Abstract
Cellular adhesion and growth on substrates with differing surface topography are known to induce unusual cell behaviors. Here, we investigated the adhesion and growth of human embryonic kidney 293T cells on vertically aligned silicon nanowires. Varying the diameter of nanowires affected their elasticity, which in turn caused variations in cell morphology and adhesion. Calculation of their elastic modulus revealed that long and thin nanowires are easier to deflect and thus provide more focal adhesion sites for adherent cells. And, induced mechanical tension triggered cellular anisotropic growth in the direction of tension, creating a greater rate of filopodium growth than was observed with a bare glass substrate devoid of mechanical tension. This nanotopographical approach provides important insights into the role of artificial substrates in the modulation of cell behavior and a conceptual framework for further analysis of substrate-induced changes in cellular activity.
Similar content being viewed by others
Introduction
In recent years, a significant amount of research has focused on the biophysical cues provided by extracellular matrices that regulate cell phenotype and function.1, 2, 3 In particular, surface topography has been shown to modulate cell adhesion, growth, proliferation and differentiation by altering intracellular signal transduction and gene expression.4, 5 Nanostructured materials with various morphologies have also received much attention over the past few decades in the field of biological research, owing to advances in nanotechnology that have enabled precision controlled nanofabrication. As direct interaction between such in vitro artificial nanostructures and living cells is inevitable, it is necessary to understand the complicated molecular mechanisms occurring at their interface. By engineering the surface topography of substrates to provide more favorable environments that enhance cell adhesion, growth and proliferation, it may be possible to control essential biological responses.6, 7
Although modulation of cell adhesion and growth using various topographical cues is well studied, most published data address or exploit the topographical confinement that is imposed by the nanostructured architecture.8, 9 In other words, overall control of cellular response is performed by cell isolation in the structure through geometry, and is not influenced by surface topography. It is therefore, essential for surface topography to be studied through controlling the shape, size and density of the nanostructure. Given these demands, there have been several studies into nanotopography, such as anodized aluminum oxide, which have different pitch and provided diverse degrees of roughness,10 and vertically aligned silicon nanopillars controlled interpillar spacing, which influenced on cell morphology.11 Interestingly, the accelerated development of hippocampal neurons was observed on well-packed structures of silica beads with diameters in excess of 200 nm, which is comparable with the thickness of filopodia.12, 13
In addition to surface topography, much research has also been focused on surface elasticity. The relationship between contractile forces and the mechanical elasticity, or stiffness, of extracellular matrices can have a major influence on cell behaviors, such as differentiation of stem cells,2, 14 migration,15 apoptosis16 and proliferation.17 A previous study of a tunable hydrogel system, which is decorated with gold nanostructures and can be ‘programmed’ to assume varying degrees of stiffness, found that the adhesion and migration of plasmodium sporozoites is more efficient on stiff, bonelike interfaces than on soft, skin-like ones.18 In addition, stem cells in early stage are impacted by micropost rigidity, resulting in cell morphology, focal adhesion and stem cell differenciation.19
In this study, surface topographies of vertically aligned silicon nanowires (SiNWs) with different diameters were synthesized on Si(111) wafers by chemical vapor deposition, as shown in Figure 1a. Human embryonic kidney (HEK) 293T cells were subsequently cultured on these substrates and their responses in terms of adhesion and growth behavior were investigated for 2 days. Particular attention was given to the promotion of focal adhesion and anisotropic cell growth induced by mechanical tension of elastic nanotopography.
(a) Schematic of the furnace used for the growth of vertically aligned silicon nanowires (SiNWs). (b) Illustration of the growth process for vertically aligned SiNWs on Au-covered Si(111) wafer used as catalyst. Au film and SiNWs are represented by yellow and red, respectively. Scanning electron microscopy images (20o tilt-view) of vertically aligned SiNWs after vapor–liquid–solid growth on an Si(111) wafer covered with a (c) 1-nm- or (d) 10-nm-thick Au film at 860 °C for 10 min.
Materials and methods
Chemicals
Silicon tetrachloride (SiCl4, 99.998%) and ammonium fluoride (NH4F, ~40% in water) were purchased from Aldrich (St Louis, MO, USA). Hydrofluoric acid (48–52% solution in water) was purchase from Acros (Pittsburgh, PA, USA). Acetone and isopropyl alcohol were purchased from Samchun (Seoul, South Korea). Si(111) wafers were purchased from LG siltron (Seoul, South Korea).
