Abstract
Miniaturization has been the key driver for many remarkable technological developments in recent decades. Miniaturization has now also extended into biology, thereby setting the stage for high-throughput single-cell analysis. This advancement is important because, despite detailed molecular information on individual cell subtypes, virtually no information is available on the functional capacities of individual cells. Typical in vivo animal models, as well as in vitro laboratory test tube experiments, only yield a global outcome of interactions of often millions of cells rather than providing insight into the functional contribution of individual cells. Reaction volumes of biological experiments have generally been reduced from milliliters to microliters. Tools and methods that study single-cell behavior have become increasingly important, but often do not allow for high-throughput manipulation. Recent advances in (droplet-based) microfluidics enable systematic high-throughput analyses of individual cells in a highly controlled manner. The implementation of microfluidic technologies in single-cell analysis is one of the most promising approaches that not only offers new information and high-throughput screening but also enables the creation of innovative conditions that are impractical or impossible by conventional methods. In this review, we provide a comprehensive overview of recent developments in droplet-based microfluidics for single-cell studies.
Similar content being viewed by others
Introduction—Single-cell studies
Cells are life’s atoms—the basic units of all life; a single cell can grow and develop into a whole organism; a single cell cultured on an agarose plate can form a bacterial colony; a single cell can cause a disease; a single cell can give us insight into how life really functions. Arguably, the very first observation of algae and flagellates in water droplets by Antonie van Leeuwenhoek1 showed that single-cell studies can result in interesting discoveries. The importance of studying individual cells has long been realized. However, available technology has persistently remained a limiting factor and hampered the study of large numbers of cells in sufficient quantitative detail. The development of new technologies in the area of high-throughput single-cell analysis has revolutionized our understanding of how complex cellular and molecular systems operate.2, 3, 4, 5
The most widely used tool for high-throughput single-cell studies is fluorescence-based flow cytometry.6 Flow cytometry provides information on cellular size and expression levels of surface and intracellular molecules at a very high throughput (50 000–100 000 cells). Although flow cytometry is mainly used for end point measurements, it also possesses the unique feature of sorting cells according to their characteristics. The development of ‘CyTof’ and the ‘Imagestream’ has greatly enhanced the power of flow cytometry.7, 8, 9, 10 The CyTof apparatus integrates single-cell fluidics with mass spectrometry, which increases the number of detectable parameters to be measured on a single cell to over a hundred non-overlapping metal-isotope labeled antibodies.7
At the same time, ‘traditional’ fluorescence microscopy can now be automated using high-throughput image analysis and used, for example, to obtain expression profiles for thousands of single cells.11 However, one of the major drawbacks of these techniques and approaches is that they preclude dynamic monitoring of (single) cellular responses over time and do not allow for the manipulation of single cells. Nanoliter-scale wells, developed by the Love group,12, 13, 14 are emerging as a promising tool for studying single-cell behavior that overcome a number of the reported drawbacks of more conventional techniques. These technologies have been extensively reviewed recently and therefore will not be further addressed here.15 In recent years, microfluidic tools that allow for the study of hundreds to a few thousands of individual cells under precisely defined conditions have also emerged.16, 17, 18 The microfluidic technology offered by Fluidigm is the current state-of-the-art platform for single-cell analysis. This elastomeric valves-based microfluidic technology enables highly parallel RNA and DNA analysis of samples containing only a few hundred cells.19 By integrating and simplifying multiple steps in the single-cell workflow, the technology allows for the rapid isolation, processing and profiling of individual cells for multiple genomics applications, such as, performing parallel measurements of expression levels up to 96 genes per single cell20 and preparing samples for single-cell RNA sequencing experiments.21 The strong advantages of this system are sample and reagent efficiency, a precision impossible to achieve with liquid-dispensing methods and a broad spectrum of available assays. Although the throughput is relatively high in comparison with that offered by traditional methods of studying single cells, the main challenge will be in scaling up this platform.
