Despite an increasing repertoire of cellular activities that have been found to involve PLD, the apoptosis-related molecular mechanisms of PLD remain largely unknown. In this report, we demonstrate for the first time that PLD1 is cleaved by caspases and that the resulting loss of PLD activity contributes to apoptosis. The dominant-negative action of the N-terminal proteolytic product of PLD1 amplifies p53 signaling and apoptosis through phosphorylation of check point kinase-2 (Chk2) and p53, which enhances p53 stability and inhibits binding of p53 to mouse double minute 2 (MDM2). This study provides a novel mechanism by which cleavage of PLD1 by caspases facilitates apoptosis via a p53-dependent cell death pathway.
Results
PLD1 undergoes caspase-mediated cleavage in vitro and in vivo
To investigate the regulation of PLD expression during apoptosis, we first monitored the fate of the PLD protein in various cell lines originating from different species, using antibodies to a C-terminal PLD1 peptide. In human U87-MG glioma and HEK-293 cells, treatment with etoposide, a DNA-damaging agent, decreased the levels of intact PLD1 and generated a truncated form of PLD1 that migrated at approximately 56 kDa based on western blot analysis. As the PLD1 antibody used in this experiment was raised against a C-terminal epitope, the recognized band was a C-terminal fragment of PLD1 (CF-PLD1; Figure 1a). We also observed time- and dose-dependent PLD1 cleavage in mouse fibroblasts (NIH3T3; Figure 1b). PLD1 cleavage was induced by other types of anticancer drugs including adriamycin and cisplatin (Supplementary Figure S1). These results suggested that PLD1 underwent degradation during apoptosis and supported the existence of a post-translational regulatory mechanism for PLD1.
PLD1 was cleaved by caspases in vitro and in vivo. (a) Human U87MG and HEK293 cells were treated with etoposide for 18 h. The cell lysates were analyzed by western blotting using an antibody to a C-terminal peptide of PLD1. (b) NIH3T3 mouse fibroblasts were treated with 25 μM etoposide for the indicated time periods or the indicated doses for 16 h. The cell lysates were analyzed by western blotting using the indicated antibodies. (c) pcDNA3.1-hPLD1b was transcribed and translated in vitro with [35S] methionine and incubated with or without purified recombinant caspase-1, -3 or -8 as described in Materials and methods. The reaction mixtures were analyzed by SDS-PAGE and autoradiography. (d) Following incubation with or without 20 μM of the indicated caspase inhibitors, 50 μM of the indicated calpain inhibitors or 20 μM cathepsin inhibitor, U87-PLD1b cells were treated with etoposide (50 μM) for 18 h. The lysates were analyzed by western blotting using the indicated antibodies. (e) Alignments of the regions containing the caspase cleavage site of PLD1 from different species. (f) HEK293 cells were transfected with WT-PLD1 or D545A-PLD1, each N-terminally tagged with GFP, and were treated with or without etoposide for 18 h. The lysates were analyzed by western blotting with the indicated antibodies. *Nonspecific unknown protein. FL, full length; NF, N-terminal fragment; CF, C-terminal fragment; WT, wild-type. These blots are representative of three independent experiments
To determine whether PLD1 is a caspase substrate, we performed an in vitro protease assay using isotope-labeled PLD1 and several recombinant caspases. As shown in Figure 1c, all caspases tested (caspase-1, -3 and -8) were capable of cleaving PLD1.
Two fragments of 64 and 56 kDa were observed and were deduced to correspond to the N-terminal (NF-PLD1) and C-terminal fragments of PLD1, respectively.
To extend this analysis, we examined the in vivo digestion of PLD1 following the treatment of cells with caspase-specific inhibitors. From this experiment, we found that caspase-3, -8 and -9, but not caspase-1, were responsible for PLD1 digestion in vivo (Figure 1d). However, other noncapsase proteases, such as calpain or cathepsin, were not involved in PLD1 cleavage (Figure 1d).
As the consensus caspase cleavage sequence is DXXD, and PLD1 was cleaved at a single site into two fragments (64 and 56 kDa), we deduced the putative cleavage sites within PLD1 and generated 11 point mutants (Supplementary Figure S2a). Among the mutants that we tested, the D545A mutant was resistant to cleavage as determined by the in vitro caspase assay (Supplementary Figure S2b).
Interestingly, the caspase-3 cleavage site, DDVD545, was located within the loop region, which is not present in PLD2, and is highly conserved in mammalian PLD1 from various species (Figure 1e).
To further confirm this result, we transfected multiple vectors, each expressing a different PLD1 point mutant, and we examined the resulting transfectants for etoposide-induced cleavage of PLD1 (Supplementary Figure S2c). Consistent with our in vitro result, the D545A mutant was resistant to cleavage. We also observed a novel band at approximately 50 kDa in all the lanes of etoposide-treated samples. However, considering our previous observations, this is likely to be a nonspecific crossreactive band induced by etoposide. Although it would be very interesting to identify this protein, it seemed to be beyond the scope of this study.
We further confirmed the presence of the two fragments during apoptosis using an N-terminally tagged GFP-PLD1 construct. Cleavage of PLD1 was concomitant with an increase in the level of a GFP-tagged N-terminal fragment following induction of apoptosis, but the mutant (D545A) was resistant to cleavage (Figure 1f). The CF-PLD1 was detected with an anti-PLD antibody in wild-type PLD1-transfected cells, but not in the D545A-PLD1-transfected cells. As bands of approximately 110 and 90 kDa were detected with an antibody to GFP, we examined whether these bands are specific or not. Only NF-PLD1 was detected with an antibody to GFP in short exposures. Upon overexposure, the presumed nonspecific bands were detected by the GFP antibody in both the absence and the presence of the caspase inhibitor, but pretreatment with caspase inhibitors led to the loss of the NF-PLD1 band only, suggesting that the other proteins recognized by the GFP antibody are nonspecific bands (Supplementary Figure S2d). Taken together, these results demonstrated that the DDVD545 sequence in PLD1 underwent caspase-mediated cleavage in vitro and in vivo.