Preparation of Au-covered Si substrate
The Si(111) wafer was cleaned with acetone and isopropyl alcohol, and dried by using nitrogen gas. It was subsequently dipped for 4 min to buffered hydrofluoric acid (aqueous; 9% hydrofluoric acid and 32% NH4F solution in water) to remove native oxide layer, followed by rinsing with water. And, Au was deposited on the cleaned Si(111) wafer by e-beam evaporation. The deposited Au thickness was 1 and 10 nm, respectively. And then, the substrate was diced into small pieces with 1 cm (width) × 1 cm (length). The Au-covered Si substrate was used for SiNWs growth. The patterned SiNWs can be synthesized using Au patterns with hole size of 400 nm and height of 30 nm on Si(111) wafer, which were fabricated by KrF stepper lithography.
Synthesis of Si nanowires
Vertically aligned SiNWs were synthesized by chemical vapor deposition in a 12-inch horizontal tube furnace (Lindberg/Blue M, Waltham, MA, USA) equipped with a 1-inch diameter quartz tube as shown in Figure 1a. In general, the Au nanoparticles (AuNPs) determine the diameter and position of the SiNWs.20 The prepared Au-covered substrate was placed at the center of the heating zone in the tube furnace. Before the experiment, the quartz tube was evacuated and flushed repeatedly with Ar gas (high purity, 99.999%) to minimize oxygen contamination. The furnace was then heated to the reaction temperature of 860 °C with 20 °C min−1 during the ramping period (see Supplementary Figure 1 for details). Here, there are two roles of 10% H2 gas in Ar gas used in the reaction. The one is carrier gas (C), which passes bubbler contained SiCl4 solution as used for Si precursor and carries Si precursor into the reactor. The other is dilution gas (D), which directly gets in the reactor and adjusts the concentration of Si precursor. Before the reaction of SiNWs growth, dilution gas is only flowed with 100 s.c.c.m. during the ramping period, then carrier gas and dilution gas is flowed with 100 and 750 s.c.c.m., respectively, during the growth period. After the growth reaction for 10 min, the reactor is quickly cooled until reaching room temperature.
Cell culture
HEK-293T cells were cultured in Dulbecco’s modified Eagle’s medium containing L-glutamine and supplemented with antibiotics (100 U ml−1 penicillin and 100 μg ml−1 streptomycin) and 10% fetal bovine serum. Cells were maintained in a humidified 5% CO2 incubator at 37 °C.
Scanning electron microscopy measurement
Following culture on bare glass and vertically aligned SiNWs for 48 h, HEK-293T cells were fixed in 4% formaldehyde in phosphate-buffered saline, and then washed with phosphate-buffered saline buffer. Subsequently, the cells were dehydrated in a series of concentrations of ethanol and hexamethyldisilazane. Before the sample was measured using a field effect scanning electron microscopy, it was coated with platinum using a sputter coater.
Immunofluorescence analysis
HEK-293T cells were seeded at density of 2 × 104 cells onto bare glass or SiNWs substrate, and incubated in a humidified 5% CO2 incubator at 37 °C for 24 h. Cells were fixed in 4% paraformaldehyde for 15 min at room temperature, and then permeabilized in 0.2% Triton X-100 for 15 min at room temperature. The samples were incubated with primary mouse anti-vinculin antibody at room temperature for 3 h, followed by incubation with secondary antibodies conjugated with tetramethylrhodamine (TRITC) for 1 h. Cells were then incubated with F-actin-specific phalloidin antibody for 1 h at room temperature. Finally, immunofluorescence was visualized using an upright bright-field microscope (Olympus, BX51WI, Center Valley, PA, USA).
Immunoblot assay
HEK-293T cells were lysed with lysis buffer from Cell Signaling (Danvers, MA, USA) on ice for 30 min. Samples were then mixed with 5 × SDS sample buffer, and vortexed for 15 s. The cell lysate samples were heated at 99 °C for 10 min and separated electrophoretically on a 10% SDS-polyacrylamide gel. Subsequently, proteins were transferred onto a 0.45-μm nitrocellulose membrane (GE Healthcare, Buckinghamshire, UK) for 2 h. The membrane was then incubated with anti-mouse IgG antibody conjugated to horseradish peroxidase (Assay Designs, Ann Arbor, MI, USA) at room temperature for 2 h. The proteins were visualized using an enhanced chemiluminescent substrate (Thermo Fisher Scientific, Logan, UT, USA) and analyzed using a LAS3000 luminescent image analyzer (Fuji Film, Tokyo, Japan). The protein bands were quantified using NIH ImageJ software (NIH, Bethesda, MD, USA).