In this review, we wish to focus on an alternative microfluidic tool for single-cell analysis that has emerged in recent years. Water-in-oil droplets have been used to compartmentalize individual cells and to study antibody production by single cells dating back to the 1950s,22 and recent advances in droplet microfluidics have greatly expanded their potential for single-cell studies.23, 24, 25, 26, 27 Indeed, these picoliter droplets can be considered ‘nanolabs’ that accommodate both single cells and all factors secreted by those cells.16,28,29 These technologies complement and augment the types of data generated by flow-cytometry-based systems. Single-cell analysis typically requires a complex workflow involving nucleic acid isolation, amplification and fluorescence-based detection. Fluidic tools must therefore be equipped to carry out multistep procedures. Droplet-based microfluidics is technically less advanced than valve one-phase microfluidics, but many of the individual steps have already been demonstrated. First, droplets can be collected from a microfluidic device and incubated off-chip.30 Incubation on-chip is also possible either in delay lines,31 in trap32 or in chamber devices.33 The possibility of increasing the residence time for droplets in microfluidic devices is crucial for studying biological processes that occur on a timescale of minutes to hours, or even days. Emulsions collected after off-chip incubation can be reinjected into another microfluidic device for further manipulation.34 Moreover, additional reagents can be added to a droplet at well-defined points in space and time by fusion35 or picoinjection.36 Splitting of droplets, in turn, offers the opportunity to perform multiple assays on a single droplet and to dilute the content of droplets without increasing the reaction volume.37 Droplets can also be sorted passively because of the difference in their size38,39 or actively25 via fluorescence-activated droplet sorting, which allows for the separation of droplets of interest from the rest of the droplet stream. Moreover, a wide spectrum of on-chip detection methods is available for the droplet format, including fluorescence measurements,40 ultraviolet-visible spectroscopy41 and electrochemical detection.42 These ‘microtools’ provided by droplet microfluidics, alone or in conjunction with traditional single-cell technologies, are opening many new avenues of research into the function of single cells (Figure 1). In this review, we provide a comprehensive overview of recent developments in droplet-based microfluidics for single-cell studies.
What can we do with droplets containing single cells and to single cells inside droplets?
Single cell encapsulation
The most straightforward method for encapsulating single cells is by simply forming emulsions of cell suspensions and an oil phase containing surfactants (Figure 2b).22,43,44 The undeniable advantage of simplicity offered by this method, however, is outweighed by the polydispersity of the resulting emulsion, which makes quantitative comparisons between individual cell-containing droplets rather complex.43 Therefore, in the majority of current studies on single cells in droplets, researchers opt to employ microfluidic droplet generators for single-cell encapsulation. Although the monodispersity of droplet emulsions can be tightly controlled by microfluidic devices, there are certain minor experimental obstacles that need to be addressed in this respect. Certain types of cells have a tendency to aggregate, leading to blockages and irregular compartmentalization of cells in droplets. Furthermore, regardless of cellular aggregation, the sedimentation of cells poses a problem because it can lead to the loss of single-cell encapsulation due to the formation of plugs of cells in microfluidic tubing. By matching of the density of the supporting fluid, sedimentation times can be increased and aggregation reduced.28 Several elegant microfluidic designs have been suggested to support the elimination of this type of problem, for example by employing polydimethylsiloxane (PDMS) discs with a biopsy punched hole instead of needles as a means to prevent cell loss in the needle45 or to incorporate mixing during injection by placing a miniature stir bar inside the syringe.46,47
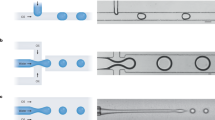
Methods of encapsulation of single cells in droplets. (a) Stochastic encapsulation inside flow-focusing device.48 (b) Emulsification via macro-scale agitation. (The mean droplet diameter is 43 μm.)44 (c) Dielectrophoretic droplet sorting after encapsulation. When an electric field is applied across the electrodes, the droplets are deflected into the positive arm. In the absence of a field, the droplets flow into the negative arm owing to the lower hydraulic resistance (inset).25 (d) Encapsulation based on cell-triggered Rayleigh-Plateau instability.39 (e) Ordered cell encapsulation.48 The length of the scale bars in each figure is 100 μm, except for (b) and (d), where the length is specified otherwise. All figures have been reproduced with permission.
