Abstract
Aim:
To investigate the strand preference of the initial cleavage of human pre-miRNAs by human Dicer in vitro.
Methods:
We used a series of in vitro transcribed pre-miRNAs that were radioactively labeled at their 5′ or 3′ ends in cleavage reactions with recombinant human Dicer or HeLa cytoplasmic S100 extracts. Pre-miRNAs samples were purified by denaturing and native PAGE and only the stem-loop structures were used in the experiments. Products of cleavage reactions were resolved by denaturing PAGE, and scanned by phosphor-imaging.
Results:
Recombinant hDicer performs a biased first-cleavage in the pre-let-7b and hsa-pre-miR-17 3′ strand. This result is recapitulated in HeLa S100 cytoplasmic extracts.
Conclusion:
The differential first-nick is observed in cleavage reactions only when stem-loops are substrates for hDicer.
Similar content being viewed by others
Introduction
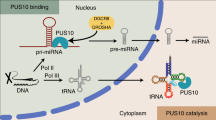
In mammals, microRNA (miRNAs) biogenesis begins when primary microRNAs (pri-miRNAs) are processed by the microprocessor complex (Drosha/DGCR8) to form ∼60−80 nt hairpin-like precursors (pre-miRNAs) inside the nucleus. These precursors are subsequently exported to the cytoplasm via exportin 51, 2, 3, where they are further processed by Dicer, a type III RNase that cleaves the stem-loop to yield shorter, double-stranded miRNAs (duplex miRNAs) with characteristic 2 nt 3′ overhangs, 3′ OH groups and 5′ phosphates4, 5.
Human Dicer (hDicer) is a ∼220 kDa protein that contains an N-terminal DExH-box RNA helicase-like domain, a domain of unknown function (DUF283), a PIWI-Argonaute-Zwille (PAZ) domain, two RNAse-III domains (RIIIa and RIIIb), and a double stranded RNA binding domain (dsRBD)6. Biochemical and crystallographic analyses of hDicer indicate that Mg2+, but not ATP, is required for RNA cleavage7, 8, 9. Analysis of dsRNA processing by hDicer enzymes that are engineered with specific RIII domain mutations revealed that hDicer contains a single dsRNA processing center, which is formed via intramolecular dimerization of the two RIII domains; these domains function independently to cleave phosphodiester bonds on opposite strands of the dsRNA substrate8, 10. hDicer has been shown to associate with the human immunodeficiency virus transactivating response RNA-binding protein (TRBP) to process pre-miRNAs.
Despite our current knowledge of hDicer biochemistry and its activity on perfectly matched double-stranded RNAs, little is known about the first step in the pre-miRNA processing. Some studies have used radioactive pre-miRNAs to show that when hDicer finds and cleaves a pre-miRNA, cleavage does not occur simultaneously in both strands11, 12, 13 due to the fact that Dicer contains only one catalytic center. In those studies, an intermediate band (∼50 nt) between the intact pre-miRNA and the mature miRNA was visualized, ... suggesting that hDicer cleaves (or makes a first-nick in) a specific strand. However, it is not clear whether hDicer cleaves the 5′ or 3′ strand preferentially, or whether its preference is dependent on the substrate. Using a series of precursor stem-loops and differential labeling, we show that recombinant hDicer makes a first-nick in a specific strand and that this activity is recapitulated in S100 HeLa cytoplasmic extracts. We also show that 3′ overhangs influence the strand selection for the first-nick. Our in vitro results suggest that hDicer selects the pre-miRNA strand by first positioning the 3′ overhang.
Materials and methods
DNA templates for in vitro transcription
DNA oligonucleotides were chemically synthesized by SIGMA/Genosys (St Louis, MO) or IDT (Coralville, IA) and purified by PAGE. Each oligonucleotide contains a DNA sequence complementary to the microRNA precursor sequences and to the sequence of T7 RNA polymerase promoter at the 3′ end (Table 1). The double-stranded templates for in vitro transcription were prepared with the Klenow fragment of DNA polymerase I. The DNA template reaction for each precursor contained 6 μL of 50 μmol/L T7 primer, 6 μL of 50 μmol/L precursor oligonucleotide, 20 μL 10×Klenow reaction buffer, 5 μL of 10 μmol/L dNTPs and 158 μL of water. The mixture was heated for 2 min at 94 °C and slowly cooled to 37 °C. Once cooled, 2 units of Klenow enzyme were added and incubated for 1 h at 37 °C. Finally, the reactions were phenol/chloroform extracted and diluted in 50 μL of DEPC-treated water.