Results and Discussion
The growth process for vertically aligned SiNWs, using Au thin film as a catalyst, is described in Figure 1b.21, 22 By increasing the temperature of substrates, the Au thin film on the Si(111) wafer starts to randomly crack and then evaporate, migrating from the wafer and forming liquid-phase Au nanodroplets of various sizes. This cracking allows Au to be randomly deposited onto a Si(111) wafer and diffuse over the substrate by the two well-known mechanisms of coalescence and Ostwald ripening. This ultimately leads to the formation of AuNPs.23 After reaching the reaction temperature, vapor-phase Si precursors are introduced into the liquid-phase AuNPs by 10% H2 in an Ar carrier gas, thereby forming a single crystal of Si by precipitation from the Au-Si alloy upon reaching supersaturation.24 Finally, vertically aligned SiNWs are synthesized by vapor–liquid–solid growth from the AuNPs.25
We next regulated the thickness of the Au film to either 1 or 10 nm, as this allows considerable control over the diameter of the SiNWs. Representative scanning electron microscopy images of SiNWs grown from 1 nm- to 10 nm-thick Au films, denoted as SiNWs-01 and SiNWs-10, respectively, are shown in Figures 1c and d. Consistent with the size of AuNPs formed during the ramping process (see Supplementary Figure 2 for details), the average diameter and density of SiNWs-01 is 72±9 nm and 2.5±0.3 μm−2; whereas for SiNWs-10, these values are 218±48 nm and 2.0±0.3 μm−2. The diameter distribution of SiNWs-01 and SiNW-10 is shown in Supplementary Figure 3. Interestingly, although the densities of SiNWs-01 and SiNWs-10 are similar, the diameter of the latter is about three times thicker than observed with SiNWs-01. This confirms that varying the thickness of the Au film allows a considerable degree of control over the diameter of synthesized SiNWs. The morphology of the SiNWs was also characterized by transmission electron microscopy analysis (see Supplementary Figure 4 for details), which shows representative low-magnification (a) and high-magnification (b) transmission electron microscopy images of a SiNW synthesized from a 10-nm-thick Au film. The SiNW surface consists of a native oxide layer, which are the same component with bare glass.
The adhesion of HEK-293T cells on bare glass and on the synthesized SiNWs was investigated over a 48-h period; the results are shown in Figure 2 and in Supplementary Figure 5. HEK-293T cells cultured on bare glass exhibited flat and extensive spreading, a diverse morphology and also did not display any preferential alignment (Figure 2a). Cells cultured on SiNW substrates could also adhere regardless of the diameter of the nanowire used, and their high density prevented them from being pierced by the SiNWs (Figures 2b and c).26, 27 When cells adhered to the SiNWs-10 substrate, they rounded up extensively and exhibited a radial alignment (Figure 2b), whereas those on a SiNWs-01 substrate displayed an anisotropic morphology (Figure 2c). As they were associated with different degrees of spreading, the morphologies of the adhered cells had a significant influence on the cell area (N=72 cells), as shown in Figure 2g. Thus, the area of cultured cells was three times greater on bare glass than on either SiNW substrates. However, the area occupied by cells was more extensive on the SiNWs-01 substrate than the SiNWs-10 substrate. High-magnification images revealed that the filopodia of cells grown on each substrate were different, as shown in Figures 2d–f. The filopodia of cells on bare glass were thick and straight (Figure 2d), whereas those on SiNW substrates were thin and exhibited a unique behavior of wrapping around the nanowires regardless of the nanowire diameter (insets of Figures 2b and c).28 As a result of this wrapping, the SiNWs were deflected by the cell filopodia, with the degree of bending depending on the diameter of the SiNW (Figures 2e and f). Furthermore, the number of filopodia of cells cultured on both SiNW substrates (N=72 cells) was twofold that of cells grown on bare glass, as shown in Figure 2h. In addition, we cultured the cells on SiNW arrays from which the Au tip was removed (see Supplementary Figure 6 for details). This had essentially no effect on morphology, as the cells were similar to those shown in Figures 2a–f. Furthermore, we investigated the effect of different density between SiNWs-10 and SiNW-01 substrates and the morphology of cells grown on high density of SiNWs-10 substrate and low density of SiNWs-01 substrate22 compared with Figures 2b and c, respectively. Although the density of SiNWs-10 substrate increases, the cell morphology is similar in Figure 2b and cell attachment area is still small, whereas in the case of low density of SiNWs-01 substrate, there are cells that spread over the SiNWs-01 substrate (see Supplementary Figure 7).
