Abstract
Aim:
Cytochrome P450 oxidoreductase (POR) is the only flavoprotein that donates electrons to all microsomal P450 enzymes (CYP), and several POR SNPs have been shown to be important contributors to altered CYP activity or CYP-mediated drug metabolism. In this study we examined the association between 6 POR SNPs and tacrolimus concentrations in Chinese renal transplant recipients.
Methods:
A total of 154 renal transplant recipients were enrolled. Genotyping of CYP3A5*3 and 6 POR SNPs was performed. All patients received a triple immunosuppressive regimen comprising tacrolimus, mycophenolate mofetil and prednisone. Dose-adjusted tacrolimus trough concentrations were obtained on d 7 (C0D7/D) after transplantation when steady-state concentration of tacrolimus was achieved (dosage had been unchanged for more than 3 d).
Results:
Tacrolimus C0D7/D in CYP3A5*3/*3/ POR rs1057868–rs2868177 GC-GT diplotype carriers was 1.62- and 2.72-fold higher than those in CYP3A5*3/*3/ POR rs1057868–rs2868177 GC-GT diplotype non-carriers and CYP3A5*1 carriers (220.17±48.09 vs 135.69±6.86 and 80.84±5.27 ng/mL/mg/kg, respectively, P<0.0001). Of CYP3A5*3/*3/ POR rs1057868-rs2868177GC-GT diplotype carriers, 85.71% exceeded the upper limit of the target range (8 ng/mL), which was also significantly higher compared with the latter two groups (14.29% and 0.00%, respectively, P<0.0001). The CYP3A5*3 and POR rs1057868–rs2868177 GC-GT diplotype explained 31.7% and 5.7%, respectively, of the inter-individual variability of tacrolimus C0D7/D, whereas the POR rs1057868–rs2868177 GC-GT diplotype could explain 10.9% of the inter-individual variability of tacrolimus C0D7/D in CYP3A5 non-expressers.
Conclusion:
The CYP3A5*3 and POR rs1057868–rs2868177 GC-GT diplotype accounted for the inter-individual variation of tacrolimus C0D7/D. Genotyping of POR rs1057868–rs2868177 diplotypes would help to differentiate initial tacrolimus dose requirements and to achieve early target C0 ranges in Chinese renal transplant recipients.
Similar content being viewed by others
Introduction
The calcineurin inhibitor tacrolimus (FK506; Prograf) is the first-line immunosuppressant that is widely used in solid-organ transplant recipients for prophylaxis against allograft rejection1. However, the narrow therapeutic window (trough concentration should be controlled in the range of 5 to 8 ng/mL in the first 3 months post-transplant in our routine2) and large inter-individual pharmacokinetic variability are the major therapeutic challenges. Over-immunosuppression causes toxicity and under-immunosuppression leads to rejection, characterizing the highest risks during the initial period, especially within the 7 d post-transplantation3. Therefore, identifying key predictors for earlier tacrolimus concentration and achieving individualized dosing have great potential for improving the procedure's safety and efficacy.
Tacrolimus is predominantly metabolized by CYP3A5, and its intrinsic clearance is 1.6- to two-fold higher for CYP3A5 than for CYP3A44. It is now generally accepted that tacrolimus pharmacokinetics is closely correlated with a nonfunctional splicing-defect in CYP3A5*3 allele (rs776746; 6985A>G)5,6,7. CYP3A5*3/*3 carriers (CYP3A5 function deficient, denominated as CYP3A5 nonexpressers) require a lower dose of tacrolimus to reach target concentrations compared with CYP3A5*1 allele carriers (CYP3A5 expressers). However, CYP3A5*3 genotype can explain only one-third of the inter-individual variability in tacrolimus pharmacokinetics6. Moreover, there are still obvious differences among both CYP3A5 nonexpressers and expressers. Hence, exploring the unaccounted genetic variation in tacrolimus pharmacokinetics on top of what is already known on CYP3A5 variants is needed.