In vitro transcription and RNA purification
The pre-microRNAs used in this study were prepared by in vitro transcription with T7 RNA polymerase (New England Biolabs, Ipswich, MA). Transcription reactions were carried out in a 120 μL volume containing 150 pmol of DNA template, 4 μL of each rNTP at 100 mmol/L, 20 μL of MgCl2 at 50 mmol/L, 12 μL of 10×T7 reaction buffer and 500 units of T7 RNA polymerase. The transcription reactions were incubated at 37 °C overnight. [α-32P] UTP was added to internally labeled RNAs (100 μCi each). Then, 10 units of RNAse-free DNAse (Roche, Indianapolis, IN) and alkaline phosphatase (Roche, Indianapolis, IN) were added and the reaction was incubated for 1 h at 37 °C to remove the 5′ triphosphates added by the T7 transcription. Transcripts were run on denaturing 8 mol/L urea, 10% PAGE (acrylamide/bisacrylamide, 19/1) gels in 0.5×TBE, gel-excised, eluted from the gel in 0.3 mol/L sodium acetate, pH 5.2, 0.5 mmol/L EDTA, and 0.1% SDS, and precipitated with ethanol. All transcripts were phosphorylated at the 5′-end by T4 polynucleotide kinase with cold or [γ-32P] ATP (4500 Ci/mmol; Amersham) and phenol/chloroform extracted. For 3′ end labeled RNAs, RNA ligase (Ambion, Austin, TX) and [α-32P] pCp were used according to the manufacturer's recommendations. As pCp labeling leaves a 3′ phosphate, we removed the extra phosphate using either T4 kinase (New England Biolabs, Ipswich, MA) or alkaline phosphatase. The samples were then phenol/chloroform extracted, precipitated, and subjected to another round of 5′ labeling with cold ATP.
Stem-loop structure purification by native PAGE
We analyzed the RNA structure homogeneity of all investigated transcripts by native 10% PAGE (acrylamide/bisacrylamide, 19/1) in 0.5×TBE. Only the bands containing hairpin-like structures were excised, eluted overnight and ethanol precipitated. To prevent the disruption of the stem-loop structures, the samples were maintained at 4 °C until the experiments with the Dicer enzyme were carried out. Secondary structures were predicted using the Mfold algorithm (release 3.2)14.
Hsa-pre-miR-17 5′ and 3′ end mapping
The ends of hsa-pre-miR-17 were identified according to the procedure used previously for human let-7 precursors15. Briefly, total RNA was obtained from HeLa cells using Trizol (Ambion, Austin, TX), and the ≤100 nt fraction was collected using FlashPage electrophoresis (Ambion, Austin, TX). The RNA was self-ligated with T4 RNA ligase. Following ligation, the RNA was phenol/chloroform extracted and precipitated with ethanol. The hsa-pre-miR-17 sequence was amplified by RT-PCR using specific hsa-pre-miR-17 primers (Table 1). PCR products were resolved in a 4% LMP agarose gel (Invitrogen, Carlsbad, CA) and the band was excised, column-purified (Invitrogen, Carlsbad CA) and cloned with the pDrive cloning kit (Qiagen). Plasmid DNA from seven individual clones derived from three independent RNA extractions were sequenced and found to be identical (data not shown).