Scanning electron microscopy images of human embryonic kidney (HEK)-293T cells on bare glass and silicon nanowire (SiNW) substrates after 48 h of incubation (false-colored red). (a) Cell spreading on the bare glass and (d) high-magnification image of the cell edge. (b, e) Cells on the SiNWs-10 substrate. (c, f) Cells on the SiNWs-01 substrate. The insets of b and c show the cells wrapping around SiNWs-10 and SiNWs-01, respectively. (f) The deflection of SiNWs-01 by the interaction of cells with the SiNWs, (e) the minimal deflection observed with SiNWs-10. Analysis of (g) area and (h) number of filopodia of cells cultured on each of the three substrates (mean±standard error, N=72 cells).
A study by Brammer et al. reported that SiNW arrays prevented MC3T3-E1 mouse osteoblast adhesion and spreading;29 consistent with this, we observed that cells grown on SiNWs also failed to spread. Furthermore, the SiNW substrates also decreased the adhesion ability of the cells, which is directly related to focal adhesion. This, in turn, is influenced by the distribution of contact sites between cells and the substrate, with cell attachment by mechano-transductive signaling most likely mediating the relation between topography and cell.30 This distinctive behavior of filopodia on the SiNW substrates can therefore be explained as a result of cells seeking to adhere more stably and strongly by attaching their ends to a nanowire.31 We note that filopodia function as antennae for environmental exploration.12, 13, 32 This may explain why the cells that adhered to uneven SiNWs substrates generated more filopodia, as they likely seek out more contact points.
We performed immunofluorescent staining of vinculin and F-actin, a cytoskeletal protein that associates with integrins and actin. Vinculin localization in cells grown on SiNW substrates differed from localization in cells cultured on bare glass (Figures 3a–c). Specifically, vinculin and F-actin were co-localized throughout the spreading regions of cells grown on bare glass, indicating that the cells were forming continuous focal adhesions. Cells on SiNW substrates could only contact the tips of the vertically aligned nanowires. Therefore, the narrow and separate contacts between cells and the SiNW substrates triggered a weak spherical cell adhesion; this prevented cells from forming stable focal adhesion complexes.9 Vinculin was localized to the contacts with the tips of nanowires, and had a linear distribution on the SiNWs-01 substrate. Furthermore, F-actin was localized to a restricted area at one edge of most cells. Together, these data suggest that SiNW substrates in general, SiNWs-10 substrate in particular, are less effective promoters of cell adhesion. We then used ImageJ analysis software to confirm that the relative intensity of staining (N=56 cells) as lower in cells grown on SiNW substrates compared with those grown on bare glass (Figure 3d). Next, we performed immunoblot assays (N=2 × 104 cells) to determine whether the levels of vinculin and F-actin decreased in cells cultured on SiNW substrates. This was indeed the case, with the effect being more pronounced on SiNWs-10 when compared with that on the SiNWs-01 substrate, as shown in Figures 3e and f. As vinculin is an actin filament cross-linking protein, co-staining of F-actin and vinculin protein shows that the proteins are located in accordance with actin location.9, 33 Therefore, the reduction in F-actin and vinculin staining on the SiNWs-10 when compared with that on the SiNWs-01 substrate is consistent with the lower adhesion capability of the cells; this is because cells require high levels of each protein in order to form stable adhesion complexes.
Fluorescence images of human embryonic kidney (HEK)-293T cells stained with fluorescein isothiocyanate (FITC; green) and phalloidin (red), after 48 h incubation on the following: (a) bare glass, (b) SiNWs-10 substrate and (c) SiNWs-01 substrate. Green=vinculin-FITC, red=F-actin, yellow=co-localization. (d) ImageJ analysis of relative fluorescence intensity of cells cultured on each of the three substrates (N=56 cells). (e) Vinculin and F-actin were detected using an immunoblot assay (N=2 × 104 cells). β-Actin was used as loading control. (f) Quantitative comparison of vinculin and F-actin levels after normalization with β-actin.
Focal adhesion provides the essential structural components for anchoring the actin cytoskeleton to an extracellular matrix.34 As described above, the SiNWs-10 substrates were deficient in focal adhesion sites; this type of insufficiency can induce remodeling of the cell membrane in order to acquire more focal adhesion sites.35 However, the narrow intervals between nanowires would lead to severe deformation of the cell membrane during any remodeling process. This would also require far more energy than is available,36 thereby precluding cells from modifying their membranes on SiNW substrates. The need to increase focal adhesion; therefore, results in alternative interactions between cells and the substrate, which is influenced by the presence of nanowires.