Cytochrome P450 oxidoreductase (POR) is known as the unique electron donor to all microsomal cytochrome P450 (CYP) enzymes in human8. Because POR's activity is necessary for CYP functions, it is reasonable to infer that genetic variations in POR gene could affect the functions of broad ranges of CYPs and thus alter therapeutic efficiency and toxicity of many drugs. The contribution of POR*28 (rs1057868C>T; A503V) to inter-individual variability of tacrolimus metabolism has been studied, but the results are conflicting. In addition to POR*28, several other POR SNPs have also been reported to be associated with altered cytochrome P450 activity9,10,11,12. Because the distribution profile of POR SNPs in the Chinese population has rarely been reported, we performed a pilot study to assess the variant frequencies of 11 functional candidate SNPs in 96 healthy Chinese volunteers. The results are shown in Supplementary Table S1. Five SNPs—rs2302429, rs17685, rs2868177, rs2286823, and rs41301394—were prevalent in our healthy volunteers. Intronic SNPs-rs2302429, rs2286823 (IVS11±20G>A) and rs41301394 (831-35C>T) have been significantly associated with the altered CYP1A2, CYP2C19, or CYP3A4 activities13,14. Rs2868177 has been found to be associated with the cardiotoxicity induced by daunorubicin, which is also a substrate of CYP3A15. Moreover, rs2868177 and other two SNPs (rs17148944 and rs17685) have been brought into warfarin pharmacogenetic research and reported to be correlated with warfarin maintenance dose16. Interestingly, a number of POR genetic variants have been shown to have substrate-dependent effects on CYP enzyme activities, probably owing to conformational changes induced by the substrate, depending on its size, electrical charge and chemical structure17. It suggests that although not all the SNPs mentioned above were related to CYP3A activity in the previous studies, they might influence CYP3A-mediated tacrolimus metabolism; a comprehensive investigation is needed.
The objective of this study was to comprehensively evaluate the influence of the popular studied POR*28 and other 5 common POR SNPs on tacrolimus concentrations during the initial period after transplantation in a group of Chinese renal-transplant recipients.
Materials and methods
Ethics statement
The study was performed in accordance with the Declaration of Helsinki and guidelines on good clinical practice, and ethical approval of this study was obtained from the ethics committee of the First Affiliated Hospital of Sun Yat-Sen University (No 200823). Written informed consent was obtained from all subjects before participation.
Patients and therapy
We reviewed the medical records and laboratory results of patients who underwent renal transplantation in the Kidney Transplant Department of the First Affiliated Hospital of Sun Yat-sen University between July 2008 and December 2013 and received maintenance treatment with tacrolimus afterward. Adult male and female recipients undergoing single primary renal transplantation were eligible. Patients who received medication known to affect tacrolimus blood levels other than prednisone, such as verapamil, ketoconazole, itraconazole, erythromycin, clarithromycin, or diltiazem, were excluded. Patients with abnormal hepatic function and combined organ transplantations were also excluded. A total of 154 renal transplant recipients were enrolled.
All patients received a triple immunosuppressive regimen comprising tacrolimus (Prograft™, Astellas, Killorglin, Ireland); mycophenolate mofetil (Cellcept™, Roche, Basel, Switzerland), 1.0–1.5 g per day; and prednisone (Guangdong Huanan Pharmacy Ltd, Dongguan, China), 30 mg per day. According to the routine in the Kidney Transplant Department at the First Affiliated Hospital of Sun Yat-sen University, the initial dose of tacrolimus (0.05–0.075 mg/kg twice daily) was given from the morning of the first day post-transplant, and doses were subsequently adjusted to achieve a target trough concentration (C0) of 5–8 ng/mL in the first 3 months post-transplant.
Data collection
Body weight, tacrolimus dosage and whole-blood trough concentrations (C0) were obtained on d 7 after transplantation when steady-state concentration of tacrolimus was achieved (dosage had been unchanged for more than 3 d). Venous blood samples (2 mL) were collected before 8:00 AM, prior to administering the morning dose. The quantification of tacrolimus in human whole blood was achieved by chemiluminescent microparticle immuno assay of Abbott ARCHITECT tacrolimus assay (AbbottDiagn, Chicago, IL, USA). The dose-adjusted tacrolimus trough concentration on d 7 after transplantation is the ratio of the measured tacrolimus trough concentration divided by dose expressed as mg/kg body weight (C0D7/D). The corresponding laboratory parameters including alanine aminotransferase, aspartate aminotransferase and serum creatinine were also obtained.