In vitro reactions using hDicer and cytoplasmic HeLa extracts
For in vitro reactions using recombinant hDicer, we incubated 5×104−10×104 cpm. of labeled precursor with 1 U of rDicer (Genlantis) according to the manufacturer's instructions. For in vitro reactions using HeLa cell S100 cytoplasmic extracts (kindly provided by Phillip D ZAMORE, University of Massachusetts), we incubated 5×104−1×105 cpm. of labeled precursor with 25 μg of cell extract, 3.2 mmol/L MgCl2 and 0.5 mmol/L ATP at 37 °C for the indicated times. Reactions were stopped by the addition of loading buffer containing 7.5 mol/L urea, heated at 95 °C for 5 min, and immediately run in 15%–20% denaturing PAGE. After electrophoresis, the gel containing the separated RNAs was exposed to a phosphorscreen and analyzed by phosphorimaging. The processing efficiency for each hairpin RNA (Figure 1) was determined as previously reported16: processing efficiency=matureIDV/(matureIDV+precursorIDV). IDV: Integrated Density Value.
MicroRNA precursors used in this study. Secondary structures were predicted using Mfold (version 3.2)1. (A) Human miRNA precursors — pre-let-7a-2, pre-let-7b, and pre-mir-30a — 5′ and 3′ ends were used based on previous studies whereas human pre-mir-17 5′ and 3′ ends were identified in this work. (B) Precursors used in (A) having a 5′ overhang. To produce 5′ overhangs, indicated with grey boxes, we used the naturally nucleotides appearing in each primary precursor as templates (miRBase 12.0)2, 3, 4 to overhang the 5′ ends, and cropped the 3′ ends leaving 2-nucleotide, 5′ overhangs. Nucleotides in red indicate mature miRNA sequences.
Results
Precursor miRNAs used in this study
miRBase is the public database where the sequences of all newly discovered mature microRNAs are deposited for classification17, 18, 19. However, precursor miRNAs (either pre-miRNAs or pri-miRNAs) are not annotated for actual sequences; the predicted stem-loop sequences in the database are not strictly pre-miRNAs, but include the pre-miRNA and some flanking sequence from the presumed primary transcript. Currently, ends have been mapped for only a few pre-miRNAs. For this reason, we mapped the 5′ and 3′ ends of hsa-pre-miR-17 from HeLa cells using a method reported previously (see Methods and Figure 1A). This pre-miRNA was initially reported to contain only one mature miRNA (miR-17, 23 nt), but another miRNA was later identified in the complementary strand (miR-17*, 22 nt)17, 18, 19. Notably, we found that pre-miR-17 contains an extra cytosine that is not found in the mature mir-17* sequence. Recent studies have reported variability in pre-miRNA ends due to Drosha processing6, 20, 21, which may explain why we observed this extra nucleotide. We used this sequence as the actual pre-miR-17 throughout this study. In addition to pre-miR-17, we included three other pre-miRNAs, the ends of which have been mapped previously: hsa-pre-miR-30 (pre-miR-30a), hsa-pre-let-7a-2 (pre-let-7b), and hsa-pre-let-7b (pre-let-7b)15 (Figure 1A).
We aimed to determine whether differential cleavage occurs in the first step of cleavage by hDicer on the miRNA or miRNA* strands when hDicer finds a stem-loop. For this reason we double-purified all RNAs by denaturation and native PAGE (Figure 2). Following transcription, we purified the RNAs by denaturing PAGE and tested the samples for degradation and purity (Figure 2A). After this step, RNA molecules adopted one of two major structures, an RNA stem-loop or an RNA dimer (Figure 2B and 2C), which we then resolved by native PAGE (Figure 2D, left). Denaturing gel extraction did not entirely prevent RNA dimer formation, even when we heated the samples at 95 °C and slowly cooled them to RT. On the contrary, heating the samples promoted dimer formation. Therefore, we subsequently gel extracted only the stem-loops and verified their integrity by a subsequent native PAGE prior to each experiment (Figure 2D, right).