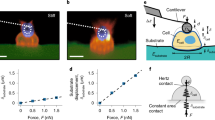
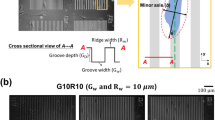
From the known mechanical behavior of NWs, the force required for bending can be derived from the degree of deflection, provided the force–displacement relationship is quantitatively established.37 To investigate the force–displacement relationship of SiNWs with differing diameter, we measured lateral force by atomic force microscopy in contact mode, during which a constant force between the tip and SiNWs is applied, as shown in Figure 4a.38 To deflect a SiNW with length ‘L’ to ‘h’, the pitch of the SiNW array (p) must be sufficiently large. In this regard, we synthesized patterned SiNW arrays (Figure 4c, Materials and Methods section). The direction of the deflected SiNW varied according to the tip movement (Figure 4d). To investigate the effects of the diameter and length of SiNWs on deflection, we measured lateral force for three samples, which were d=200 nm, L=5 μm (Figure 4e), d=200 nm, L=10 μm (Figure 4f) and d=400 nm, L=10 μm (Figure 4g). With increasing SiNW length, the lateral force tended to decrease from 8.4 to 0.2 V, implying that longer SiNWs are easier to deflect (Figures 4e and f). On the other hand, when the diameter of SiNW was larger, the lateral force increased from 0.2 to 5.7 V, indicating that thicker SiNWs are more difficult to deflect (Figures 4f and g). As mentioned above, the different diameters of the SiNWs are directly related to their elasticity. The average elastic modulus of SiNWs is determined by their length and diameter, and the lengths of SiNWs-01 and SiNWs-10 are quite similar at around 5 μm. The diameter of SiNWs is inversely proportional to the average elastic modulus; given that the diameter of SiNWs-10 is 3 times greater than SiNWs-01, the average elastic modulus is therefore 81 times smaller (see Supplementary Information for the calculation of the average elastic modulus of SiNWs). This means that while the thick nanowires of the SiNWs-10 substrate are poorly deflected by interaction with cells, the thin nanowires of the SiNWs-01 substrate are much more easily deflected (Figure 4f). This deflection of the nanowires allows the cells on a SiNWs-01 substrate to secure themselves at contact points and produce more focal adhesion. Thus, changing the nanostructure of the SiNW substrate increased the focal adhesion of cells, although the absolute level of focal adhesion will still be lower than what is achievable on bare glass. In agreement with a previous study,39 the deflection degree of nanowires increased depending on their length. Consistent with this, increasing the length of SiNWs leads to a greater deflection of the SiNWs-10 substrate (Figure 5 and Supplementary Figure 8). The increased SiNW length creates a greater cell adhesion area, thereby leading to a more ‘spread-out’ morphology.
Schematic diagrams of (a) the silicon nanowire (SiNW) bending process following tip movement during the measurement of lateral force and (b) the patterned SiNW array. L, h, d and p are the length, height, the diameter of the SiNWs and the distance between them, respectively. (c) The patterned array of vertically aligned SiNW is shown. (d) Topography images of the SiNWs according to the atomic force microscopy (AFM) scan directions. R=right, L=left. (e–g) Lateral force images of the SiNWs measured by AFM contact mode.
Scanning electron microscopy images of cells cultured on the SiNWs-10 substrate with different lengths. (a, b) The adherent cells on SiNWs-10 substrates with 9.50- and 12.79-μm-long SiNWs, respectively, are shown. (c) The diameter of SiNWs are constant; however, the length of the SiNWs (denoted by L) increases from 5.10 (L1) to 9.50 (L2), and then 12.79 μm (L3), respectively. The length of SiNWs is proportional to the SiNW growth time. Although the diameter of the SiNWs is thick (d=200 nm), degree of deflection of SiNWs and mechanical tension (Tm) increase as the length of SiNW increases.
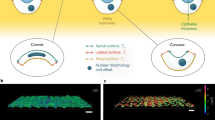
This greater spreading of cells on the SiNWs-01 substrate is further explained by the fact that the former provides more contact points on the side of SiNWs. The morphology and focal adhesion of cells on these three different substrates are depicted in Figure 6. Focal adhesion complexes consisting of F-actin and α-actinin were formed continuously and densely on the flat bare glass surface. In the case of the SiNWs-10 substrate, there was a sparse formation of focal adhesion complexes at the tip of thick SiNWs; however, with the SiNWs-01 substrate, adhesion complexes also formed at the tip as well as on the side of deflected SiNWs. Ligand density influences cell adhesion strength and focal adhesion formation.40, 41, 42 In light of this observation, we infer that the deflection of SiNWs brings adjacent SiNWs in close proximity, thus facilitating the formation of focal adhesion complexes.