DNA extraction and genotyping
After informed consent had been obtained, 2 mL venous blood samples for genotyping were collected in the outpatient clinic. Total genomic DNA was extracted from the peripheral leukocytes according to a previously described method18. CYP3A5*3 was determined by using published polymerase chain reaction restriction-fragment length polymorphism (PCR-RFLP) methods5,19. 6 POR SNPs were detected by Agena Bioscience MassARRAY® system (Agena Bioscience, San Diego, CA, USA).
Statistical analysis
The Hardy-Weinberg equilibrium test was performed using χ2 test or Fisher's exact test (two-sided). Linkage disequilibrium (LD) maps were constructed using online software SHEsis20. Haplotypes were statistically inferred using an algorithm based on Bayesian inference by the program PHASE 2.121. For analysis of continuous pharmacologic variables, we used patient genotypes as categorical independent variables. Mann-Whitney U-test was used for comparisons between two groups and the Kruskal-Wallis H-test for comparisons among several groups. Multiple linear regression models considering the contribution of the genetic factors on tacrolimus C0D7/D were evaluated using a stepwise variable selection method; only the variables with a P value of less than 0.1 in the univariate analysis were included in the multivariate analysis. Tacrolimus C0D7/D ratios were log-transformed to reduce skewness of the distribution. The statistical significance of the differences of the target tacrolimus C0 achievement ratio between groups was calculated by χ2 test or Fisher's exact test (two-sided). Statistical analysis was performed using SPSS (Statistical Package for the Social Sciences) software (version 21; SPSS, IBM, Armonk, NY, USA). Data are expressed as the median and range or mean±SD, depending on data type. Statistical power of the sample size was calculated by using software PASS (Power Analysis and Sample Size) software (version 11.0.7; PASS, NCSS, LLC). All the results met the requirement to have more than 80% power to detect the difference within/between groups with two-sided type one error at 5%.
Results
Patient characteristics and POR polymorphisms
The average age of the patients was 40.00±10.93 years (range 18–68 years), and the average body weight was 59.80±10.67 kg (range 35–105 kg). Age, gender, hepatic function and renal function were not found to be correlated with tacrolimus C0D7/D (data not shown).
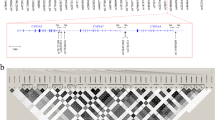
The MA(F) (minor allele frequency) of CYP3A5*3, POR rs1057868, rs2302429, rs17685, rs2868177, rs2286823, and rs41301394 were 25.2%, 34.3%, 30.8%, 39.3%, 38.7%, 40.1%, and 45.0%, respectively. All SNPs were in Hardy-Weinberg equilibrium. Genotype frequencies of CYP3A5*3 and POR rs1067868 were in agreement with those reported in previous publications in Chinese healthy volunteers or patients6,21.
There was a linkage disequilibrium between POR rs2868177 and rs1057868. The r2 and |D′| values between the two SNPs were 0.189 and 0.651, respectively. Haplotypes were constructed, and the frequencies were 25.4% for A–C, 29.3% for A–T, 39.5% for G–C and 5.8% for G–T, respectively.
Influence of CYP3A5*3, POR genotypes on tacrolimus C0D7/D
A total of 7 SNPs were examined. In accord with our earlier study6, CYP3A5 nonexpressers had a significantly higher tacrolimus C0D7/D compared with CYP3A5 expressers [137.96±8.40 vs 89.16±7.02 (ng·mL−1)/(mg·kg−1), P<0.0001] (Figure 1), and stratification analysis was performed to eliminate the confounding effect of CYP3A5*3.
Influence of (A) CYP3A5*3, (B) POR rs2868177 and (C) POR rs1057868–rs2868177 GC-GT diplotypes on tacrolimus trough concentration on d 7 after transplantation. *P<0.05, **P<0.01, ***P<0.001. C0D7/D: Dose-adjusted tacrolimus trough concentration on d 7 after transplantation.
A significantly higher tacrolimus C0D7/D was observed in POR rs2868177 GG genotype carriers than in carriers of AA plus AG genotypes [147.32±18.21 vs 105.95±4.84 (ng·mL−1/(mg·/kg−1), P=0.041] (Figure 1). None of the other 5 POR SNPs demonstrated a significant association with C0D7/D, with or without stratifying for the CYP3A5*3 (Table 1).