Caspase-mediated cleavage of PLD1 reduced its enzymatic activity
Next, we investigated the relationship between PLD1 cleavage and PLD1 activity. A caspase-3 inhibitor (z-DEVD) blocked the etoposide-induced cleavage of PLD1 and caused a loss in PLD1 enzymatic activity in a dose-dependent manner (Figure 2a and b). Furthermore, we found that D545A-PLD1 showed resistance to the etoposide-induced reduction in PLD1 activity (Figure 2c). These results suggested that the loss of PLD activity might be due to cleavage of PLD1 by caspases.
Caspase-mediated cleavage of PLD1 led to a reduction in enzymatic activity. (a) NIH3T3 cells were treated with etoposide (50 μM) in the absence or presence of z-DEVD. Two tissue-culture dishes of cells were treated with each condition. One set was labeled with [3H] myristic acid, and PLD activity levels were measured. The values are represented as the mean±S.D. of five independent experiments. *P<0.01 versus treatment with etopside only. (b) The other set of culture dishes from (a) were analyzed by western blotting using the indicated antibodies. (c) HEK293 cells transfected with either of the GFP-tagged vectors, FL-WT-PLD1 or FL-D545A-PLD1, were treated with etoposide for the indicated times. The cells were labeled with [3H] myristic acid, and PLD activity levels were measured. The values are the mean±S.D. of five independent experiments. *P<0.01 versus vector-only control cells
Cleavage of PLD1 was required for apoptosis
To monitor the relationship between PLD1 cleavage and apoptosis, we compared general apoptotic characteristics including capsase activity (Figure 3a), annexin staining (Figure 3b), morphological changes (Figure 3c and d) and nuclear condensation (Supplementary Figure S3) of D545A-PLD1- and WT-PLD1-transfected cells. All of these indicators showed that PLD1 cleavage was required for proper apoptotic processes.
PLD1 cleavage was required for apoptosis. (a and b) HEK293 cells were transfected with either of the GFP-tagged vectors, FL-WT-PLD1 or FL-D545A-PLD1, and were treated with or without etoposide (50 μM) for 18 h. Forty hours after transfection, two culture dishes of cells subjected to each condition were analyzed by western blotting with the indicated antibodies and were also stained with annexin V. The percentage of annexin V-positive cells relative to the percentage of GFP-expressing cells was determined. (c and d) HEK293 cells were transfected with the same constructs as above and cells were analyzed by Hoechst staining. The percentage of apoptotic membrane blebbing over nuclei-condensed cells expressing GFP was determined. At least 100 cells were counted per well in three different wells. (e) HEK293 cells were transfected with the indicated constructs and were then treated with or without etoposide. Apoptosis was measured by flow cytometry after propidium iodide staining. The indicated percentages represent the ratio of cells in sub-G1 phase/total no. of cells. The values are the mean±S.D. of five independent experiments. *P<0.01 versus vector-only control cells, &hash nonspecific unknown protein
N-terminal fragment of PLD1-induced apoptosis
We further investigated whether the fragments produced by cleavage of PLD1 were required for apoptosis. Interestingly, ectopic expression of NF-PLD1, but not CF-PLD1, significantly induced apoptosis even in the absence of apoptotic stimuli as indicated by annexin V-positive GFP-expressing cells (Figure 4a), caspase-3 activation (Figure 4b) and a reduction in cell viability (Figure 4c). We further confirmed that NF-PLD1 suppressed cell viability in a colony-forming assay (Supplementary Figure S4a and b). To avoid technical interference due to experimental and cell-to-cell variations, we examined NF-PLD1-induced apoptosis in human breast cancer cells (MDA-MB361) and mouse embryonic fibroblasts. In both cell lines, NF-PLD1-induced apoptosis was observed (Figure 4d and e).
N-terminal PLD1 fragment-induced apoptosis. (a) HEK293 cells were transfected with either of the GFP-tagged vectors, FL-PLD1, NF-PLD1 or CF-PLD1. Left panel: 40 h after transfection, nuclei were detected using Hoechst staining and annexin V-positive cells were identified. A representative cell in each image was enlarged for visual enhancement. Right panel: the percentage of GFP-expressing cells that were annexin V-positive and exhibited apoptotic nuclei was determined. At least 100 cells were counted per well in three different wells. Arrows indicate annexin V-positive NF-PLD1-expressing cells. (b) HEK293 cells were transfected with the indicated expression vector. Cells from each condition were analyzed by western blotting with the indicated antibody. The relative density of pro-caspase-3 to β-tubulin is indicated. (c) HEK293 cells were transfected with increasing quantities (0, 0.5, 1 and 3 μg) of plasmids encoding the indicated expression vector. Two days later, photographs were taken through a microscope. (d and e) MDA-MB361 and MEF cells were transfected with the indicated GFP-tagged constructs. Forty hours after transfection, nuclei were detected using Hoechst staining. The percentage of GFP-expressing cells exhibiting apoptotic nuclei were determined. At least 100 cells were counted per well in three different wells. The values are represented as the mean±S.D. of five independent experiments. *P<0.01 versus vector-only control cells
Suppression of PLD1 activity by NF-PLD1 was critical for apoptosis
As suppression of PLD1 activity appeared to be linked to apoptosis, we investigated the effect of NF-PLD1 on intact PLD1. First, we examined the binding between NF-PLD1 and intact PLD1 based on previous observations that PLD1 exerted its enzymatic activity through homo- or heterodimerization.12 We found that exogenous NF-PLD1 interacted with endogenous PLD1 (Figure 5a).