Although the encapsulation of cells in droplets is relatively easy and the encapsulation efficiency can be high, single-cell encapsulation has certain inherent challenges. Diluted cell suspensions and random cell encapsulation results in a Poisson distribution of encapsulated cells, and if the average number of cells per droplets is low enough, the fraction of droplets containing more than one cell is negligible. Unfortunately, at the same time, the fraction of empty droplets is significant, lowering the throughput of the method and making analysis more difficult (Figure 2a).48 Stochastic encapsulation becomes even more problematic in experiments in which two single objects need to be encapsulated in a single droplet.49 Therefore, attempts have been made to overcome the abovementioned difficulties.
Active droplet sorting
One possible way to increase the ratio of cell-containing to empty droplets without increasing the fraction containing multiple cells is to actively sort droplets after encapsulation (Figure 2c).25 Droplet sorting is possible by charging droplets and sorting them in an electric field,50 by dielectrophoresis,51 or by localized heating by a laser.52 Active sorting of droplets containing cells, however, has to date only been demonstrated by means of dielectrophoresis. The presence of cells inside droplets can be detected in several ways. One possibility is to use fluorescently labeled cells for encapsulation and subsequently perform fluorescence-activated droplet sorting.25,28 Additionally, discrimination between empty and cell-containing droplets can also be achieved by electrical impedance measurements53,54 or by sorting cells in microfluidic channels based on the Clausius–Mossotti effect, which has been demonstrated for both continuous-phase55 and droplet microfluidics.56 Although the active sorting of cells requires a relatively complex setup, the number of recently published studies in this field indicates the feasibility of this approach for single-cell droplet experiments.
Passive methods for increasing single-cell encapsulation efficiency
Another possible way to increase the efficiency of single-cell encapsulation by means of droplet microfluidics is by employing hydrodynamic methods enabling the control over the number of cells encapsulated per droplet or self-sorting of droplets containing cells from smaller, empty droplets. These methods utilize physical phenomena such as cell-triggered Rayleigh-Plateau instability (Figure 2d)39 or self-organization occurring when a high-density suspension is forced to travel rapidly through a microchannel (Figure 2e)48 and provide an efficiency of single-cell encapsulation of approximately 80%. Size-based sorting can also be achieved by exploiting cell-induced droplet shrinking, in which the osmotic water flow that occurs between droplets containing metabolically active yeast cells and empty droplets leads to shrinkage of cell-containing droplet reservoirs.38,57 Nonetheless, the only method used to date in droplet-based functional studies on single cells is fluorescence-activated droplet sorting.
In general, improvements in single-cell encapsulation and manipulation introduced by microfluidics have aided in the development of new analytical techniques, such as assays for detecting antibodies secreted from single mouse hybridoma cells developed by Mazutis et al.28 However, more work still needs to be invested in the development of methods for increasing single-cell encapsulation efficiency to make more of these methods applicable for single-cell analysis protocols.
Manipulation of single cells inside droplets
Cell culture inside droplets
Encapsulation of cells in small droplets might at first seem at odds with ensuring that cells remain healthy, but there is now ample evidence that demonstrates the biocompatibility of droplets for cellular studies compared to that of cells cultivated in bulk suspensions, provided that suitable media, non-ionic surfactants and gas-permeable carrier fluids are used,58,59 or, when necessary, a solid support is provided inside the droplet in the form of a hydrogel matrix. Examples include multiple human cell lines (Figure 3a),58, 59, 60 yeast (Figure 3b),61 numerous types of bacteria (Figure 3c)32,44,62 and algae (Figure 3d).63 Several designs of microfluidic devices facilitating this type of culture have been published. Such designs include the on-chip microdroplet chemostat, which allows for the automated exchange of culture media during droplet cell culture without interfering with compartmentalization and monitoring of bacterial growth inside droplets64 and an integrated modular system for on-chip cell immobilization, culturing, transfection and transfer into droplets.65 However, of special interest is the possibility of culturing monoclonal microbial populations and single cells in isolation from other cells by droplet microfluidics.44,60,62,63
Cell culture inside droplets. (a) Fluorescence microscopy image of drops (660 nl) hosting lacZ-expressing human cells (HEK293T) converting the fluorogenic substrate FDG after 16 h of incubation.58 (b) Droplets containing Saccharomyces cerevisiae after encapsulation.38 (c) Image of Escherichia coli cells in 36-μm diameter agarose microparticles after 8 h of culture.83 (d) Chlamydomonas reinhardtii cells cultured in microdroplets.63 All figures have been reproduced with permission.