Influence of POR haplotypes on tacrolimus C0D7/D
When the effects of POR rs1057868 and rs2868177 were combined, the C0D7/D in each diplotype of POR rs2868177–rs1057868, with or without stratifying for the CYP3A5*3, were as shown in Table 2. Carriers of the POR 1057868–rs2868177 GC-GT diplotype had a considerably higher tacrolimus C0D7/D than noncarriers [220.17±48.09 vs 109.88±5.15 (ng·mL−1)/(mg·kg−1), P=0.002) in the total cohort (Figure 1). The POR 1057868-rs2868177 GC-GT diplotype was not detected in CYP3A5 expressers. In CYP3A5 nonexpressers, tacrolimus C0D7/D was also significantly higher in POR 1057868–rs2868177 GC-GT diplotype carriers than in noncarriers [220.17±48.09 vs 135.69±6.86 (ng·mL−1)/(mg·kg−1, P=0.01].
Multiple variable analysis for association with tacrolimus C0D7/D
When the separated POR SNPs were considered, a multiple linear regression model for log-transformed tacrolimus C0D7/D was developed that included CYP3A5*3 and POR rs2868177. In the final model, shown in Table 3, CYP3A5*3 accounted for 33.7% (P<0.001) of total variation in tacrolimus C0D7/D, whereas POR rs2868177 was excluded.
When POR haplotype was considered, the CYP3A5*3 and POR 1057868–rs2868177 GC-GT diplotype explained 31.7% (P<0.001) and 5.7% (P=0.014) of inter-individual variability of tacrolimus C0D7/D, respectively, and together explained 36.9% of the variability in the total cohort (Table 3). In CYP3A5 nonexpressers, the POR GC-GT diplotype accounted for 10.9% of total variation in log-transformed tacrolimus C0D7/D (P=0.006) (Table 3).
Combined influence of CYP3A5*3 and POR diplotypes on tacrolimus C0D7/D
Considering the combined effects of CYP3A5*3 and POR rs1057868–rs2868177 diplotypes, we divided patients into three groups: fast metabolizers (CYP3A5*1 carriers), slow metabolizers (CYP3A5*3/*3/POR rs1057868–rs2868177 GC-GT diplotype noncarriers) and ultra-slow metabolizers (CYP3A5*3/*3/POR rs1057868–rs2868177 GC-GT diplotype carriers).
Tacrolimus C0D7/D was 220.17±48.09 ng/mL/mg/kg for ultra-slow metabolizers, which was 1.62-fold higher than for slow metabolizers (135.69±6.86 ng/mL/mg/kg, P=0.01) and 2.72-fold higher than for fast metabolizers (80.84±5.27 ng/mL/mg/kg, P<0.0001) (Figure 2). Achievement of target ranges was analyzed on d 7 after transplantation; tacrolimus trough concentration in ultra-slow metabolizers (10.92±2.90 ng/mL) was also significantly higher than that in slow metabolizers (7.29±2.02 ng/mL, P=0.004) and in fast metabolizers (5.46±2.04 ng/mL, P<0.0001). A total of 39.6% of fast metabolizers did not achieve the lower limit of target tacrolimus C0 of 5 ng/mL compared with 7.4% of slow metabolizers and 0% of ultra-slow metabolizers. Conversely, 85.71% of ultra-slow metabolizers exceeded the upper limit of the target range (8 ng/mL) in comparison with 29.63% of slow metabolizers and 10.42% of fast metabolizers (P<0.0001) (Table 4).
Influence of CYP3A5*3 and POR rs1057868–rs2868177 GC-GT diplotype combinations on tacrolimus trough concentration on d 7 after transplantation. **P<0.01, ***P<0.001. Fast metabolizers: CYP3A5*1 carriers. Slow metabolizers: CYP3A5*3/*3/POR rs1057868-rs2868177 GC-GT diplotype noncarriers. Ultra-slow metabolizers: CYP3A5*3/*3/ POR rs1057868–rs2868177 GC-GT diplotype carriers. C0D7/D: Dose-adjusted tacrolimus trough concentration on d 7 after transplantation.
Discussion
In the current study, the associations of 6 POR SNPs with tacrolimus C0D7/D were examined. The positive role of POR rs1057868-rs2868177 haplotypes was first observed, which could explain 5.7% and 10.9% of inter-individual variability of tacrolimus C0D7/D in the entire cohort and in CYP3A5 nonexpressers, respectively. POR rs1057868–rs2868177 GC-GT diplotype carriers also represent a higher-risk group because they have a significantly higher likelihood of exceeding the upper limit of the target range. Therefore, genotyping of POR rs1057868–rs2868177 diplotypes, would help to further differentiate initial tacrolimus dose requirements and control the toxicity of over-immunosuppression.