Suppression of PLD1 activity by NF-PLD1 was critical for apoptosis. (a) HEK293 cells were transfected with GFP vector and GFP-NF-PLD1. The lysates were immunoprecipitated with anti-GFP antibody and immunoblotted with antibody to PLD1. The lysates were also immunoblotted with antibody to GFP to assess its expression level. (b) Alignment of the amino acids present in the two HKD regions of PLD1 from different species. The amino acids that were mutated in this study are marked by asterisks. GFP-tagged PLD constructs are represented. (c) HEK293 cells were transfected with GFP vector or various GFP-NF-PLD1 constructs. The lysates were immunoprecipitated as in (a). (d) HEK293 cells were transiently transfected with the indicated expression vectors and labeled with [3H] myristic acid, and then the PLD activity was measured. (e) HEK293 cells were transfected with the indicated GFP constructs. After transfection for 40 h, the nuclei were stained with Hoechst and viewed by fluorescence microscopy. The percentage of GFP-expressing cells exhibiting apoptotic nuclei was determined. At least 100 cells were counted per well in three different wells. These experiments are representative of the results obtained from three experiments. The values are represented as the mean±S.D. of five independent experiments. *P<0.01 versus vector-only control cells
Patches of hydrophobic amino acids (leucine, isoleucine and valine) found either within or close to the HKD motifs, which participate in the catalytic activity, are highly conserved among mammalian PLD isozymes as outlined in Figure 5b. To investigate the role of this motif, we generated three vectors (H464E-, L467A- and I470A-NF-PLD1) and examined the binding of the mutants encoded by each of these vectors with endogenous PLD1. Two NF-PLD1 mutants (L467A and I470A) exhibited a lower affinity for PLD1 binding (Figure 5c) than wild-type NF-PLD1, suggesting that these hydrophobic amino acids are important for the intermolecular association between NF-PLD1 and endogenous intact PLD1. Next, we examined the effect of NF-PLD1 binding on PLD1 activity. Treatment of cells with phorbol 12-myristate 13-acetate (PMA), a well-known activator of PLD, increased PLD activity as expected.6 Transfection with WT-NF- or H464E-NF-PLD1 dramatically inhibited both basal and PMA-induced PLD activity. However, the inhibitory effects of L467A- and I470A-NF-PLD1 on basal and PMA-induced PLD activity were considerably lower than those of WT-NF- and H464E-NF-PLD1 (Figure 5d). In contrast, CF-PLD1 did not have any affect on PLD activity.
Moreover, NF-PLD1 was mainly localized in the cytoplasm and colocalized with endogenous PLD, whereas CF-PLD1 was localized in the nucleus but was not colocalized with endogenous PLD (Supplementary Figure S5), suggesting that the different subcellular localization patterns of both of these cleavage products may affect their activity and association with endogenous PLD. The reduced activity of PLD caused by ectopic expression of NF-PLD1 correlated closely with the effects on the intermolecular association with intact PLD1. Thus, it is postulated that NF-PLD1 exerts a dominant-negative effect by suppressing PLD activity due to intermolecular association with endogenous PLD1 through these hydrophobic amino acids. This may occur as a result of the dominant-negative NF-PLD1 preventing the activation of PLD1 by its regulators (PIP2, ARF, Rho, PKC) or impeding access to its membrane substrate.6
We next investigated whether the association of exogenous NF-PLD1 with endogenous PLD1 and the resulting inhibition of PLD1 activity are related to the pro-apoptotic effects of the PLD1 fragment. Expression of WT-NF- and H464E-NF-PLD1 induced significant levels of apoptosis as compared to the vector-only control based on the level of GFP-positive nuclear condensation (Figure 5f) and FACS analysis (Supplementary Figure S6). L467A- and I470A-NF-PLD1, which showed reduced association with endogenous PLD1 and lower potency in the inhibition of PLD activity, were found to significantly suppress apoptosis, compared with WT-NF- and H464E-NF-PLD1. These results suggested that suppression of PLD1 activity by NF-PLD1 was critical for apoptosis.
NF-PLD1-induced apoptosis was achieved through a p53-dependent pathway
Although NF-PLD1 induced apoptosis through the suppression of PLD1 activity, it was unclear how the reduction in PLD1 activity promoted apoptosis. Elevated PLD activity suppresses DNA damage-induced increases in p53 levels due to increased expression of MDM2.13 Thus, we examined the role of p53 in NF-PLD1-induced apoptosis using HCT116 p53-positive and -negative isogenic cell lines. As shown in Figure 6a, expression of NF-PLD1 enhanced p53 expression and suppressed MDM2 levels. However, in p53-deficient cells, a reduction in MDM2 levels was undetected. We did not observe NF-PLD1-induced apoptosis in p53-deficient cells as measured by nuclear condensation (Figure 6b) and FACS analysis (Supplementary Figure S7). We further confirmed that NF-PLD1 did not induce apoptosis in another p53-null PC3 prostate cancer cell line. Ectopic expression of p53 in the null cells significantly enhanced NF-PLD1-induced apoptosis, and the p53 protein levels were increased by exogenous NF-PLD1 (Figure 6d and e). Furthermore, ectopic expression of WT-NF- and H464E-NF-PLD1 in HCT116 cells significantly elevated p53 protein levels as well as phosphorylation of p53 at Ser-20, which is known to render the protein resistant to degradation by MDM2 (Figure 6f). However, the increase in p53 levels was suppressed in L467A- and I470A-NF-PLD1-expressing cells compared with cells expressing WT-NF- or H464E-NF-PLD1. We next examined whether the effect of NF-PLD1 on the induction of p53 was also reflected at the level of p53-targeted gene expression. The induction of p53 target proteins including p21, Bax, Egr-1 and PTEN in p53-elevated samples was observed (Figure 6f). Therefore, it was suggested that NF-PLD1-induced apoptosis correlated with the extent of hydrophobic interactions with endogenous PLD1 and may be intimately related to an increase in p53 levels and downstream target genes.
