Abstract
Background:
Nasopharyngeal carcinoma (NPC) has a distinctive geographic distribution and is characterised by its strong tendency of metastasis. We aimed to examine the microRNA (miRNA) expression profiles in plasma samples of NPC patients to explore their clinical significance in disease development and progression.
Methods:
This study was divided into four steps: (1) confirmation of differentially expressed miRNAs using microarray analysis and quantitative PCR validation; (2) comparison of plasma miR-9 levels during NPC progression; (3) evaluation of the predictive performance of plasma miR-9 as a biomarker for NPC metastasis; and (4) comparison of plasma miR-9 levels between pre- and post-treatment samples.
Results:
Plasma microarray profiling identified 33 differentially expressed miRNAs between NPC patients and healthy volunteers. The significantly declined level of miR-9 in NPC patients was confirmed through two-stage validation. The low level of plasma miR-9 was significantly correlated with worse lymphatic invasion and advanced TNM stage. The plasma miR-9 could distinguish locoregional from metastatic NPC cases with a high sensitivity and specificity. Furthermore, the plasma miR-9 level was significantly elevated in post-treatment plasma compared with those pre-treatment samples.
Conclusion:
Our study reports that plasma miR-9 may serve as a useful biomarker to predict NPC metastasis and to monitor tumour dynamics.
Similar content being viewed by others
Main
Nasopharyngeal carcinoma (NPC) is a tumour of the head and neck with a complex aetiology. The incidence is rare in the western world (0.5 per 100 000); however, in the Guangdong region of southeast China the incidence peaks at 50 per 100 000 (Yu and Yuan, 2002). Unlike other head and neck tumours, NPC has a high rate of local or locoregional recurrence or metastasis. Thirty to sixty percent of patients with NPC will eventually develop distant metastasis, which is the key contributor to NPC mortality (Hui et al, 2004). Because of the high incidence and frequent fatal metastasis, NPC is considered as a significant health problem in parts of the world where it is endemic.
Currently, computed tomography (CT) and magnetic resonance (MR) imaging are essential for detection of early NPC, staging of the primary tumour and evaluation of associated retropharyngeal and cervical lymphadenopathy (Abdel Khalek Abdel Razek and King, 2012). However, the imaging findings are inadequate for definition of prognosis because they classify the extent of disease chiefly on the basis of anatomical information. Further studies are urgently needed to identify more effective approaches to monitor and block NPC progression. Previous studies reported that persistently elevated post-treatment plasma Epstein-Barr virus (EBV) DNA level was a robust predictor of relapse in NPC (Lin et al, 2004). Nevertheless, some issues need to be overcome before the plasma EBV DNA assay can become a universal clinical routine, including discrepancies between different institutes and DNA degradation over time (Le et al, 2005).
MicroRNAs (miRNAs) are small non-coding RNAs, which inhibit gene expression by interacting with the 3′ untranslated regions (3′UTR) of target mRNAs (Bartel, 2004). There is a large body of evidence that dysregulation of miRNAs is a hallmark of cancer (Esquela-Kerscher and Slack, 2006). MicroRNA-expression profiling of human tumours has identified signatures associated with diagnosis, staging, progression, prognosis and response to treatment (Calin and Croce, 2006). Recent evidences show that circulating miRNAs might have great potential to serve as novel, non-invasive biomarkers for cancer diagnosis, progression and prognosis (Wittmann and Jack, 2010; Cortez et al, 2011). Although circulating miRNA profiling has been investigated in many cancers, such as lymphoma, breast, prostate, ovarian, pancreatic, gastric, colorectal and lung cancer (Schwarzenbach et al, 2011; Zhang et al, 2012), rare data are available in NPC. More recently, Zeng et al (2012) reported that the serum four-miRNA-based biomarker model might provide a novel strategy for NPC diagnosis. However, whether circulating miRNAs can be used as prognostic biomarkers in NPC remains unknown. Here, we aimed to examine the miRNA expression profiles in plasma samples of NPC patients to explore their clinical significance in disease development and progression, and provide information for personalised therapy.