POR is the unique electron donor to all microsomal CYPs, and the possibility of POR as a potential rate-limiting step in CYPs-mediated drug metabolism has been considered since 196922. Thus, the contribution of common POR variants to variability of tacrolimus metabolism is of great interest. POR*28, the most common sequence variant in POR gene, induces an amino acid substitution (C>T, p.Ala503Val) and influence the electron binding moiety of POR23. A previous in vitro study showed that this mutation could decrease CYP3A4 activity. By contrast, POR*28 has been reported to be associated with lower tacrolimus concentrations in CYP3A5 expressers24,25 or nonexpressers26,27, suggesting that POR*28 variant may enhance CYP3A4 or CYP3A5 enzyme activity. Moreover, no significant association between POR*28 and tacrolimus concentration was found in the present study, which is in line with a study also conducted in Chinese early post-renal transplant recipients28 and two studies conducted in Caucasian early or stable renal transplant recipients15,29. The reason for the discrepancy among these studies is not clear, but several possibilities may be involved. First, different doses or types of corticosteroids were co-administered with tacrolimus in different studies. For instance, prednisone is a CYP3A and P-gp inducer30,31, whereas prednisolone is a weak competitive inhibitor of CYP3A432. Thus the different influences on tacrolimus metabolism between prednisone and prednisolone may interfere with the analysis of the effect of POR*28. Furthermore, the dose of co-administered corticosteroids is reduced gradually during the post-transplant period; thus, the extent of drug-drug interaction between corticosteroids and tacrolimus may also be a confounding factor. Second, Saito et al33 found that there were significant interethnic differences in the haplotype profile of the POR gene containing POR*28. POR*28 may be associated with other functional polymorphisms, and the interethnic haplotype difference may lead to different results. We speculated that these confounding effects may complicate the influence of POR*28 on tacrolimus, and further investigation is required.
The tacrolimus C0D7/D was higher in POR rs2868177 GG genotype carriers than in AA plus AG carriers (P=0.041). We speculated that POR rs2868177 GG genotype may be associated with decreased in vivo CYP3A activity and lead to increased tacrolimus concentration. In a genome-wide association study (GWAS) in acute myeloid leukemia patients, the minor allele (G) was found to be associated with a higher incidence of cardiotoxicity induced by daunorubicin, which is also a substrate of CYP3A, suggesting that the G allele could lead to slower daunorubicin metabolism34. However, the underlying mechanism needs further clarification.
Interestingly, when the effects of POR rs1057868 and rs2868177 were combined, a more obvious correlation was observed, namely, that tacrolimus C0D7/D in carriers of the POR 1057868–rs2868177 GC-GT diplotype was approximately two-fold higher than in noncarriers (220.17±48.09 vs 109.88±5.15 (ng·mL−1)/(mg·kg−1, P=0.002). Moreover, in the multiple linear regression analysis, the POR rs1057868–rs2868177 GC-GT diplotype was the only contributor accounting for the inter-individual variation of tacrolimus C0D7/D besides CYP3A5*3. Such findings suggest that haplotype analysis may be superior to SNP analysis in describing genotype-phenotype associations; this was confirmed in MDR1 pharmacogenetic studies19,35,36. However, further study with a larger sample size is warranted.
Among CYP3A5 nonexpressers, the POR 1057868–rs2868177 GC-GT diplotype could explain 10.9% of total variation in tacrolimus C0D7/D. Because the metabolism of tacrolimus depends mainly on CYP3A4 activity in CYP3A5 nonexpressers6, we presumed that the POR 1057868–rs2868177 GC-GT diplotype might be associated with decreased CYP3A4 enzymatic activity with tacrolimus as a substrate. Further studies identifying the functional characterization of the POR rs1057868–rs2868177 diplotypes and their impacts on the metabolism of tacrolimus are also needed.