Schematic illustrations of focal adhesion formation on (a) bare glass, (b) SiNWs-10 substrate and (c) SiNWs-01substrate. The insets demonstrate different aspects of focal adhesion complexes and the interface between cells and their substrate. (a) Cells spread extensively and flatly, with their focal adhesion complexes formed impregnably. Conversely, the cells of b and c are rounded up and form weak focal adhesion complexes relative to those of a. These weak focal adhesion complexes are improved through deflection of SiNWs, which reduces the distance between SiNWs and thus produces more contact points. The cells adhered to the SiNWs-01 substrate have a higher deflection spread than those on the SiNWs-10 substrate. Mechanical tension is denoted by Tm.
The rate of neurite elongation is a linear function of pulling tension, provided either by the growth cone43 or by experimental manipulation.44 Furthermore, Lamoureux et al. reported that mechanical tension using calibrated glass needles can break symmetry in neurons, and thereby specify which process becomes the axon.45 SiNWs were deflected by cells cultured on SiNW substrates by being first pulled into the center of cell. The deflected SiNWs have restoration force, which pulls the cells into the center of the SiNW. This restoration force creates mechanical tension (Tm), which is induced in the cells by the SiNWs. This tension then triggers anisotropic cell growth in the direction of tension, thereby creating a polarization of cell morphology. In the case of the SiNWs-10 substrate, cell polarity was less significant compared with the SiNWs-01 substrate because of its reduced mechanical tension.
Conclusion
Varying the diameter of the SiNWs affected their elasticity, resulting in a difference in cell morphology and adhesion. In agreement with our calculation of the elastic modulus, the SiNWs-01 substrates proved easier to deflect and therefore provided more cell adhesion sites. The nanotopography of the substrate influenced the distribution of focal adhesion, which is either continuous or discrete, and this subsequently influenced cell adhesion and morphology. Our results also indicate that mechanical tension applied by the SiNWs induced filopodium growth. This suggests that ongoing research should focus on the induction of cell adhesion to SiNWs by controlling nanowire spacing. The effects of SiNWs arrays in guiding cell adhesion and spreading behavior may prove useful for the development of cell microarrays, tissue engineering scaffolds and molecule delivery vehicles, in which strong cell substrate adhesion and reduced cell–cell communication are beneficial.
References
Guilak, F., Cohen, D. M., Estes, B. T., Gimble, J. M., Liedtke, W. & Chen, C. S. Control of stem cell fate by physical interactions with the extracellular matrix. Cell Stem Cell 5, 17–26 (2009).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Yao, X., Peng, R. & Ding, J. Cell–material interactions revealed via material techniques of surface patterning. Adv. Mater. 25, 5257–5286 (2013).
Dalby, M. J., Gadegaard, N., Tare, R., Andar, A., Riehle, M. O., Herzyk, P., Wilkinson, C. D. W. & Oreffo, R. O. C. The control of human mesenchymal cell differentiation using nanoscale symmetry and disorder. Nat. Mater 6, 997–1003 (2007).
Chen, W., Villa-Diaz, L. G., Sun, Y., Weng, S., Kim, J. K., Lam, R. H. W., Han, L., Fan, R., Krebsbach, P. H. & Fu, J. Nanotopography influences adhesion, spreading, and self-renewal of human embryonic stem cells. ACS Nano 6, 4094–4103 (2012).
Tsimbouri, P. M., McMurray, R. J., Burgess, K. V., Alakpa, E. V., Reynolds, P. M., Murawski, L., Kingham, E., Oreffo, R. O. C., Gadegaard, N. & Dalby, M. J. Using nanotopography and metabolomics to identify biochemical effectors of multipotency. ACS Nano 6, 10239–10249 (2012).
Hu, B., Shi, W., Wu, Y.-L., Leow, W. R., Cai, P., Li, S. & Chen, X. Orthogonally engineering matrix topography and rigidity to regulate multicellular morphology. Adv. Mater. 26, 5786–5793 (2014).
Hällström, W., Prinz, C. N., Suyatin, D., Samuelson, L., Montelius, L. & Kanje, M. Rectifying and sorting of regenerating axons by free-standing nanowire patterns: a highway for nerve fibers. Langmuir 25, 4343–4346 (2009).
Schulte, V. A., Díez, M., Möller, M. & Lensen, M. C. Surface topography induces fibroblast adhesion on intrinsically nonadhesive poly(ethylene glycol) substrates. Biomacromolecues 10, 2795–2801 (2009).
Cho, W. K., Kang, K., Kang, G., Jang, M. J., Nam, Y. & Choi, I. S. Pitch-dependent acceleration of neurite outgrowth on nanostructured anodized aluminum oxide substrates. Angew. Chem. Int. Ed. 49, 10114–10118 (2010).