Materials and methods
Patients and study design
All blood samples of NPC patients (n=294) and healthy control subjects (n=109) were collected from Nanfang Hospital (Southern Medical University, Guangzhou, China) between January 2009 and December 2012. The healthy donors were carefully selected to match the gender and age distribution of NPC patients. None of the patients had received radiotherapy or chemotherapy before blood collecting. The TNM classification for NPC was according to the definitions of the seventh edition of the UICC-American Joint Committee on Cancer staging criteria. Informed consent was obtained from all individuals, and the research protocols were approved by the Ethics Committee of Nanfang Hospital.
The patients received a uniform protocol of image-guided intensity modulated radiotherapy and cisplatin-based concurrent chemotherapy following induction chemotherapy according to the National Comprehensive Cancer Network (NCCN) Guidelines (Version 1. 2012). After the completion of all treatment, patients were followed once every 3 months during the first and second year and every 6 months thereafter. All tumour recurrences and metastases were documented by imaging studies along with pathological verification if the lesions were accessible and the patient agreed.
This study was divided into five stepwise phases (Figure 1). In phase I, miRNA expression profiling was performed in a discovery set (comprising 20 NPC patients and 10 healthy donors). Differentially expressed miRNAs were identified, and nine candidate miRNAs with functional or biomarker-related reports were selected for further validation. In phase II, nine candidate miRNAs were validated using quantitative reverse transcription–polymerase chain reaction (qRT–PCR) on an independent cohort (comprising 40 NPC patients and 15 healthy donors). We then focused on miR-9, one of the most profoundly reduced miRNAs in all NPC individuals. In phase III, miR-9 levels were subsequently validated in a training set (including 150 NPC patients and 54 control subjects). In phase IV, receiver operating characteristic (ROC) curve-based risk assessment analysis was then performed in a validation set (including 84 NPC patients and 30 control subjects). In phase V, further analysis was conducted in an evaluation set (including 40 NPC patients), who provided paired plasma samples of pre-treatment and 3-month post-treatment in the long-term follow-up study, to explore the temporal trend of miR-9 levels.
Plasma collection, RNA isolation and miRNA microarray analysis
Whole blood (8 ml) was drawn into EDTA-containing tubes and separated into plasma and cellular fractions by centrifugation at 1500 g for 5 min. Total plasma RNA was harvested with the TRIzol LS reagent (Invitrogen, Carlsbad, CA, USA) and the RNeasy Mini kit (Qiagen, Valencia, CA, USA) according to the manufacturers’ instructions. In detail, 500 μl plasma was mixed thoroughly with 1.5 ml TRIzol LS reagent, incubated for 5 min at room temperature and subsequently mixed with 400 μl chloroform. The aqueous phase containing RNA was carefully removed, and RNA was precipitated by addition of 100% ethanol. The mixture was applied to an RNeasy Mini spin column and washed several times, and RNA was eluted and stored at −80 °C until further processing.
Small RNA extracted from the plasma samples were labelled using the miRCURY Hy3/Hy5 Power labelling kit and hybridised on the miRCURY LNA Array (Version 16.0, Exiqon, Vedbaek, Denmark). The threshold value for differentially expressed miRNAs was a fold change >1.5 with a value of P<0.05. The raw and normalised miRNA data were available through GEO accession number GSE48442. The candidate miRNAs were further filtered on the basis of expression levels.
Quantitative reverse transcription–PCR analysis of miRNA expression levels
Quantitative reverse transcription–PCR was used to validate the profiling results. Reverse transcription of the total RNA was performed using NCode miRNA First Strand cDNA Synthesis kit (Invitrogen) according to the manufacturer’s protocol. Real-time PCR was performed by using TaqMan miRNA Assay kit (Applied Biosystems Inc., Carlsbad, CA, USA) on an ABI 7500HT System. All reactions were run in triplicate. All samples were normalised to internal control U6 small nuclear RNA and fold changes were calculated through relative quantification (2−ΔΔCt, Livak and Schmittgen, 2001).
Statistical analysis
SPSS 15.0 software was used for statistical analysis (SPSS, Chicago, IL, USA). A Mann–Whitney U-test was performed to identify differentially expressed miRNAs and detect differences of miR-9 levels between NPC and control groups. Characteristics of study participants between the training and validation set were also compared using the Mann–Whitney U-test. The Wilcoxon test was used to compare the paired plasma samples obtained before and after treatment. Significant associations between the miR-9 level and clinicopathological parameters were assessed with a pearson χ2 test. Receiver operating characteristic curve analysis and the area under the curve (AUC) were used to assess the predictive performance of miR-9. Two-sided P-values of <0.05 were considered statistically significant.