It is now generally accepted that patients who carry CYP3A5*3/*3 genotype are slow metabolizers for tacrolimus. However, in this study, we separated a new sub-group from conventional slow metabolizers: CYP3A5*3/*3 and POR GC-GT diplotype carriers, who have been found to have the highest C0D7/D and need 38.2% and 63.2% dosage reduction to obtain the same level of C0 as “slow metabolizers” and “fast metabolizers,” respectively. Therefore, the term “ultra-slow metabolizers” was used to dissociate this group from the conventional “slow metabolizers.” Moreover, 85.71% of ultra-slow metabolizers exceeded the upper limit of the target range (8 ng/mL) despite concentration-controlled dose adjustments. It has been found that tacrolimus C0 recorded within 7 d post-transplantation was significantly associated with rates of rejection and toxicity and a significant trend for increasing toxicity with increasing maximum tacrolimus C03. Therefore, timely dosage reduction is in urgent need for the ultra-slow metabolizers who have the highest risk of over-immunosuppression and toxicity, and identifying the POR rs1057868–rs2868177 diplotype before tacrolimus administration may have great potential for improving the procedure's safety.
In conclusion, this observational pharmacogenetic analysis demonstrated for the first time that POR rs1057868–rs2868177 GC-GT diplotype significantly increased tacrolimus C0D7/D in recipients in the early stage after renal transplantation. Patients who are CYP3A5 nonexpressers and carry POR rs1057868–rs2868177 GC-GT diplotype might be at an elevated risk of early tacrolimus overexposure and need timely dosage adjustment. The results could be useful for personalized medicine for organ-transplant recipients.
Author contribution
Jia-li LI and Chang-xi WANG were involved in study design and manuscript writing, and they are the corresponding authors; Shu LIU was involved in study design, research performance, data analysis and manuscript writing; Qian FU and Jun LI were involved in data collection and analysis; Yu ZHANG, Xue-ding WANG, Ling-yan CHEN, and Xiao-man LIU were involved in laboratory analysis; Rong-xin CHEN, Min HUANG and Hong-bing HUANG were involved in data analysis.
References
Peterson LB, Cryan JG, Rosa R, Martin MM, Wilusz MB, Sinclair PJ, et al. A tacrolimus-related immunosuppressant with biochemical properties distinct from those of tacrolimus. Transplantation 1998; 65: 10–8.
Chen SY, Li JL, Meng FH, Wang XD, Liu T, Li J, et al. Individualization of tacrolimus dosage basing on cytochrome P450 3A5 polymorphism — a prospective, randomized, controlled study. Clin Transplant 2013; 27: E272–81.
Laskow DA, Vincenti F, Neylan JF, Mendez R, Matas AJ . An open-label, concentration-ranging trial of FK506 in primary kidney transplantation: a report of the United States Multicenter FK506 Kidney Transplant Group. Transplantation 1996; 62: 900–5.
Kamdem LK, Streit F, Zanger UM, Brockmoller J, Oellerich M, Armstrong VW, et al. Contribution of CYP3A5 to the in vitro hepatic clearance of tacrolimus. Clin Chem 2005; 51: 1374–81.
Li JL, Wang XD, Chen SY, Liu LS, Fu Q, Chen X, et al. Effects of diltiazem on pharmacokinetics of tacrolimus in relation to CYP3A5 genotype status in renal recipients: from retrospective to prospective. Pharmacogenomics J 2011; 11: 300–6.
Li JL, Liu S, Fu Q, Zhang Y, Wang XD, Liu XM, et al. Interactive effects of CYP3A4, CYP3A5, MDR1 and NR1I2 polymorphisms on tracrolimus trough concentrations in early postrenal transplant recipients. Pharmacogenomics 2015; 16: 1355–65.
Haufroid V, Wallemacq P, VanKerckhove V, Elens L, De Meyer M, Eddour DC, et al. CYP3A5 and ABCB1 polymorphisms and tacrolimus pharmacokinetics in renal transplant candidates: guidelines from an experimental study. Am J Transplant 2006; 6: 2706–13.
Hubbard PA, Shen AL, Paschke R, Kasper CB, Kim JJ . NADPH-cytochrome P450 oxidoreductase. Structural basis for hydride and electron transfer. J Biol Chem 2001; 276: 29163–70.
Chen X, Pan LQ, Naranmandura H, Zeng S, Chen SQ . Influence of various polymorphic variants of cytochrome P450 oxidoreductase (POR) on drug metabolic activity of CYP3A4 and CYP2B6. PLoS One 2012; 7: e38495.