NF-PLD1-induced apoptosis was achieved through a p53-dependent pathway. (a) HCT116 p53-null and HCT116 p53 wild-type cells were transfected with GFP vector or GFP-NF-PLD1. The lysates were immunoblotted with the indicated antibodies. (b and c) Under the same conditions as in (a), nuclei were detected using Hoechst staining for the measurement of nuclear condensation. The percentage of GFP-expressing cells exhibiting apoptotic nuclei was determined. At least 100 cells were counted per well in three different wells. Arrows indicate apoptotic nuclear condensation in NF-PLD1-expressing cells. (d and e) PC3 p53-null cells were transfected with a GFP vector, GFP-NF-PLD1 or a myc-tagged p53 vector. The lysates were immunoblotted with the indicated antibodies. Under the same conditions, apoptosis was measured by nuclear condensation. (f) HCT116 cells were transfected with the indicated GFP constructs. After transient transfection, cells were analyzed by immunoblotting with the indicated antibodies. (g) Upper panel: HCT116 cells were transfected with GFP vector or GFP-NF-PLD1. Twenty-four hours later, the cells were treated with cyclohexamide (100 μg/ml) for the indicated times, and the lysates were analyzed by western blotting with the indicated antibodies. Lower panel: densitometer analysis of the ratio of p53/tubulin. (h) HEK293 cells were transfected with GFP vector or GFP-NF-PLD1. The lysates were immunoprecipitated and/or immunoblotted with the indicated antibodies. (i) HCT116 cells were cotransfected with GFP-PLD1 and flag-tagged MDM2 (0, 1 and 3 μg) and then the lysates were analyzed by immunoblotting with the indicated antibodies. These blots are representative of results obtained from three experiments. The values are represented as the mean±S.D. of five independent experiments. *P<0.01 versus vector-only control cells
The levels of p53 are commonly regulated at the level of protein stabilization. To examine whether NF-PLD1 signaling had an impact on the stability of p53, we examined the levels of p53 protein in cells treated with cyclohexamide, which inhibits new p53 protein synthesis. The p53 protein levels were significantly stabilized by the expression of exogenous NF-PLD1 as compared to control cells expressing vector only during cyclohexamide treatment (Figure 6g). The stability of p53 is known to be mediated by the E3 ubiquitin ligase, MDM2, which facilitates the ubiquitination of p53 and targets it for degradation by the proteosome.14 Like p53, MDM2 is usually regulated at the level of protein stabilization. During treatment with cyclohexamide, the levels of MDM2 were significantly suppressed in NF-PLD1-expressed cells relative to those of the vector-only control cells (Figure 6g), suggesting that a decrease in the half-life of MDM2 is induced by NF-PLD1. In addition, MG132 abolished the NF-PLD1-mediated induction of p53 and reduction of MDM2 (Supplementary Figure S8), suggesting that NF-PLD1 regulates the protein stability and degradation of p53. We also found that NF-PLD1 disrupted the interaction between p53 and MDM2 by enhancing the phosphorlyation of p53 at both Ser-15 and Ser-20 (Figure 6h). As a reduction in MDM2 levels would be critical for NF-PLD1-mediated p53 stabilization, we added MDM2 into NF-PLD1-expressing cells. The exogenous MDM2 blocked the NF-PLD1-mediated p53 stabilization (Figure 6i). Taken together, these data strongly suggested that NF-PLD1-induced apoptosis might be achieved through a p53-dependent pathway.
Knockdown of PLD1 enhances p53 stabilization and sensitivity to DNA damage-induced apoptosis
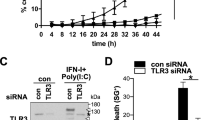
To investigate whether there is a correlation between PLD1 cleavage and p53 phosphorylation status, we examined the time course of etoposide-treated cells after transfection with siRNA specific for the vector or PLD1. Etoposide time-dependently enhanced both PLD1 cleavage and phosphorylation of p53 at both Ser-15 and Ser-20 residues, which prevents the binding of p53 to MDM2.15, 16 Upon DNA damage, p53 is rapidly activated through the ATM/ATR-dependent phosphorylation pathway.17 The phosphorylation of Chk2 (Thr-68) as a downstream event in this pathway was slightly enhanced by etoposide (Figure 7a). Moreover, the activation of caspase-3 by etoposide was concomitant with an increase in p53 stability (Figure 7a). This phenomena has also been reported by other research groups.18, 19, 20 In addition, it has been reported that caspase activation occurs in a p53-independent manner,21, 22, 23, 24, 25, 26 and substrate cleavage by caspases regulates p53-mediated signaling.27, 28 Thus, it is suggested that p53-independent apoptotic pathways induced by etoposide might induce a partial activation of caspases at early time points and then caspases are fully activated by a p53-dependent pathway. Interestingly, suppression of PLD1 using siRNA-mediated gene silencing enhanced etoposide-induced p53 stabilization, phosphorylation of Chk2 and p53, and caspase-3 activation, compared with control siRNA (Figure 7a). Moreover, the PLD1 cleavage fragment induced by etoposide was also weakly detected in p53-deficient cells, as compared with p53-positive cells (Figure 7b). Although etoposide very weakly induced apoptosis in p53-null cells as compared with p53-positive cells, knockdown of PLD1 significantly increased apoptosis in p53-positive cells but not in p53-null cells (Figure 7c). However, p53-independent PLD1 cleavage or caspase activation seems to be insufficient to induce apoptosis. Instead, partial PLD1 cleavage-induced p53 activation might be required for apoptosis. In Figure 7a, PLD1 is weakly cleaved after 3 h of etoposide treatment, whereas the phosphorylation of p53 is strong. As mentioned above, the initial cleavage of PLD1 is a p53-independent event and was only partial. Therefore, the early response of PLD1 cleavage induced by DNA damage is weak. However, this cleavage can enhance the phosphorylation of p53. Thus, different kinetics should be observed (PLD1 cleavage occurs first, followed by phosphorylation of p53 and then complete cleavage of PLD1). Finally, we examined the involvement of PA, the end product of PLD1 activity, on the check point kinase–p53 pathway. Treatment with PA suppressed DNA damage-induced p53 induction and phosphorylation of Chk2 (Figure 7d). This result suggested that a reduction in PA levels derived from PLD1 digestion or other causes could facilitate p53 activation and apoptosis. Taken together, these results demonstrate that endogenous PLD1 plays an antiapoptotic role via the modulation of a p53-dependent pathway.