Results
Patient characteristics
The characteristics of the study participants were presented in Table 1 and Supplementary Table 1. There was no significant difference in the distribution of age and gender between the training and validation set. The clinical staging also exhibited no statistically significant difference between the training and validation set.
MicroRNA microarray profiling and validation studies using qRT–PCR
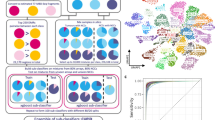
To characterise the miRNA expression signature in NPC patients, we performed microarray analysis. A two-dimensional hierarchical clustering analysis revealed 33 miRNAs that showed significantly altered levels (Figure 2A). Among these 33 miRNAs, 5 were downregulated and 28 were upregulated in the plasma of NPC patients (Supplementary Table 2). There were five miRNAs, including miR-892b, miR-941, miR-3921, miR-4257 and miR-143-3p, with significantly higher levels in NPC patients than in healthy controls (fold change=2.0–3.3, P<0.01). In contrast, miR-9-5p (also known as miR-9) was identified as the most profoundly repressed miRNA in NPC patients (fold change=0.07, P=0.0144).
Global profiling of plasma miRNA in NPC patients. (A) Heat map of miRNA microarray expression between NPC group (n=20) and healthy control group (n=10). Thirty-three miRNAs were identified (P<0.05). A vertical branch showed the expression pattern of the candidate miRNAs in each individual. The relative expression was depicted according to the colour scale above. Red indicated upregulation; green, downregulation. Numbers with t indicated NPC patients; numbers with c indicated healthy control subjects. (B) Validation of deregulated miRNAs by qRT–PCR. An independent validation cohort included 40 NPC patients and 15 healthy control subjects. MicroRNA abundance was normalised to U6 RNA. Quantification was presented as mean values (error bars corresponded to s.d.) relative to control from three independent experiments. *P<0.05, **P<0.01.
To confirm the findings of the miRNA profile, the levels of nine differentially expressed miRNAs were tested using an independent cohort (including 40 NPC patients and 15 healthy volunteers) with qRT–PCR. The relative level of each miRNA was shown in Figure 2B. The results demonstrated a 2.5-fold increase in miR-214-3p, a two-fold increase in miR-3135a and a 90% decrease in the miR-9 level in NPC patients, respectively. Although the levels of miR-92a-2-5p and miR-892b were elevated in miRNA profiling, their levels decreased by 30% and 40% in NPC patients by our qRT–PCR assay, respectively. Among the candidate miRNAs, miR-9 had the lowest level assessed by miRNA profiling and qRT–PCR assay. We then focused on miR-9 in further analysis.
Comparison of miR-9 levels during NPC progression
We further validated miR-9 levels in the plasma samples of the training set using qRT–PCR. MiR-9 level was significantly reduced in NPC patients as compared with normal control subjects (P<0.001, Figure 3A). We also found that the miR-9 level was higher in stage 1 patients, whereas stages 2–4 had lower levels, showing a significant correlation of miR-9 level with clinical stages (P<0.001, Figure 3B). Correlation analysis further demonstrated that low level of miR-9 was positively associated with a more advanced clinical stage of NPC (Table 2). Remarkably, miR-9 level in the subgroup of metastatic NPC patients (n=120) was reduced by 50% compared with the group of locoregional NPC patients (n=30) (P<0.001, Figure 3C). All these data suggested that loss of miR-9 expression was associated with invasive disease and higher tumour stage.
MiR-9 levels during NPC progression. (A) Validation of miR-9 level in the plasma samples of the training set. (B) The plasma level of miR-9 decreased with increasing NPC stage. Each column showed the mean for triplicate experiments; bars, s.d. (C) Examination of miR-9 level in the plasma samples of locoregional NPC patients (n=30) and metastatic NPC patients (n=120).
Validation study of miR-9 as a predictive plasma biomarker for NPC metastasis
We used the data from the training set (which consisted of 120 metastatic and 30 locoregional NPC cases) to construct a ROC curve with an AUC of 0.743 (95% CI=0.619–0.867, sensitivity=73.7% and specificity=74.2%, Supplementary Figure 1). To test the accuracy of the parameters estimated from the training set, we assayed miR-9 level by qRT–PCR in the validation set (which consisted of 63 metastatic and 21 locoregional NPC cases). Using the previously determined cutoff level, we found that 47 of 63 metastatic NPC cases tested positive (sensitivity=74.6%), and that 4 of 21 locoregional NPC cases tested positive (specificity=81.0%).