Bonina TA, Gilep AA, Estabrook RW, Usanov SA . Engineering of proteolytically stable NADPH-cytochrome P450 reductase. Biochemistry (Mosc) 2005; 70: 357–65.
Agrawal V, Huang N, Miller WL . Pharmacogenetics of P450 oxidoreductase: effect of sequence variants on activities of CYP1A2 and CYP2C19. Pharmacogenet Genomics 2008; 18: 569–76.
Miller WL, Agrawal V, Sandee D, Tee MK, Huang N, Choi JH, et al. Consequences of POR mutations and polymorphisms. Mol Cell Endocrinol 2011; 336: 174–9.
Gomes AM, Winter S, Klein K, Turpeinen M, Schaeffeler E, Schwab M, et al. Pharmacogenomics of human liver cytochrome P450 oxidoreductase: multifactorial analysis and impact on microsomal drug oxidation. Pharmacogenomics 2009; 10: 579–99.
Dobrinas M, Cornuz J, Pedrido L, Eap CB . Influence of cytochrome P450 oxidoreductase genetic polymorphisms on CYP1A2 activity and inducibility by smoking. Pharmacogenet Genomics 2012; 22: 143–51.
Kurzawski M, Malinowski D, Dziewanowski K, Drozdzik M . Impact of PPARA and POR polymorphisms on tacrolimus pharmacokinetics and new-onset diabetes in kidney transplant recipients. Pharmacogenet Genomics 2014; 24: 397–400.
Zhang X, Li L, Ding X, Kaminsky LS . Identification of cytochrome P450 oxidoreductase gene variants that are significantly associated with the interindividual variations in warfarin maintenance dose. Drug Metab Dispos 2011; 39: 1433–9.
Agrawal V, Choi JH, Giacomini KM, Miller WL . Substrate-specific modulation of CYP3A4 activity by genetic variants of cytochrome P450 oxidoreductase. Pharmacogenet Genomics 2010; 20: 611–8.
Loparev VN, Cartas MA, Monken CE, Velpandi A, Srinivasan A . An efficient and simple method of DNA extraction from whole blood and cell lines to identify infectious agents. J Virol Methods 1991; 34: 105–12.
Zhang Y, Li JL, Fu Q, Wang XD, Liu LS, Wang CX, et al. Associations of ABCB1, NFKB1, CYP3A, and NR1I2 polymorphisms with cyclosporine trough concentrations in Chinese renal transplant recipients. Acta Pharmacol Sin 2013; 34: 555–60.
Shi YY, He L . SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res 2005; 15: 97–8.
Zhang JJ, Zhang H, Ding XL, Ma S, Miao LY . Effect of the P450 oxidoreductase 28 polymorphism on the pharmacokinetics of tacrolimus in Chinese healthy male volunteers. Eur J Clin Pharmacol 2013; 69: 807–12.
Ullrich V . On the hydroxylation of cyclohexane in rat liver microsomes. Hoppe Seylers Z Physiol Chem 1969; 350: 357–65.
Huang N, Agrawal V, Giacomini KM, Miller WL . Genetics of P450 oxidoreductase: sequence variation in 842 individuals of four ethnicities and activities of 15 missense mutations. Proc Natl Acad Sci U S A 2008; 105: 1733–8.
de Jonge H, Metalidis C, Naesens M, Lambrechts D, Kuypers DR . The P450 oxidoreductase *28 SNP is associated with low initial tacrolimus exposure and increased dose requirements in CYP3A5-expressing renal recipients. Pharmacogenomics 2011; 12: 1281–91.
Gijsen VM, van Schaik RH, Soldin OP, Soldin SJ, Nulman I, Koren G, et al. P450 oxidoreductase *28 (POR*28) and tacrolimus disposition in pediatric kidney transplant recipients — a pilot study. Ther Drug Monit 2014; 36: 152–8.
Elens L, Hesselink DA, Bouamar R, Budde K, de Fijter JW, De Meyer M, et al. Impact of POR*28 on the pharmacokinetics of tacrolimus and cyclosporine A in renal transplant patients. Ther Drug Monit 2014; 36: 71–9.
Pulk RA, Schladt DS, Oetting WS, Guan W, Israni AK, Matas AJ, et al. Multigene predictors of tacrolimus exposure in kidney transplant recipients. Pharmacogenomics 2015; 16: 841–54.