Bucaro, M. A., Vasquez, Y., Hatton, B. D. & Aizenberg, J. Fine-tuning the degree of stem cell polarization and alignment on ordered arrays of high-aspect-ratio nanopillars. ACS Nano 6, 6222–6230 (2012).
Kang, K., Choi, S. −E., Jang, H. S., Cho, W. K., Nam, Y., Choi, I. S. & Lee, J. S. In vitro developmental acceleration of hippocampal neurons on nanostructures of self-assembled silica beads in filopodium-size ranges. Angew. Chem. Int. Ed. 51, 2855–2858 (2012).
Kang, K., Yoon, S. Y., Choi, S.-E., Kim, M.-H., Park, M., Nam, Y., Lee, J. S. & Choi, I. S. Cytoskeletal actin dynamics are involved in pitch-dependent neurite outgrowth on bead monolayers. Angew. Chem. Int. Ed. 53, 6075–6079 (2014).
Ye, K., Wang, X., Cao, L., Li, S., Li, Z., Yu, L. & Ding, J. Matrix stiffness and nanoscale spatial organization of cell-adhesive ligands direct stem cell fate. Nano Lett. 15, 4720–4729 (2015).
Guo, W.-H., Frey, M. T., Burnham, N. A. & Wang, Y.-L. Substrate rigidity regulates the formation and maintenance of tissues. Biophys. J. 90, 2213–2220 (2006).
Wang, J. H.-C. & Thampatty, B. P. Mechanobiology of adult and stem cells. Int. Rev. Cell Mol. Biol. 271, 301–346 (2008).
Hadjipanayi, E., Mudera, V. & Brown, R. A. J. Close dependence of fibroblast proliferation on collagen scaffold matrix stiffness. Tissue Eng. Regen. Med 3, 77–84 (2009).
Perschmann, N., Hellmann, J. K., Frischknecht, F. & Spatz, J. P. Induction of malaria parasite migration by synthetically tunable microenvironments. Nano Lett. 11, 4468–4474 (2011).
Fu, J., Wang, Y.-K., Yang, M. T., Desai, R. A., Yu, X., Liu, Z. & Chen, C. S. Mechanical regulation of cell function with geometrically modulated elastomeric substrates. Nat. Methods 7, 733–736 (2010).
Cui, Y., Lauhon, L. J., Gudiksen, M. S., Wang, J. & Lieber, C. M. Diameter-controlled synthesis of single-crystal silicon nanowires. Appl. Phys. Lett. 78, 2214–2216 (2001).
Ge, S., Jiang, K., Lu, X., Chen, Y., Wang, R. & Fan, S. Orientation-controlled growth of single-crystal silicon-nanowire arrays. Adv. Mater. 17, 56–61 (2005).
Park, Y.-S., Jung, D. H., Kim, H. J. & Lee, J. S. Annealed Au-assisted epitaxial growth of Si nanowires: control of alignment and density. Langmuir 31, 4290–4298 (2015).
Bechelany, M., Maeder, X., Riesterer, J., Hankache, J., Lerose, D., Christiansen, S., Michler, J. & Philippe, L. Synthesis mechanisms of organized gold nanoparticles: influence of annealing temperature and atmosphere. Crystal Growth Design 10, 587–596 (2010).
Hochbaum, A. I., Fan, R., He, R. & Yang, P. Controlled growth of Si nanowire arrays for device integration. Nano Lett. 5, 457–460 (2005).
Wagner, R. S. & Ellis, W. C. Vapor–liquid–solid mechanism of single crystal growth. Appl. Phys. Lett. 4, 89–90 (1964).
Shalek, A. K., Robinson, J. T., Karp, E. S., Lee, J. S., Ahn, D.-R., Yoon, M.-H., Sutton, A., Jorgolli, M., Gertner, R. S., Gujral, T. S., MacBeath, G., Yang, E. G. & Park, H. Vertical silicon nanowires as a universal platform for delivering biomolecules into living cells. Proc. Natl Acad. Sci. USA 107, 1870–1875 (2010).
Kim, W., Ng, J. K, Kunitake, M. E., Conklin, B. R. & Yang, P. Interfacing silicon nanowires with mammalian cells. J. Am. Chem. Soc. 129, 7228–7229 (2007).
Qi, S., Yi, C., Ji, S., Fong, C.-C. & Yang, M. Cell adhesion and spreading behavior on vertically aligned ailicon nanowire arrays. ACS Appl. Mater. Interfaces 1, 30–34 (2009).
Brammer, K. S., Choi, C., Oh, S., Cobb, C. J., Connelly, L. S., Loya, M., Kong, S. D. & Jin, S. Antibiofouling, sustained antibiotic release by Si nanowire templates. Nano Lett. 9, 3570–3574 (2009).