Dynamic changes of plasma miR-9 level after treatment
The dynamic changes of plasma miR-9 level were detected in an evaluation set (including 40 pre- and 3-month post-treatment paired plasma samples) by qRT–PCR. As shown in Figure 4A, the level of miR-9 in post-treatment plasma was elevated in 37 cases (37 out of 40, 92.5%) compared with that in the pre-treatment sample. Additionally, the level of miR-9 was significantly increased in post-treatment plasma compared with those pre-treatment samples (P<0.001, Figure 4B). In a typical patient, the level of miR-9 was fluctuating over time. As shown in Figure 4C, there was a prominent increase for miR-9 level with the time distance from treatment completion, and re-reduction of the plasma miR-9 level was found when distant metastasis occurred.
Comparison of plasma miR-9 levels in pre- and 3-month post-treatment paired samples. (A) The levels of miR-9 in 40 paired samples were examined by qRT–PCR. The levels of miR-9 in post-treatment plasma samples were higher than those pre-treatment in 92.5% of NPC patients. Each column showed the mean for triplicate experiments; bars, s.d. Marked high data were indicated by numerals and columns were abbreviated. (B) Box plots of plasma miR-9 levels in pre- and 3-month post-treatment paired samples. (C) Changes of the plasma miR-9 level in a typical patient. In one patient, who developed distance metastasis after treatment, re-reduction of plasma miR-9 level was found when distant metastasis occurred.
Discussion
The aim of this study was to identify differentially expressed miRNAs in the plasma of NPC patients, and investigate the potential of plasma miRNA as a biomarker for predicting NPC metastasis. We found the significantly declined level of plasma miR-9 in NPC patients through systematic microarray-based screening and two-stage validation. The plasma miR-9 level could distinguish locoregional from metastatic NPC cases and showed a decreasing trend during NPC progression. To the best of our knowledge, this is the first population-based study in exploring the dynamic changes of plasma miRNAs associated with NPC progression in a high-risk area of NPC.
More recently, Zeng et al (2012) performed microarray-based serum miRNA profiling on the serum of 20 NPC patients and 20 non-cancerous individuals as controls. The profiles showed that 39 and 17 miRNAs were exclusively expressed in the serum of non-cancerous volunteers and NPC patients, respectively. Followed by qRT–PCR validation, 18 serum miRNAs were identified as differentially expressed miRNAs, including miR-92a, miR-106a, miR-143 and miR-17, which also displayed higher levels in NPC plasma on our array, whereas the striking decline of miR-9 in our study was not observed by them. The partially inconsistent results may reflect the differences in sample types, screening tools or quantification methods.
Among the 33 differentially expressed miRNAs, miR-9 had the lowest level assessed by miRNA profiling and qRT–PCR assay. We then focused on miR-9, one of the most profoundly reduced miRNAs in all NPC individuals. In fact, there were many reports of aberrant miR-9 expression in several cancers, suggesting that miR-9 could act as an oncogene or as a tumour suppressor. Upregulation of miR-9 was reported in Hodgkin’s lymphoma (Nie et al, 2008), primary brain tumours (Nass et al, 2009) and endometrial cancer (Myatt et al, 2010), whereas downregulation of miR-9 was observed in cervical (Hu et al, 2010), colorectal (Bandres et al, 2009), non-small-cell lung (Heller et al, 2012) and ovarian cancer (Guo et al, 2009) and in hepatocellular (Tan et al, 2010) and gastric carcinoma (Luo et al, 2009). Decreased miR-9 expression was considered as a poor survival marker in cervical cancer (Hu et al, 2010), lung squamous cell carcinoma (Heller et al, 2012) and acute lymphoblastic leukaemia (Rodriguez-Otero et al, 2011), and it was also associated with the development of cancer metastasis (Lujambio et al, 2008; Hildebrandt et al, 2010; Ma et al, 2010; Tan et al, 2010). Moreover, Shigehara et al (2011) examined the human bile miRNAome and identified miR-9 as a potential diagnostic biomarker for biliary tract cancer. However, the role of miR-9 in NPC is largely unknown. To date, only one study has reported that miR-9 modulates the expression of interferon-regulated genes and MHC class I molecules in NPC cells (Gao et al, 2013).