Li CJ, Li L, Lin L, Jiang HX, Zhong ZY, Li WM, et al. Impact of the CYP3A5, CYP3A4, COMT, IL-10 and POR genetic polymorphisms on tacrolimus metabolism in Chinese renal transplant recipients. PLoS One 2014; 9: e86206.
Bruckmueller H, Werk AN, Renders L, Feldkamp T, Tepel M, Borst C, et al. Which genetic determinants should be considered for tacrolimus dose optimization in kidney transplantation a combined analysis of genes affecting the CYP3A locus. Ther Drug Monit 2015; 37: 288–95.
Pichard L, Fabre I, Daujat M, Domergue J, Joyeux H, Maurel P . Effect of corticosteroids on the expression of cytochromes P450 and on cyclosporin A oxidase activity in primary cultures of human hepatocytes. Mol Pharmacol 1992; 41: 1047–55.
Joy MS, Nickeleit V, Hogan SL, Thompson BD, Finn WF . Calcineurin inhibitor-induced nephrotoxicity and renal expression of P-glycoprotein. Pharmacotherapy 2005; 25: 779–89.
Lam S, Partovi N, Ting LS, Ensom MH . Corticosteroid interactions with cyclosporine, tacrolimus, mycophenolate, and sirolimus: fact or fiction? Ann Pharmacother 2008; 42: 1037–47.
Saito Y, Yamamoto N, Katori N, Maekawa K, Fukushima-Uesaka H, Sugimoto D, et al. Genetic polymorphisms and haplotypes of POR, encoding cytochrome p450 oxidoreductase, in a Japanese population. Drug Metab Pharmacokinet 2011; 26: 107–16.
Lubieniecka JM, Graham J, Heffner D, Mottus R, Reid R, Hogge D, et al. A discovery study of daunorubicin induced cardiotoxicity in a sample of acute myeloid leukemia patients prioritizes P450 oxidoreductase polymorphisms as a potential risk factor. Front Genet 2013; 4: 231.
Johne A, Kopke K, Gerloff T, Mai I, Rietbrock S, Meisel C, et al. Modulation of steady-state kinetics of digoxin by haplotypes of the P-glycoprotein MDR1 gene. Clin Pharmacol Ther 2002; 72: 584–94.
Chowbay B, Cumaraswamy S, Cheung YB, Zhou Q, Lee EJ . Genetic polymorphisms in MDR1 and CYP3A4 genes in Asians and the influence of MDR1 haplotypes on cyclosporin disposition in heart transplant recipients. Pharmacogenetics 2003; 13: 89–95.
Acknowledgements
The authors appreciate the financial support provided by National Major Projects for Science and Technology Development from the Science and Technology Ministry of China (grant No 2012ZX09506001-004), the National Natural Science Foundations of China (grants 81102515 and 81320108027), the Key Laboratory Foundation of Guangdong Province (grant No 2011A060901014) and the Major Scientific and Technological Project of Guangdong Province (grants 2011A080300001 and 2012A080202013).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary information is available on the website of Acta Pharmacologica Sinica.
Supplementary information
Supplementary Table S1
Comparisons of 11 detected POR SNPs allelic frequencies between Chinese Han population and existing data. (DOC 38 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Liu, S., Chen, Rx., Li, J. et al. The POR rs1057868–rs2868177 GC-GT diplotype is associated with high tacrolimus concentrations in early post-renal transplant recipients. Acta Pharmacol Sin 37, 1251–1258 (2016). https://doi.org/10.1038/aps.2016.77
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2016.77
Keywords
This article is cited by
-
Effects of CYP3A4*22 and POR*28 variations on the pharmacokinetics of tacrolimus in renal transplant recipients: a meta-analysis of 18 observational studies
BMC Nephrology (2024)
-
Population Pharmacokinetic Analysis for Model-Based Therapeutic Drug Monitoring of Tacrolimus in Chinese Han Heart Transplant Patients
European Journal of Drug Metabolism and Pharmacokinetics (2023)
-
Impact of single nucleotide polymorphisms on P450 oxidoreductase and peroxisome proliferator-activated receptor alpha on tacrolimus pharmacokinetics in renal transplant recipients
The Pharmacogenomics Journal (2019)
-
Whole exome sequencing for the identification of CYP3A7 variants associated with tacrolimus concentrations in kidney transplant patients
Scientific Reports (2018)