RNA preparation and purification. MicroRNA precursors were synthesized using T7 polymerase. (A) Following transcription, each RNA was resolved by UREA-PAGE (eg, pre-let-7a-2), gel-extracted and tested again for integrity by denaturing PAGE (left and right). (B) While the RNA molecules aligned, two major folding types appeared: a precursor RNA dimer (B), and a precursor RNA hairpin (C). (D) Following denaturing gel extraction, we resolved all RNAs by Native-PAGE and only stem-loop precursor structures were gel-extracted (left). Each hairpin structure was tested again by Native-PAGE to confirm that did not form precursor RNA dimers (right; pre-let-7a-2 and pre-let-7b are shown as examples). To test whether dimer structures could be avoided, samples were heated to 95 °C and slowly cooled down to RT (B, boiled; N, not boiled). Heating the samples did not disrupt RNA dimer structures but, in contrast, promoted their formation.
First-nick step in pre-miRNA processing by recombinant hDicer
We incubated internally labeled pre-miRNAs with recombinant hDicer and allowed the cleavage reactions to proceed until the indicated time points (Figure 3A–3D). All precursor RNAs were processed by hDicer to produce two bands near 20 nt (gray arrows), which correspond to mature miRNAs and might also correspond to a piece of or the entire loop sequence. The abundance of mature miRNAs increased in a time-dependent manner as a result of hDicer activity. In addition to the mature miRNAs, bands near ∼40−50 nt were observed (Figure 3A–3D, black arrows). The length of the fragments obtained indicates that these bands correspond to a first-nick step in the processing that occurs near the loop of the pre-miRNAs. Interestingly, the first-nick intermediates decreased in abundance over the time course, in contrast to the mature miRNAs, which continued to accumulate for the entire incubation time (Figure 3A–3D, right graphs).
Precursor microRNA processing by, recombinant hDicer in vitro. Internally labeled ([α-32P] UTP) pre-miRNAs were incubated with recombinant hDicer. Reactions were stopped at different time points (5, 15, 30, 60, and 480 min) and resolved by 20% (v/v) denaturing PAGE (A−D). Grey arrows indicate the bands expected from cleavage reactions (mature miRNAs); black arrows indicate the cleavage first nick product. Note that the first nick product (∼40−50 nt) reaches its maximum at ∼60 min and decreases at ∼480 min whereas the mature products increment. C, 0-time point; M, decade marker. Asterisks indicate precursor loops.
To test whether the first nick was made in the miRNA or miRNA* strand (namely 5′ or 3′ strand), we labeled either the 5′ or the 3′ end of each pre-miRNA and incubated the samples with hDicer (Figure 4A–4D). Time course experiments indicated that recombinant hDicer is able to discriminate the 5′ strand from the 3′ strand. First-nick intermediates for pre-let-7a-2, pre-let-7b and pre-miR-17 appeared to be labeled at the 5′ end (Figure 4A, 4B and 4D, asterisks), indicating that hDicer made the first nick in the 3′ strand of the stem-loop. In the case of pre-miR-30a, hDicer cleaved both strands with similar efficiency (Figure 4C, asterisks), indicating that recombinant hDicer can make the first nick on either strand. Next, we determined whether these results could be recapitulated in HeLa S100 cytoplasmic extracts. Pre-let-7b and pre-miR-17 were preferentially cleaved first in their 3′ strands (Figure 5B and 5D, asterisks); the 5′ strand of pre-let-7b was barely cleaved, and the first-nick in the 5′ strand of pre-miR-17 was negligible. Experiments with pre-let-7a-2 revealed a tendency to make the first nick on the 5′ strand. By contrast, pre-miR-30a was preferentially nicked on the 3′ strand (Figure 5A and 5C, asterisks). To determine the identity of the pre-let-7b and pre-miR-17 recombinant hDicer reaction products, we cloned and sequenced all reaction products from 19–60 nt (data not shown). This analysis revealed that first-nick intermediates possessed the 5′ strand and the loop in a 5:1 ratio. These results indicate that in vitro hDicer discriminates between strands when making the first nick, and that its choice of strand depends on the specific pre-miRNA. The first-nick bias does not seem to be related to the position of the mature miRNA or miRNA* (Figure 1A).