In our current study, we found that plasma miR-9 could distinguish locoregional from metastatic NPC cases with a high sensitivity and specificity. We were also interested in exploring the associations between plasma miR-9 levels and clinical stages of NPC. By comparing miR-9 levels in the plasma of NPC patients with various stages, we found that advanced stage NPC patients had lower plasma miR-9 levels than those with early stage disease. This result revealed that low level of plasma miR-9 in NPC patients was significantly correlated with worse lymphatic invasion and advanced TNM stage. However, the exact reason for this phenomenon is not clear. Kosaka et al (2010), revealed a secretary machinery of circulating miRNAs and their intercellular transfer, and suggested that these circulating miRNAs might function as a signalling molecule. Further analysis should clarify the origin of extracellular circulating miRNAs and shed light on the causation of the correlations.
By examining the levels of plasma miR-9 in 40 pre- and 3-month post-treatment paired samples, we found that plasma miR-9 level was significantly elevated in post-treatment plasma compared with those pre-treatment samples. This finding suggested that the plasma miR-9 level reflected tumour dynamics to some extent. Concerning monitoring cancer, in a representative patient with distant metastasis, re-reduction of plasma miR-9 level was found when metastasis occurred. Without further information on the origin of plasma miRNAs, any attempt to arrive at an appropriate understanding of the fluctuating miRNA pattern appears premature. One possible explanation is that some miRNAs might be released selectively from cancer cells to stroma and circulation (Ohshima et al, 2010; Pigati et al, 2010), and another possible theory is that the plasma miRNAs might also be released from normal tissues by unknown mechanisms (Chen et al, 2008; Hunter et al, 2008). We speculated that the elevated miR-9 levels after chemo-radiotherapy might be due to the damage of intact cells as a result of chemo-radiotherapy. Those cells might then release their content including nucleic acids in the surroundings and subsequently into the blood stream.
The presented study has several limitations, one being that we lack long-term clinical follow-up data of each patient, which currently limits the ability to explore the prognostic value of miR-9. An expanded patient cohort with long time follow-up will probably validate its prognostic effect. Another limitation is that the results of this preliminary study with small sample size necessitate further confirmation in large prospective studies and the functional relevance of the newly identified miRNAs needs to be further clarified.
In conclusion, this study clearly demonstrated that miR-9 could be a useful blood-based biomarker for predicting NPC metastasis and monitoring tumour dynamics. This non-invasive blood-based biomarker could have great potential to predict the clinical behaviour of NPC and to monitor therapeutic response. Further prospective clinical trials should be carried out to define the benefit of the assay for each potential application.
Accession codes
Change history
21 January 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Abdel Khalek Abdel Razek A, King A (2012) MRI and CT of nasopharyngeal carcinoma. AJR Am J Roentgenol 198: 11–18.
Bandres E, Agirre X, Bitarte N, Ramirez N, Zarate R, Roman-Gomez J, Prosper F, Garcia-Foncillas J (2009) Epigenetic regulation of microRNA expression in colorectal cancer. Int J Cancer 125: 2737–2743.
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: 281–297.
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6: 857–866.
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, Zhang Y, Wang J, Jiang X, Xiang Y, Xu C, Zheng P, Zhang J, Li R, Zhang H, Shang X, Gong T, Ning G, Wang J, Zen K, Zhang J, Zhang CY (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18: 997–1006.
Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA (2011) MicroRNAs in body fluids—the mix of hormones and biomarkers. Nat Rev Clin Oncol 8: 467–477.
Esquela-Kerscher A, Slack FJ (2006) Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 6: 259–269.
Gao F, Zhao ZL, Zhao WT, Fan QR, Wang SC, Li J, Zhang YQ, Shi JW, Lin XL, Yang S, Xie RY, Liu W, Zhang TT, Sun YL, Xu K, Yao KT, Xiao D (2013) miR-9 modulates the expression of interferon-regulated genes and MHC class I molecules in human nasopharyngeal carcinoma cells. Biochem Biophys Res Commun 431: 610–616.