Knockdown of PLD1 enhances p53 stability and sensitivity to DNA damage-induced apoptosis. (a) HCT116 cells were transfected with control or PLD1 siRNA. Forty hours after transfection, the cells were treated with etoposide (100 μM) for the indicated times, and the lysates were immunoblotted with the indicated antibodies. (b) After transfection with the indicated siRNA, two isogenic HCT116 cells (WT and p53-null) were treated with or without etoposide, and the lysates were immunoblotted with the indicated antibodies. (c) Under the same conditions as in (b), apoptosis was measured by nuclear condensation. The percentage of GFP-expressing cells exhibiting apoptotic nuclei was determined. At least 100 cells were counted per well in three different wells. *P<0.01 versus vector-only control cells. (d) MCF7 cells were pretreated with PA (50 μM) for 30 min followed by treatment with vehicle (Veh) or etoposide (Eto, 10 μM) for 2 h. The lysates were analyzed by immunoblotting using the indicated antibodies
Discussion
PLD activity and protein levels play important roles in inhibiting apoptosis by acting at multiple levels of the apoptotic cascade. In the present study, we demonstrate that cleavage of PLD1 results in the loss of its activity and induces apoptosis. We provide the first evidence that the DDVD545 motif of PLD1 is cleaved by caspases in vitro and in vivo during apoptosis. Interestingly, the DDVD545 caspase cleavage motif in PLD1 has been highly conserved from mouse to humans.
We also observed elevated expression of CF-PLD1 in Alzheimer’s disease (data not shown), implying that cleavage of PLD1 is a physiological event.
Interestingly, it has been reported that hPLD1c exists as a splice variant of PLD1 with a molecular weight of 69 kDa (1–597 amino acids). Moreover, the amino-acid sequence of hPLD1c from residues 1 to 513 is identical to that of NF-PLD1b (1–545 amino acids). However, the amino-acid sequence of hPLD1c from residues 514 to 597 is altered by splicing, resulting in premature transcriptional termination.29 Therefore, the endogenous splice variant, hPLD1c, containing one HKD motif might have pro-apoptotic activity like NF-PLD1, suggesting the physiological relevance of the cleavage fragment of PLD1.
In general, apoptosis-related caspase activity serves two different purposes: one is eliminating functional proteins by digestion into inactive forms (for example, poly(ADP-ribose) polymerase (PARP) and structural proteins) and the other is the generation of new biological activities (for example, Bid truncation and pro-caspase activation). In the case of PLD1, both purposes are achieved. First, caspase-mediated cleavage of PLD1 may act as an inactivating process as PLD activity in WT-PLD1-expressing cells, but not in caspase-resistant PLD1-expressing cells, is significantly decreased during apoptosis. It is also suggested that the loss of PLD activity due to the cleavage of PLD1 is involved in the progression of apoptosis. Second, the NF-PLD1 fragment induces pro-apoptotic activity, which might be due to its dominant-negative effect through hydrophobic interactions with endogenous PLD1 and subsequent inhibition of its activity. This is further supported by our observation that mutations in the hydrophobic residues within the HKD motif rescue the dominant-negative effect. In this paper, we have provided evidence that the dominant-negative action of NF-PLD1 inhibits the interaction of MDM2 with p53 through phosphorylation of p53. Ultimately, NF-PLD1-induced p53 stabilization, which elevated downstream p53 target genes, promoted apoptosis. However, NF-PLD1 did not affect apoptosis in p53-deficient cells. Although CF-PLD1 had no effect on apoptosis, the fragment was localized to the nucleus and is postulated to play a role in other cellular physiological processes such as transcriptional regulation after nuclear import. The role of CF-PLD1 needs to be elucidated in future studies.