Schlie–Wolter, S., Deiwick, A., Fadeeva, E., Paasche, G., Lenarz, T. & Chichkov, B. N. Topography and coating of platinum improve the electrochemical properties and neuronal guidance. ACS Appl. Mater. Interfaces 5, 1070–1077 (2013).
Xie, C., Hanson, L., Xie, W., Lin, Z., Cui, B. & Cui, Y. Noninvasive neuron pinning with nanopillar arrays. Nano Lett. 10, 4020–4024 (2010).
Albuschies, J. & Vogel, V. The role of filopodia in the recognition of nanotopographies. Sci. Rep. 3, 1658–1666 (2013).
Anselme, K. Osteoblast adhesion on biomaterials. Biomaterials 21, 667–681 (2000).
Pennisi, C. P., Dolatshahi-Pirouz, A., Foss, M., Chevallier, J., Fink, T., Zachar, V., Besenbacher, F. & Yoshida, K. Nanoscale topography reduces fibroblast growth, focal adhesion size and migration-related gene expression on platinum surfaces. Coll. Surf. B 85, 189–197 (2011).
Kunzler, T. P., Huwiler, C., Drobek, T., Vörös, J. & Spencer, N. D. Systematic study of osteoblast response to nanotopography by means of nanoparticle-density gradients. Biomaterials 28, 5000–5006 (2007).
Zimmerberg, J. & Kozlov, M. M. How proteins produce cellular membrane curvature. Nat. Rev. Mol. Cell Biol. 7, 9–19 (2006).
Li, Z., Song, J., Mantini, G., Lu, M.-Y., Fang, H., Falconi, C., Chen, L.-J. & Wang, Z. L. Quantifying the traction force of a single cell by aligned silicon nanowire array. Nano Lett. 9, 3575–3580 (2009).
Song, J., Wang, X., Riedo, E. & Wang, Z. L. Elastic property of vertically aligned nanowires. Nano Lett. 5, 1954–1958 (2004).
Gordon, M. J., Baron, T., Dhalluin, F., Gentile, P. & Ferret, P. Size effects in mechanical deformation and fracture of cantilevered silicon nanowires. Nano Lett. 9, 525–529 (2009).
Arnold, M., Hirschfeld-Warneken, V. C., Lohmüller, T., Heil, P., Blümmel, J., Cavalcanti–Adam, E. A., López–García, M., Walther, P., Kessler, H., Geiger, B. & Spatz, J. P. Induction of cell polarization and migration by a gradient of nanoscale variations in adhesive ligand spacing. Nano Lett. 8, 2063–2069 (2008).
Deeg, J. A., Louban, I., Aydin, D., Selhuber–Unkel, C., Kessler, H. & Spatz, J. P. Impact of local versus global ligand density on cellular adhesion. Nano Lett. 11, 1469–1476 (2011).
Wang, X., Yan, C., Ye, K., He, Y., Li, Z. & Ding, J. Effect of RGD nanospacing on differentiation of stem cells. Biomaterials 34, 2865–2874 (2013).
Lamoureux, P., Buxbaum, R. E. & Heidemann, S. R. Direct evidence that growth cones pull. Nature 340, 159–162 (1989).
Zheng, J., Lamoureux, P., Santiago, V., Dennerll, T., Buxbaum, R. E. & Heidemann, S. R. Tensile regulation of axonal elongation and initiation. J. Neurosci. 11, 1117–1125 (1991).
Lamoureux, P., Ruthel, G., Buxbaum, R. E. & Heidemann, S. R. Mechanical tension can specify axonal fate in hippocampal neurons. J. Cell Biol. 159, 499–508 (2002).
Acknowledgements
This work was supported by Nano-Material Technology Development Program (2012M3A7B4034986) funded by the National Research Foundation and the Pioneer Research Center Program through the National Research Foundation of Korea funded by the Ministry of Science, ICT and Future Planning (2012-0009562). In addition, it was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT and Future Planning (2015R1A2A2A01005556).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the NPG Asia Materials website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Park, YS., Yoon, S., Park, J. et al. Deflection induced cellular focal adhesion and anisotropic growth on vertically aligned silicon nanowires with differing elasticity. NPG Asia Mater 8, e249 (2016). https://doi.org/10.1038/am.2016.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/am.2016.5
This article is cited by
-
Tutorial: using nanoneedles for intracellular delivery
Nature Protocols (2021)
-
Nanobiotechnology: 1D nanomaterial building blocks for cellular interfaces and hybrid tissues
Nano Research (2018)