Guo LM, Pu Y, Han Z, Liu T, Li YX, Liu M, Li X, Tang H (2009) MicroRNA-9 inhibits ovarian cancer cell growth through regulation of NF-kappaB1. FEBS J 276: 5537–5546.
Heller G, Weinzierl M, Noll C, Babinsky V, Ziegler B, Altenberger C, Minichsdorfer C, Lang G, Dome B, End-Pfutzenreuter A, Arns BM, Grin Y, Klepetko W, Zielinski CC, Zochbauer-Muller S (2012) Genome-wide miRNA expression profiling identifies miR-9-3 and miR-193a as targets for DNA methylation in non-small cell lung cancers. Clin Cancer Res 18: 1619–1629.
Hildebrandt MA, Gu J, Lin J, Ye Y, Tan W, Tamboli P, Wood CG, Wu X (2010) Hsa-miR-9 methylation status is associated with cancer development and metastatic recurrence in patients with clear cell renal cell carcinoma. Oncogene 29: 5724–5728.
Hu X, Schwarz JK, Lewis JS Jr., Huettner PC, Rader JS, Deasy JO, Grigsby PW, Wang X (2010) A microRNA expression signature for cervical cancer prognosis. Cancer Res 70: 1441–1448.
Hui EP, Leung SF, Au JS, Zee B, Tung S, Chua D, Sze WM, Law CK, Leung TW, Chan AT (2004) Lung metastasis alone in nasopharyngeal carcinoma: a relatively favorable prognostic group. A study by the Hong Kong Nasopharyngeal Carcinoma Study Group. Cancer 101: 300–306.
Hunter MP, Ismail N, Zhang X, Aguda BD, Lee EJ, Yu L, Xiao T, Schafer J, Lee ML, Schmittgen TD, Nana-Sinkam SP, Jarjoura D, Marsh CB (2008) Detection of microRNA expression in human peripheral blood microvesicles. PLoS One 3: e3694.
Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T (2010) Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem 285: 17442–17452.
Le QT, Jones CD, Yau TK, Shirazi HA, Wong PH, Thomas EN, Patterson BK, Lee AW, Zehnder JL (2005) A comparison study of different PCR assays in measuring circulating plasma epstein-barr virus DNA levels in patients with nasopharyngeal carcinoma. Clin Cancer Res 11: 5700–5707.
Lin JC, Wang WY, Chen KY, Wei YH, Liang WM, Jan JS, Jiang RS (2004) Quantification of plasma Epstein-Barr virus DNA in patients with advanced nasopharyngeal carcinoma. N Engl J Med 350: 2461–2470.
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25: 402–408.
Lujambio A, Calin GA, Villanueva A, Ropero S, Sanchez-Cespedes M, Blanco D, Montuenga LM, Rossi S, Nicoloso MS, Faller WJ, Gallagher WM, Eccles SA, Croce CM, Esteller M (2008) A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci USA 105: 13556–13561.
Luo H, Zhang H, Zhang Z, Zhang X, Ning B, Guo J, Nie N, Liu B, Wu X (2009) Down-regulated miR-9 and miR-433 in human gastric carcinoma. J Exp Clin Cancer Res 28: 82.
Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D, Teruya-Feldstein J, Reinhardt F, Onder TT, Valastyan S, Westermann F, Speleman F, Vandesompele J, Weinberg RA (2010) miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol 12: 247–256.
Myatt SS, Wang J, Monteiro LJ, Christian M, Ho KK, Fusi L, Dina RE, Brosens JJ, Ghaem-Maghami S, Lam EW (2010) Definition of microRNAs that repress expression of the tumor suppressor gene FOXO1 in endometrial cancer. Cancer Res 70: 367–377.
Nass D, Rosenwald S, Meiri E, Gilad S, Tabibian-Keissar H, Schlosberg A, Kuker H, Sion-Vardy N, Tobar A, Kharenko O, Sitbon E, Lithwick Yanai G, Elyakim E, Cholakh H, Gibori H, Spector Y, Bentwich Z, Barshack I, Rosenfeld N (2009) MiR-92b and miR-9/9* are specifically expressed in brain primary tumors and can be used to differentiate primary from metastatic brain tumors. Brain Pathol 19: 375–383.