Recombinant hDicer makes differential cleavages on pre-miRNAs. Pre-miRNAs were labeled radioactively either on their 5′ or 3′ ends and incubated with hDicer. Reactions were stopped at 5, 15, 30, 60, and 480 min and resolved by 20% (v/v) denaturing PAGE. Asterisks indicate the position of the first-nick product (∼40−50 nt). hDicer cleaves pre-let-7a-2, pre-let-7b and pre-mir-17 stem-loops preferentially on one strand (3′ end strand), whereas pre-miR-30a is cleaved similarly on both strands. Arrows on the stem-loop drawings represent the preferential first-nick for each precursor. C, 0-time point; M, decade marker.
hDicer makes differential cleavages on pre-miRNAs in HeLa cytoplasmic extracts. Pre-miRNAs were labeled radioactively either on their 5′ or 3′ ends and incubated with HeLa cytoplasmic extracts. Reactions were stopped at 5, 15, 30, 60, and 480 min and resolved by 15% (v/v) denaturing PAGE. Asterisks indicate the position of the first-nick product (∼40–50 nt). hDicer cleaves pre-let-7b and pre-mir-17 stem-loops preferentially on one strand (3′ end strand), whereas the first nick on pre-let-7a-2 — in contrast to our experiments using recombinant hDicer — is preferentially on the 5′ strand. There was a tendency towards the 3 strand in pre-miR-30a. Arrow-sizes on the stem-loop drawings indicate the preferential first-nick for each precursor. C, 0-time point; M, decade marker.
Since most pre-miRNAs are Drosha products generated in the nucleus22, 23, 24, 25, pre-miRNAs bear the characteristic signature of RNase III products: 2 nt, 3′ end overhangs1, 12, 20, 26. As it is known that the PAZ domain binds double-stranded RNA by anchoring 3′ end overhangs1, 3, 5, 26, 27, we investigated whether the overhang position influenced the strand preference for the first-nick introduced by hDicer. We modified the ends of all pre-miRNAs tested to leave 2 nt 5′ overhangs (Figure 1B), incubated these modified pre-miRNAs with recombinant hDicer and resolved the reactions by denaturing PAGE (Figure 6). Our results indicate that 2 nt 5′ overhangs modified the first-nick strand preference, allowing the cleavage of both strands for pre-let-7b and pre-miR-17 (Figure 6B and 6D, asterisks) and biasing initial cleavage to the 5′ strand of pre-let-7a-2 and pre-miR-30a. Interestingly, pre-let-7b and pre-miR-30a intermediates changed in size when labeled at their 5′ or 3′ ends rather than possessing 3′ overhangs (Figures 4 and 5), indicating that the scissile phosphate was changed. These results suggest that hDicer anchors the PAZ domain as a first step in pre-miRNA cleavage, and therefore acts as a molecular ruler to cleave the precursor to a specific size by introducing the first-nick.
The first-nick made by recombinant hDicer is influenced by the overhang position in the pre-miRNAs. Pre-miRNAs having a 5′ end overhang were radioactively labeled either on their 5′ or 3′ ends, incubated with hDicer at 30 or 60 min time points and reactions were resolved by 20% (v/v) denaturing PAGE. Asterisks indicate the position of the first-nick product (∼40−50 nt). C, 0-time point; M, decade marker.
Discussion
Here, we show that hDicer differentiates between miRNA precursor strands and introduces a first-nick either in the 5′ or 3′ strand. The choice of first-nick strand can be modified by the presence of a 2 nt 5′ overhang in the precursor, supporting the model in which hDicer binds 3′ ends through its PAZ domain6. Our results suggest that, in order to act as a molecular ruler, hDicer initiates pre-miRNA processing after anchoring the 3′ end overhang. If improper binding of the 3′ overhang to the PAZ domain occurs, the processing of the pre-miRNA will be altered by a shift at the scissile phosphate (Figure 6B and 6D). This might be the result of the “wrong” positioning of the stem-loop in hDicer owing to the 5′ overhang. The stem loop adopts a helical shape10, so the incorrect anchoring of the 5′ end could cause a shift in the cleavage site; a shift of two helical turns of the RNA would cause the observed ∼22 nt product change. This in turn causes the intermediate to become longer or shorter. Lima et al reported recently that 3′ overhangs are important for the affinity of hDicer and siRNAs, as well as for biased strand loading into the RNA-induced silencing complex28. The erratic cleavage position of precursors with 5′ overhangs is interesting as none of these species has been reported endogenously. In light of our results, this suggests that the location of the first-nick might influence miRNA loading on Argonaute 2. Further studies will be required to verify this hypothesis.