How does inhibition of PLD1 activity by NF-PLD1 activate p53 and suppress the interaction between p53 and MDM2? Indeed, p53-independent caspase activation has been proposed during apoptosis.21, 22, 23, 24, 25, 26 Moreover, it has been reported that the cleavage of some substrates by caspase can regulate p53- and ATM/ATR kinase-mediated signaling upstream of caspase-3 during DNA damage-induced apoptosis.27, 28 Thus, it is suggested that p53-independent apoptotic pathways are induced by etoposide trigger caspase activation at an early time point and that full activation of caspases is a late event mediated by a p53-dependent pathway. The present study suggests that caspase activation at an early time point by etoposide induces the cleavage of PLD1 to generate NF-PLD1, which further amplifies p53 signaling and apoptosis through phosphorylation of p53 via a positive feedback loop system. Upon DNA damage, p53 is rapidly activated through the ATM/ATR-dependent phosphorylation pathway. Using RNA interference, we found that PLD1 might upregulate the upstream kinases including ATM or Chk2 that induce phosphorylation of p53 at Ser-15 and -20 and the subsequent inhibition of p53/MDM2 interactions.30 Indeed, treatment of cells with PA, the enzymatic product of PLD1, suppressed p53 and p-Chk2 induction (Figure 7d). This result implies that the enzymatic activity of PLD1 should be maintained for cell survival, and loss of PLD1 activity through caspase-mediated digestion or production of NF-PLD1 may facilitate DNA damage-induced apoptosis. Moreover, knockdown of PLD1 significantly promoted apoptosis in p53-positive cells, but not in p53-null cells, suggesting that PLD1 plays an antiapoptotic role probably through modulation of a p53-mediated cell death pathway. Our results suggest that cellular stresses such as DNA damage initially induce PLD1 cleavage and suppress PLD1 activity, which then results in the activation of p53. Activated p53 amplifies cell death signaling and this leads to complete PLD1 cleavage and apoptosis. In support of these results, the PLD1 cleavage fragment induced by etoposide was detected in U87MG and HEK293 cells, which are p53 function-deficient cell lines due to E1A, E1B or mutation, and in p53−/− cells. However, p53-independent PLD1 cleavage or caspase activation seems to be insufficient to induce apoptosis. Instead, partial PLD1 cleavage-induced p53 activation might be required for apoptosis. A proposed model for NF-PLD1-induced apoptosis is illustrated in Figure 8.
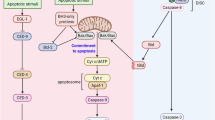
A proposed model for caspase-mediated PLD1 cleavage to promote p53-dependent apoptosis. Following treatment with DNA damage-inducing agents such as etoposide, PLD1 is cleaved by caspase-3 at an early time via a p53-independent pathway, generating NF-PLD1, which further amplifies p53 signaling and apoptosis through modulation of the ATM-dependent phosphorylation pathway
Although PLD plays an antiapoptotic role by regulating Fas versus the mitochondrial apoptotic pathway, the discrepancy between these mechanisms is unclear. It is known that PLD activity is required for protection from these apoptotic pathways. In response to Fas, PLD has been reported to prevent apoptosis by generating PA and increasing Bcl-2 and Bcl-xL expression.31 In response to the mitochondrial apoptotic signal, PLD is known to prevent apoptosis due to the elevation of PA levels and the subsequent decrease in the expression of tumor suppressor genes such as Egr-1 and PTEN.10 PLD1 cleavage may be induced by extrinsic apoptotic signals such as TNF-α and Fas. As mediators of extrinsic apoptotic pathways can sensitize cells to intrinsic apoptotic progression, the differential mechanisms of PLD in regulating Fas versus mitochondrial apoptotic pathways should be the subject of further studies.
As inhibition of PLD activity may be responsible for the increase in p53 stability and the suppression of MDM2 levels, drugs interfering with hydrophobic interactions involving PLD and its activity can inhibit the p53–MDM2 interaction and thus be an important target for cancer therapy. It is possible that hydrophobic peptides containing the HKD motif can bind to endogenous PLD and inhibit its activity, thereby inducing apoptosis. In summary, alternation in protein turnover due to PLD1 processing by caspases regulates apoptotic function, providing evidence for the existence of a novel mechanism in which caspase-dependent cleavage of PLD1 facilitates p53-mediated apoptosis.
Materials and Methods
Materials
MG132 was from AG scientific, CA, USA. z-VAD-fmk, z-YVAD-fmk, ac-DEVD-cho, ac-IETD-cho, ac-LEHD-cho, adriamycin, cisplatin and etoposide were from Alexis, Lausen. PD150606, MDL2870, pepstatin A, cathepsin B inhibitor and cycloheximide were from Calbiochem, CA, USA. The siRNA for hPLD1 (nucleotides 1486–1506, AAGGUGGGACGACAAUGAGCA) was purchased from Dharmacon Research Inc. (Lafayette, CO, USA).
Cell culture and transient transfection
HEK293, U87MG, MCF7, C6 glioma, mouse NIH3T3 fibroblasts, PC3 p53-null and HCT116 (wt and p53−/−) colon cancer were cultured at 37°C in DMEM (GIBCO BRL, CA, USA) containing 10% fetal bovine serum and 1% antibiotic–antimycotic. Cells were grown to 50–60% confluence for transient transfections using LipofectAMINE Plus (Invitrogen, CA, USA) according to the manufacturer's instructions.
Construction of plasmids and site-directed mutagenesis
Full-length constructs of hPLD1b, NF-PLD1 or CF-PLD1 cloned into pEGFP (Clontech) are described in the ‘Supplementary Materials and Methods’. The mutations in the caspase cleavage site (D545A-PLD1) and interdomain association sites (H464E, L467A and I470A) were generated from pEGFP-PLD1b or pEGFP-NF-PLD1b by PCR site-directed mutagenesis using the Quick Change Site-Directed Mutagenesis Kit (Stratagene, LaJolla, CA, USA) according to the manufacturer's instructions. Primer sequence information is described in ‘Supplementary Materials and Methods’. The mutations were confirmed by DNA sequencing.