Nie K, Gomez M, Landgraf P, Garcia JF, Liu Y, Tan LH, Chadburn A, Tuschl T, Knowles DM, Tam W (2008) MicroRNA-mediated down-regulation of PRDM1/Blimp-1 in Hodgkin/Reed-Sternberg cells: a potential pathogenetic lesion in Hodgkin lymphomas. Am J Pathol 173: 242–252.
Ohshima K, Inoue K, Fujiwara A, Hatakeyama K, Kanto K, Watanabe Y, Muramatsu K, Fukuda Y, Ogura S, Yamaguchi K, Mochizuki T (2010) Let-7 microRNA family is selectively secreted into the extracellular environment via exosomes in a metastatic gastric cancer cell line. PLoS One 5: e13247.
Pigati L, Yaddanapudi SC, Iyengar R, Kim DJ, Hearn SA, Danforth D, Hastings ML, Duelli DM (2010) Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS One 5: e13515.
Rodriguez-Otero P, Roman-Gomez J, Vilas-Zornoza A, Jose-Eneriz ES, Martin-Palanco V, Rifon J, Torres A, Calasanz MJ, Agirre X, Prosper F (2011) Deregulation of FGFR1 and CDK6 oncogenic pathways in acute lymphoblastic leukaemia harbouring epigenetic modifications of the MIR9 family. Br J Haematol 155: 73–83.
Schwarzenbach H, Hoon DS, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 11: 426–437.
Shigehara K, Yokomuro S, Ishibashi O, Mizuguchi Y, Arima Y, Kawahigashi Y, Kanda T, Akagi I, Tajiri T, Yoshida H, Takizawa T, Uchida E (2011) Real-time PCR-based analysis of the human bile microRNAome identifies miR-9 as a potential diagnostic biomarker for biliary tract cancer. PLoS One 6: e23584.
Tan HX, Wang Q, Chen LZ, Huang XH, Chen JS, Fu XH, Cao LQ, Chen XL, Li W, Zhang LJ (2010) MicroRNA-9 reduces cell invasion and E-cadherin secretion in SK-Hep-1 cell. Med Oncol 27: 654–660.
Wittmann J, Jack HM (2010) Serum microRNAs as powerful cancer biomarkers. Biochim Biophys Acta 1806: 200–207.
Yu MC, Yuan JM (2002) Epidemiology of nasopharyngeal carcinoma. Semin Cancer Biol 12: 421–429.
Zeng X, Xiang J, Wu M, Xiong W, Tang H, Deng M, Li X, Liao Q, Su B, Luo Z, Zhou Y, Zhou M, Zeng Z, Li X, Shen S, Shuai C, Li G, Fang J, Peng S (2012) Circulating miR-17, miR-20a, miR-29c, and miR-223 combined as non-invasive biomarkers in nasopharyngeal carcinoma. PLoS One 7: e46367.
Zhang J, Zhao H, Gao Y, Zhang W (2012) Secretory miRNAs as novel cancer biomarkers. Biochim Biophys Acta 1826: 32–43.
Acknowledgements
This work was supported by National Natural Science Foundation of China (U1132003, 81172053 to XPL; 81272197, 31070953 to YP; 81301849 to JL) and National Basic Science Research Program (2013CB911302 to MLH).
Author information
Authors and Affiliations
Corresponding authors
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Lu, J., Xu, X., Liu, X. et al. Predictive value of miR-9 as a potential biomarker for nasopharyngeal carcinoma metastasis. Br J Cancer 110, 392–398 (2014). https://doi.org/10.1038/bjc.2013.751
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.751
Keywords
This article is cited by
-
RETRACTED ARTICLE: Chrysophanol suppresses growth and metastasis of T cell acute lymphoblastic leukemia via miR-9/PD-L1 axis
Naunyn-Schmiedeberg's Archives of Pharmacology (2020)
-
Exosomal miR-9 inhibits angiogenesis by targeting MDK and regulating PDK/AKT pathway in nasopharyngeal carcinoma
Journal of Experimental & Clinical Cancer Research (2018)
-
BPIFB1 (LPLUNC1) inhibits migration and invasion of nasopharyngeal carcinoma by interacting with VTN and VIM
British Journal of Cancer (2018)
-
Prognostic Value of microRNA-9 in Various Cancers: a Meta-analysis
Pathology & Oncology Research (2017)
-
RBM24 suppresses cancer progression by upregulating miR-25 to target MALAT1 in nasopharyngeal carcinoma
Cell Death & Disease (2016)