Intermediates in pre-miRNAs processing have been visualized in denaturing gels previously11, 13, 16, 29; however, differential first-nicks have not been reported. We attribute our ability to see the differential first-nick to the gel-extraction and subsequent analysis of only the stem-loop precursor. Failure to perform this isolation step results in a mix of two major RNA structures in the samples ― a stem loop and a dimer. Although the proportion of the RNA dimer is small compared with the stem loop (Figure 2C), the dimer is a substrate of hDicer. Consequently, no difference in the first nick could be visualized with a denaturing PAGE gel, as hDicer is able to enter from either end and cleave the RNA dimer (Figure 7A and 7B). Therefore, a mix of dimer and stem-loop RNAs in the cleavage reaction masks the existence of the differential first-nick. This finding is particularly important because the standard method to form stem-loops is by heating the pre-miRNA samples; however, boiling the samples promotes RNA dimer formation.
First-nick in RNA dimers by hDicer. (A) When the RNA precursors are labeled at the 5′ end, hDicer enters and introduces a nick (from either end) in the corresponding strand. The second-nick yields two labeled products — a ∼21 nt and the ∼40−50 nt intermediate. (B) If the RNA precursors are labeled at the 3′ end, there is no difference in size compared to 5′ labeled precursor dimer when the products are resolved by denaturing PAGE; hDicer activity will appear as if there is no discrimination for the precursor strands.
Another interesting finding was that recombinant hDicer and HeLa cell extracts did not process pre-miRNAs identically. While a recombinant hDicer generated a differential first nick in pre-let-7b and pre-miR-17, we observed only a small bias towards one strand in pre-let-7a-2 and pre-miR-30a in HeLa cell extracts. This suggests that other pathway components, as well as their compartmentalization, might influence strand selection in endogenous pre-miRNA stem loops.
It will be important to investigate whether such differential first nick occurs in all human pre-miRNAs in living cells, as well as to identify other factors that might be involved in the processing.
Author contribution
Carlos Fabián FLORES-JASSO designed and performed research, and wrote the paper. Catalina ARENAS-HUERTERO and Jose Luis REYES performed research and contributed ideas. Cecilia CONTRERAS-CUBAS performed research. Alejandra COVARRUBIAS contributed ideas and reagents. Luis VACA contributed ideas and wrote the paper.
References
Han J, Lee Y, Yeom KH, Kim YK, Jin H, Kim VN . The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev 2004; 18: 3016–27.
Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U . Nuclear export of microRNA precursors. Science 2004; 303: 95–8.
Yi R, Qin Y, Macara IG, Cullen BR . Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev 2003; 17: 3011–6.
Elbashir SM, Lendeckel W, Tuschl T . RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev 2001; 15: 188–200.
Tomari Y, Zamore PD . MicroRNA biogenesis: drosha can't cut it without a partner. Curr Biol 2005; 15: R61–64.
MacRae IJ, Doudna JA . Ribonuclease revisited: structural insights into ribonuclease III family enzymes. Curr Opin Struct Biol 2007; 17: 138–45.
Provost P, Dishart D, Doucet J, Frendewey D, Samuelsson B, Radmark O . Ribonuclease activity and RNA binding of recombinant human Dicer. EMBO J 2002; 21: 5864–74.
Zhang H, Kolb FA, Jaskiewicz L, Westhof E, Filipowicz W . Single processing center models for human Dicer and bacterial RNase III. Cell 2004; 118: 57–68.
Zhang H, Kolb FA, Brondani V, Billy E, Filipowicz W . Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J 2002; 21: 5875–85.
Takeshita D, Zenno S, Lee WC, Nagata K, Saigo K, Tanokura M . Homodimeric structure and double-stranded RNA cleavage activity of the C-terminal RNase III domain of human dicer. J Mol Biol 2007; 374: 106–20.