Western blotting and immunoprecipitation
Cells were harvested for immunoblot analysis or immunoprecipitation as described previously.32 Antibodies to PARP (Santa Cruz, CA, USA), β-tubulin (Sigma, MO, USA), p53 (Santa Cruz), phospho-p53 (Ser-15/20) (Cell Signaling), MDM2 (Santa Cruz), phosphor-Chk2 (Thr68) (Cell Signaling), Chk2 (Cell Signaling) and GFP (Santa Cruz) were used as primary antibodies. Rabbit polyclonal anti-PLD1 antibody, which recognizes a C-terminal portion of PLD1, was generated as previously described.33
In vitro transcription/translation and caspase cleavage assays
Various PLD1 constructs were transcribed and translated in vitro using the TNT-coupled reticulocyte lysate system (Promega, WI, USA) according to the manufacturer's recommendations.
Fluorescence microscopy
Cells were transfected with various PLD1 constructs and incubated with media containing 1 mg/ml Hoechst (Molecular probes, CA, USA) for 20 min. Cells were visualized and images were collected using a fluorescence microscope (Axiovert 200M, Zwiss, Germany). For immunocytochemistry, cells were washed with PBS and fixed with 4% formaldehyde/PBS for 10 min, followed by permeabilization with 0.2% NP40/PBS for 5 min at room temperature. Cells were washed with PBS, blocked for 2 h in 20 mg/ml bovine serum albumin (BSA) and then incubated with primary antibody/BSA for 2 h. Cells were washed four times with PBS and then incubated with anti-rabbit texas red-conjugated secondary antibody or anti-mouse fluorescein isothiocyanate-conjugated secondary antibody (Jackson Immunoresearch Laboratories, PA, USA) containing 1 mg/ml Hoechst for 1 h. Cells were washed again four times with PBS and were mounted with antifading mounting medium (Vector Lab., CA, USA). Cells were visualized and images were collected using a fluorescence microscope.
PLD activity assays
PLD activity was assessed by measuring the formation of [3H] phosphatidylbutanol, the product of PLD-mediated transphosphatidylation, in the presence of 1-butanol as previously described.10
Measurement of phosphatidylserine expression using annexin V
Annexin V binding was assessed by fluorescence microscopy using a commercial kit (Invitrogen, Molecular probes, IN, USA) according to the instructions of the manufacturer.
Flow cytometry
Cells were cultured, harvested at the indicated times, stained with propidium iodide (1 μg/ml) according to the manufacturer's protocol and then analyzed using an FACScan flow cytometer. Cell cycle analysis was performed by flow cytometry using a commercially available software package (Modifit; Verity Winlist, Topsham, ME, USA).
Statistics
The results are expressed as mean±S.D. of the number of determinations indicated. Statistical significance of differences was determined by ANOVA. Significance was accepted when P<0.05 or 0.01.
Abbreviations
- CF-PLD1:
-
C-terminal fragment of PLD1
- Chk2:
-
check point kinase-2
- MDM2:
-
mouse double minute gene number 2
- NF-PLD1:
-
N-terminal fragment of PLD1
- PA:
-
phosphatidic acid
- PARP:
-
poly(ADP-ribose) polymerase
- PC:
-
phosphatidylcholine
- PLD:
-
phospholipase D
References
Singer WD, Brown HA, Sternweis PC . Regulation of eukaryotic phosphatidylinositol-specific phospholipase C and phospholipase D. Annu Rev Biochem 1997; 66: 475–509.
Freyberg Z, Siddhanta A, Shields D . ‘Slip, sliding away’ phospholipase D and the Golgi apparatus. Trends Cell Biol 2003; 13: 540–546.
Colley WC, Sung TC, Roll R, Jenco J, Hammond SM, Altshuller Y et al. Phospholipase D2, a distinct phospholipase D isoform with novel regulatory properties that provokes cytoskeletal reorganization. Curr Biol 1997; 7: 191–201.
Stuckey J, Dixon J . Crystal structure of a phospholipase D family member. Nat Struct Biol 1999; 6: 278–284.
Hammond SM, Altshuller YM, Sung TC, Rudge SA, Rose K, Engebrecht J et al. Human ADP-ribosylation factor-activated phosphatidylcholine-specific phospholipase D defines a new and highly conserved gene family. J Biol Chem 1995; 270: 29640–29643.
McDermott M, Wakelam MJ, Morris AJ . Phospholipase D. Biochem Cell Biol 2004; 82: 225–253.
Hanahan D, Weinberg R . The hallmarks of cancer. Cell 2000; 1: 57–70.
Foster DA, Xu L . Phospholipase D in cell proliferation and cancer. Mol Cancer Res 2003; 1: 789–800.
Hui L, Abbas T, Pielak RM, Joseph T, Bargonetti J, Foster DA . Phospholipase D elevates the level of MDM2 and suppresses DNA damage-induced increases in p53. Mol Cell Biol 2004; 24: 5677–5686.
Kim J, Lee YH, Kwon TK, Chang JS, Chung KC, Min DS . Phospholipase D prevents etoposide-induced apoptosis by inhibiting the expression of early growth response-1 and phosphatase and tensin homologue deleted on chromosome 10. Cancer Res 2006; 66: 784–793.
Kwun HJ, Lee JH, Min DS, Jang KL . Transcriptional repression of cyclin-dependent kinase inhibitor p21 gene by phospholipase D1 and D2. FEBS Lett 2003; 544: 38–44.
Kam Y, Exton JH . Dimerization of phospholipase d isozymes. Biochem Biophys Res Commun 2002; 1: 375–380.
Cai D, Zhong M, Wang R, Netzer WJ, Shields D, Zheng H et al. Phospholipase D1 corrects impaired APP trafficking and neurite outgrowth in familial Alzheimer's disease-linked presenilin-1 mutant neurons. PNAS 2006; 103: 1936–1940.