Microdroplet generators are also used to produce cell-laden microgels from molten hydrogels or monomer solutions, after which emulsions are broken and beads resuspended in cell culture medium. To date, various types of hydrogels have been utilized for that purpose, for example, alginate,66 agarose,67 synthetic peptides,68 polyethylene glycol69 and hyperbranched polyglycerol.70 The major advantage of cell culture in microgels, compared with cell culture in droplets, is the ease of exchanging the culture media simply by transferring porous microgels into fresh medium, which allows for prolonged cell culture without the adverse effects caused by the accumulation of metabolites. Nevertheless, this approach also poses a disadvantage, because soluble secreted molecules can cross-contaminate other cell-containing microgels, thereby influencing cellular responses. There is also the possibility of culturing adherent cells inside microgels prepared from hydrogels containing adhesive substrates for cells71 or studying the influence of environmental properties, such as hydrogel elasticity, on cell fate by using hydrogels that have different stiffnesses.67,72
For biological applications, the recovery of cells or particles from droplets after an experiment or assay is desirable. The most commonly described approach for retrieving cells from emulsions is by the addition of 1H,1H,2H,2H-perfluoro-1-octanol (Figure 4).28,72 Nonetheless, in their paper, the authors specify that particular caution needs to be exercised when performing this type of demulsification and that prolonged exposure to 1H,1H,2H,2H-perfluoro-1-octanol should be avoided.28 Another approach that circumvents the harsh chemical treatments and vigorous washing steps uses an electro-demulsification chip, which offers a mild way for extracting cells and polymer particles into an aqueous phase from droplet microfluidic water-in-oil emulsions.73 Furthermore, it is also possible to retrieve cells from cell-laden agarose gel beads by agarase treatment.74 The recovery of cells from microgels or droplets allows for upstream analysis to be performed on exactly the same cell population that was used in microfluidic experiments.
Survival rate of cells after incubation in droplets and release from the emulsion.72 (a) Phase image of human mesenchymal stem cells (hMSC) spheroids encapsulated in double emulsion droplets after 6 h. (b) Live/dead staining of spheroids after release from emulsion at 6 h using 1H,1H,2H,2H-perfluoro-1-octanol. Live cells were labeled with calcein AM (green), and dead cells were labeled with propidium iodide (red).
Cell lysis inside droplets
Cell lysis is often the first step that molecular biologists take to quantify protein or mRNA levels, intracellular enzymes activity, or the genetic material that cells contain. Cell lysis allows for insight into the complexity of intracellular processes and for studying the subject of interest, regardless of whether it is the proteome, transcriptome or genome. The lysis of cells can be carried out using physical or chemical lysis methods.75 One method for conducting the physical lysis of cells encapsulated in microfluidic droplets is optical lysis,76 in which laser-induced plasma formation and the subsequent generation of a shock wave, followed by a cavitation bubble, results in cellular disruption (Figure 5).77 The main advantage of this approach is that the speed of the lysis allows for a ‘snapshot’ of the activity of a cell to be obtained at a particular time point.76 Another method suggested for studying the release of genomic DNA within droplets is a heating step preceding downstream analysis.32,49 Commercial78,79 and home-made detergent-based or enzymatic lysis agents18,30 have been applied to study the content released from single cells. For example, to release genomic DNA or mRNA, proteinase K digestion and sodium dodecyl18 or Tween 2030 lysis have been demonstrated to be useful for the droplet-microfluidics platform. However, not all available lysis methods can be applied in this manner. Physical lysis methods in particular appear to be challenging because it is relatively easy to perturb monodisperse water-in-oil emulsions. For example, the sonication of emulsions by using high sound energy may lead to the introduction of a polydisperse population of droplets instead of the desired monodisperse population. The introduction of coalescence/splitting-induced polydispersity may influence the results of subsequent analysis by potentially cross-contaminating single-cell-containing droplets or might lead to the generation of two droplets containing the information of a single cell. In summary, the repertoire of cell lysis methods available for the droplet-microfluidics platform is broad, with many of these methods already having been successfully applied, but there is still plenty of room for the improvement and development of new methodologies.