Leuschner PJ, Martinez J . In vitro analysis of microRNA processing using recombinant Dicer and cytoplasmic extracts of HeLa cells. Methods 2007; 43: 105–9.
Lund E, Dahlberg JE . Substrate selectivity of exportin 5 and Dicer in the biogenesis of microRNAs. Cold Spring Harb Symp Quant Biol 2006; 71: 59–66.
Obernosterer G, Leuschner PJ, Alenius M, Martinez J . Post-transcriptional regulation of microRNA expression. RNA 2006; 12: 1161–7.
Zuker M . Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 2003; 31: 3406–15.
Basyuk E, Suavet F, Doglio A, Bordonne R, Bertrand E . Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucleic Acids Res 2003; 31: 6593–7.
Soifer HS, Sano M, Sakurai K, Chomchan P, Saetrom P, Sherman MA, et al. A role for the Dicer helicase domain in the processing of thermodynamically unstable hairpin RNAs. Nucleic Acids Res 2008; 36: 6511–22.
Griffiths-Jones S . The microRNA registry. Nucleic Acids Res 2004; 32: D109–11.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 2006; 34: D140–4.
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ . miRBase: tools for microRNA genomics. Nucleic Acids Res 2008; 36: D154–8.
Helvik SA, Snove O Jr, Saetrom P . Reliable prediction of Drosha processing sites improves microRNA gene prediction. Bioinformatics 2007; 23: 142–9.
Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 2006; 125: 887–901.
Berezikov E, Chung WJ, Willis J, Cuppen E, Lai EC . Mammalian mirtron genes. Mol Cell 2007; 28: 328–36.
Babiarz JE, Ruby JG, Wang Y, Bartel DP, Blelloch R . Mouse ES cells express endogenous shRNAs, siRNAs, and other microprocessor-independent, Dicer-dependent small RNAs. Genes Dev 2008; 22: 2773–85.
Okamura K, Chung WJ, Lai EC . The long and short of inverted repeat genes in animals: microRNAs, mirtrons and hairpin RNAs. Cell Cycle 2008; 7: 2840–5.
Martin R, Smibert P, Yalcin A, Tyler DM, Schafer U, Tuschl T, et al. A Drosophila pasha mutant distinguishes the canonical microRNA and mirtron pathways. Mol Cell Biol 2009; 29: 861–70.
Carmell MA, Hannon GJ . RNase III enzymes and the initiation of gene silencing. Nat Struc Mol Biol 2004; 11: 214–8.
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003; 425: 415–9.
Lima WF, Murray H, Nichols JG, Wu H, Sun H, Prakash TP, et al. Human Dicer binds short single-strand and double-strand RNA with high affinity and interacts with different regions of the nucleic acids. J Biol Chem 2009; 284: 2535–48.
Ye X, Paroo Z, Liu Q . Functional anatomy of the Drosophila microRNA-generating enzyme. J Biol Chem 2007; 282: 28373–8.
Acknowledgements
We thank P ZAMORE for helpful discussions and for kindly sharing the facilities in his laboratory. We would also like to thank the members of our lab and the Zamore lab for helpful discussions. This work was supported by CONACyT (#42469) and DEGEP.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Flores-jasso, C., Arenas-huertero, C., Reyes, J. et al. First step in pre-miRNAs processing by human Dicer. Acta Pharmacol Sin 30, 1177–1185 (2009). https://doi.org/10.1038/aps.2009.108
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2009.108
Keywords
This article is cited by
-
Secondary structure RNA elements control the cleavage activity of DICER
Nature Communications (2022)
-
A novel family of mammalian transmembrane proteins involved in cholesterol transport
Scientific Reports (2017)
-
Characterization of a naturally occurring truncated Dicer
Molecular Biology Reports (2015)
-
Two-step cleavage of hairpin RNA with 5' overhangs by human DICER
BMC Molecular Biology (2011)
-
The role of the precursor structure in the biogenesis of microRNA
Cellular and Molecular Life Sciences (2011)