Michael D, Oren M . The p53-Mdm2 module and the ubiquitin system. Semin Cancer Biol 2003; 1: 49–58.
Chehab NH, Malikzay A, Stavridi ES, Halazonetis TD . Phosphorylation of Ser-20 mediates stabilization of human p53 in response to DNA damage. Proc Natl Acad Sci USA 1999; 96: 13777–13782.
Unger T, Juven-Gershon T, Moallem E, Berger M, Vogt SR, Lozano G et al. Critical role for Ser20 of human p53 in the negative regulation of p53 by Mdm2. EMBO J 1999; 7: 1805–1814.
Shiloh Y . ATM and related protein kinases: safeguarding genome integrity. Nat Rev Cancer 2003; 3: 155–168.
Kim SJ, Ju JW, Oh CD, Yoon YM, Song WK, Kim JH et al. ERK-1/2 and p38 kinase oppositely regulate nitric oxide-induced apoptosis of chondrocytes in association with p53, caspase-3, and differentiation status. J Biol Chem 2002; 277: 1332–1339.
Erster S, Mihara M, Kim RH, Petrenko O, Moll UM . In vivo mitochondrial p53 translocation triggers a rapid first wave of cell death in response to DNA damage that can precede p53 target gene activation. Mol Cell Biol 2004; 24: 6728–6741.
Dhanalakshmi S, Agarwal C, Singh RP, Agarwal R . Silibinin up-regulates DNA-protein kinase-dependent p53 activation to enhance UVB-induced apoptosis in mouse epithelial JB6 cells. J Biol Chem 2005; 21: 20375–20383.
Martins LM, Kottke TJ, Kaufmann SH, Earnshaw WC . Phosphorylated forms of activated caspases are present in cytosol from HL-60 cells during etoposide-induced apoptosis. Blood 1998; 9: 3042–3049.
Martins LM, Mesner PW, Kottke TJ, Basi GS, Sinha S, Tung JS et al. Comparison of caspase activation and subcellular localization in HL-60 and K562 cells undergoing etoposide-induced apoptosis. Blood 1997; 11: 4283–4296.
Shimada K, Nakamura M, Ishida E, Kishi M, Yonehara S, Konishi N . c-Jun NH2-terminal kinase-dependent Fas activation contributes to etoposide-induced apoptosis in p53-mutated prostate cancer cells. Prostate 2003; 4: 265–280.
Higginbottom K, Jahnke U, Newland AC, Cotter FE, Allen PD . New alternative phosphorylation sites on the cyclin dependent kinase 1/cyclin a complex in p53-deficient human cells treated with etoposide: possible association with etoposide-induced apoptosis. Apoptosis 2007; 10: 1847–1855.
Chatterjee D, Schmitz I, Krueger A, Yeung K, Kirchhoff S, Krammer PH et al. Induction of apoptosis in 9-nitrocamptothecin-treated DU145 human prostate carcinoma cells correlates with de novo synthesis of CD95 and CD95 ligand and down-regulation of c-FLIP(short). Cancer Res 2001; 19: 7148–7154.
Matkar S, Wrischnik L, Hellmann-Blumberg U . Sanguinarine causes DNA damage and p53-independent cell death in human colon cancer cell lines. Chem Biol Interact 2008; 1: 63–71.
Marchetti A, Cecchinelli B, D’Angelo M, D’Orazi G, Crescenzi M, Sacchi A et al. p53 can inhibit cell proliferation through caspase mediated cleavage of ERK2/MAPK. Cell Death Differ 2004; 11: 596–607.
Yim HS, Hwang IS, Choi J-S, Chun KH, Jin YH, Ham Y-M et al. Cleavage of Cdc6 by caspase-3 promotes ATM/ATR kinase-mediated apoptosis of HeLa cells. J Cell Biol 2006; 174: 77–88.
Steed PM, Clark KL, Boyar WC, Lasala DJ . Characterization of human PLD2 and the analysis of PLD isoform splice variants. FASEB J 1998; 12: 1309–1317.
Toledo F, Wahl GM . Regulating the p53 pathway: in vitro hypotheses, in vivo veritas. Nat Rev Cancer 2006; 12: 909–923.
Oh KJ, Lee SC, Choi HJ, Oh DY, Kim SC, Min DS et al. Role of phospholipase D2 in anti-apoptotic signaling through increased expressions of Bcl-2 and Bcl-xL. J Cell Biochem 2007; 6: 1409–1422.
Ahn BH, Kim SY, Kim EH, Choi KS, Kwon TK, Lee YH et al. Transmodulation between phospholipase D and c-Src enhances cell proliferation. Mol Cell Biol 2003; 23: 3103–3115.
Min DS, Ahn BH, Rhie DJ, Yoon SH, Hahn SJ, Kim MS et al. Expression and regulation of phospholipase D during neuronal differentiation of PC12 cells. Neuropharmacology 2001; 41: 384–391.
Acknowledgements
This study was supported by a grant (no. R01-2006-000-10521-0; 2006) from the Basic Research Program of the Korea Science & Engineering Foundation References.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by V De Laurenzi
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Jang, Y., Namkoong, S., Kim, Y. et al. Cleavage of phospholipase D1 by caspase promotes apoptosis via modulation of the p53-dependent cell death pathway. Cell Death Differ 15, 1782–1793 (2008). https://doi.org/10.1038/cdd.2008.111
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.111
Keywords
This article is cited by
-
Phospholipase D2 is a positive regulator of sirtuin 1 and modulates p53-mediated apoptosis via sirtuin 1
Experimental & Molecular Medicine (2021)
-
Phospholipase D1 regulation of TNF-alpha protects against responses to LPS
Scientific Reports (2018)