Fast photolysis of a single cell in an aqueous droplet. (a–d) Video sequence acquired by a fast camera. Cell lysis was caused by laser-induced plasma formation and cavitation bubbles. The scale bar in (a) applies to all panels. Reproduced with permission from reference.76
Transfection of single cells
The possibility of transfecting eukaryotic cells (i.e., delivering nucleic acids such as plasmid DNA or RNA into living cells) is important for molecular biology studies because it allows for the introduction of genes of interest in target cells and for genetically modified organisms to be obtained to study specific gene functions. Similarly to cell lysis methods, methods that introduce foreign DNA into a eukaryotic cell fall into two categories: physical (such as electroporation or particle-based methods) and chemical methods.80 Transfection of cells inside microfluidic droplets has to date only been reported by two groups.81,82 Zhan et al.81 described for the first time the electroporation of cells encapsulated in droplets to transfect them with plasmid DNA coding for fluorescent proteins (Figure 6). The efficiency of the authors’ process was unfortunately not very high; in fact, the efficiency was estimated to be only 11%. Improvement in the transfection efficiency to the values known to be possible via bulk transfection (~25%) was achieved by applying chemical transfection reagents to the droplet format.82 The uptake of DNA by cells was possibly because of the formation of complexes of DNA with their carrier, in this case a cation-activated dendrimer. During incubation, those complexes were nonspecifically endocytosed by the cells. Interestingly, the authors observed a higher transfection efficiency in small droplets than in large droplets, which may be explained by the beneficial effects of confinement, which increase the probability of interaction between cells and the DNA/carrier complexes. The mechanism of this phenomenon, however, remains to be elucidated. These promising findings urge for the development and application of other transfection protocols known from bulk experiments in droplet-based microfluidics.
A microfluidic device for encapsulation of cells in aqueous droplets and their subsequent electroporation. (a) Electroporation occurs when the cell containing droplets in oil flow through a pair of microelectrodes with a constant voltage established in between. (b) The droplets with encapsulated cells after electroporation at the exit of the device. Reproduced with permission from reference.81
Functional studies on single cells
Drug studies on single cells in droplets
The first examples of single-cell analysis in droplets were demonstrated in pharmacological experiments. In those studies, researchers took advantage of the reduction in cost because of miniaturization, automation and a confinement-related increase in concentration of compounds. For example, in the study by Boedicker et al.,62 a plug-based microfluidic technology was applied for the rapid determination of antibiograms from samples featuring complex biological matrices. Antibiograms are charts of the sensitivity of a bacterial strain to different antibiotics. In their experiments, the authors co-encapsulated cells together with a viability indicator. A significant reduction in the incubation time was achieved by effectively increasing both the cell concentration and the products of viability-testing reactions because of the confinement of single cells into small-volume compartments. In general, the detection time in this system is proportional to the plug volume. Another similar agarose microparticle-based platform for antibiotic studies was suggested for determining the minimum inhibitory antibiotic concentration. This platform, which also allowed for the isolation and characterization of spontaneous mutants resistant to antibiotics, required less than 1 mg of compound and less than 3 h of incubation, thereby providing an interesting alternative to bulk bacteria cultures.83
Drug studies have also been conducted on single mammalian cells confined in droplets. The workflow presented by Brouzes et al. (Figure 7)60 allowed for cell cytotoxicity screenings at a very high throughput. To support the real-time identification of droplet composition and post-processing data analysis, the authors introduced a fluorescent coding scheme. A similar coding strategy was used in a more complex analysis in which a reporter gene assay was used to test the dose–response profiles of single Bombyx mori cells to an insecticide.45 Coding methods serve as a promising tool for droplet microfluidics because they offer a solution to the absence of spatial positioning, which is typical for microtiter plate experiments. Experiments reported by Baret et al.,45 Brouzes et al.60 or Joensson et al.84 provide evidence of the potential of fluorophore mixtures as barcodes for oil-in-water emulsions. Other currently available and possible barcoding systems for the droplet format still await benchmarking before they can be applied. DNA can be considered a promising candidate for a barcode system for single-cell experiments in droplets because its utility in single-cell studies has recently been proven by experiments involving the barcoding and sequencing of transcriptome from single cells.85 Although DNA is technically more challenging as a barcode than fluorophores because it does not allow for the collection of online data, it is certainly worth further exploration because it can overcome library-size-related limitations.86
Schematic depiction of droplet screening workflow involving four steps. (a) The droplet library is formed by emulsifying drug library using the library generation chip. Each droplet is uniquely coded with an optical label. (b) Each droplet library member is merged with a cell-containing droplet. (c) The emulsion is incubated for cell treatment. (d) The emulsion is reinjected into the assay chip, where fluorescence of each droplet is measured for drug coding readouts and viability assay. Reproduced with permission from reference.60
Single-cell genomics
Protein as well as mRNA levels show a large cell-to-cell variation both in a resting state and after exposure to stimuli, even in cells bearing the same genotype.3,87 Moreover, in many fields, minority subpopulations of cells are often the subject of interest to the scientific community. This statement applies particularly to stem cell science and tumor immunology, in which cells of interest often comprise only a small fraction of cells in a sample. One example of such cells are circulating tumor cells that are found in frequencies as low as 1–10 cells per ml of whole blood (which roughly contains five billion cells) in patients with metastatic disease.88,89 Therefore, not surprisingly, a significant amount of research involving the encapsulation of single cells in microdroplets is focused on studying the cells’ genome or transcriptome; an overview of studies performed in this area in recent years is provided in Table 1.
One of the first examples of single-cell genetic analysis in microfluidic droplets was reported by Kumaresan et al.,49 who presented a novel platform that combines the power of two techniques: droplet-based microfluidics and flow cytometry. In those experiments, single cells were encapsulated together with primer-functionalized microbeads and a PCR mix. As a result of the PCR reaction, dye-labeled product attached to the bead surface was produced. Subsequently, the emulsion was broken, and the product-bearing beads were released and either were analyzed by flow cytometry or served as a template for sequencing. Further development of this platform 2 years later enabled a higher throughput and a higher number of genes that could be analyzed per cell.90 Another improvement made for single-cell multiplex gene detection and sequencing was the use of agarose droplets as reactors for single-cell DNA amplification to address the challenge of inefficient and irreproducible DNA extraction.18 The encapsulation in agarose gel beads provided, after breaking the emulsion, the opportunity to isolate and manipulate isolated genomic DNA without mixing the genetic content of different cells. Owing to the pore size of agarose gels, large genomic DNA is prevented from diffusing out from the beads, whereas the diffusion of lysis buffer and small- and medium-sized proteins into the gel beads is still allowed.
As mentioned previously, droplet-based microfluidics has also been applied to gene expression analysis.91,92 A serious weakness of the majority of protocols for single-cell transcriptome analysis in droplets is the lack of the high throughput expected from emulsion-based approaches. This limitation is caused by the relatively large size of droplets used in assays, which is necessary to avoid lysate-driven inhibition of RT-PCR and to encapsulate together exactly one cell and primer-bearing bead. The latter demands the use of solutions at limiting dilutions and results in emulsions in which only a very small fraction is occupied by the desired combination of exactly one cell and one bead. Recently reported, however, is a new platform that overcomes the above-described problems by the introduction of a workflow, in which droplets containing single cell lysates are diluted before adding RT-PCR reagents, thus preventing lysate-driven reverse transcriptase inhibition (Figure 8).30 Another important advantage of this method is that sample compartmentalization is retained throughout the entire experiment, allowing for the elimination of false-positive results caused by cell lysis prior to encapsulation.
Microfluidic workflow for ultrahigh-throughput mammalian single-cell RT-PCR. (a, b) A mixed population of human cells is first encapsulated in the droplet generator together with lysis buffer, then incubated off-chip. Subsequently, droplets are reinjected and diluted on-chip by merging with water droplets to avoid lysate-driven RT-PCR inhibition. After dilution, a portion of the lysate is split off and subsequently picoinjected with RT-PCR reagents. (c) Picoinjected drops are collected in a tube and thermocycled. (d) After reaction, the fluorescence of drops is analyzed for transcript amplification. Reproduced with permission from reference.30
The recent developments in droplet microfluidics for single-cell studies have set the stage for employing emulsion-based techniques in common analytical laboratories. As reported by Geng et al.,93 a single-cell forensic short tandem repeat analysis was performed in droplets with improved throughput (compared with that of conventional forensic techniques for genetic single-cell analysis) and simpler sample handling. The next step will certainly be to apply droplet-based microfluidic technology for single-cell genetic analysis to more complex, ‘real-world’ samples.
Single-cell immunology
Droplet-based microfluidics has also opened new opportunities for areas of research such as immunology, providing platforms that allow for high throughput and fast screening of individual cells. Examples of such studies include experiments that have investigated individual hybridoma clones for the release of antibodies inhibiting a drug target based on fluorescence16,28 or that have studied low-abundance cell-surface biomarkers on individual human cells. Svahn and coworkers84 used enzymatic amplifications with the fluorescent product remaining confined inside each droplet and increasing in concentration over time in combination with color codes, allowing for concurrent analysis of multiple samples. This method provided a higher signal resolution above background levels than could be reached by standard flow cytometry.
Until recently, one of the major drawbacks of using droplet-based microfluidics for single-cell immunology studies was the lack of the ability to wash droplets or allow metabolites to diffuse in and out of droplets without risking cross-contamination between cell-containing droplets. Although some studies disregarded this phenomenon and investigated the secretion of molecules by cells by the direct co-encapsulation of detection antibodies, information on the sensitivity of this technique compared with that of conventional methods is currently lacking.94 A solution to this problem was provided in a recent study by Chokkalingam et al.59 in which the authors (re-)introduced the use of low-melting-point agarose during droplet production. By exploiting the thermal properties of low-melting-point agarose, single immune cells could be encapsulated and incubated for up to 36 h, provided that the cells were kept above room temperature. This technique allowed for cells to become stimulated and for secreted molecules to be captured in the same confined environment. Upon cooling the droplets, the agarose solidified, which subsequently facilitated the detection of the parameters of choice by the diffusion of fluorescently labeled detection antibodies. Another aspect that should not be disregarded when working with primary immune cells is the secretion of lipid metabolites by cells or the use of hydrophobic stimuli for cell activation. In standard water-in-oil emulsions, these substances will almost instantly diffuse into the oil layer surrounding the formed droplets. A solution to this problem was recently suggested by Zinchenko et al.,95 namely, the generation of double emulsion droplets. Such droplets are generated by using two devices in series, the first of which has hydrophobic properties and the second hydrophilic.95 This approach is technologically more demanding than using simple oil-in-water emulsion droplets; however, it will ensure that all components and metabolites remain confined.
Although still in its infancy, the implementation of the above-described approaches in combination with the various techniques available for droplet manipulation hold great potential for exploring and revealing currently unknown heterogeneities present in what have traditionally been considered to be homogeneous immune cell populations.
Evolution of microorganisms in droplets
Directed evolution experiments are of great importance not only in basic research, where they provide information about the function and structure of proteins, but also in biotechnology because they help in obtaining increasingly efficient enzymes and microorganisms. To study evolution in the laboratory, mutations are first artificially introduced into a population. Such a population is subsequently exposed to a controlled evolutionary pressure to select variants with improved characteristics. Droplets of water in oil provide the perfect platform for evolutional studies because they enable the coupling of a phenotype with a genotype, which is necessary because evolutionary information is contained in the genome and the selection is performed with regard to phenotype. The compartmentalization of cells in droplets is especially beneficial for single-cell enzyme catalysis studies, which were previously limited to studying cases in which the diffusion of the product from the cell is restricted.26 Droplets of water in oil not only allow for the accumulation of products of single-cell enzymatic reactions in the same compartment as the genome but also provide an opportunity for the amplified detection of single enzymatic events from single cells.96 In a recently reported assay, rolling circle amplification serves as an amplification method and empowers the detection of single DNA cleavage-ligation events.96 Moreover, droplet microfluidics allows for fast screening of large libraries (108 in 10 h),97 and studies focusing on the binding affinity of proteins have shown that the size of the library screened is correlated with the improvement achieved in the affinity of the selected proteins.98 Similarly, these types of correlations are also expected in studies on enzymes.97
Research began to use water-in-oil emulsions in directed evolution experiments more than 15 years ago,99 whereas cells were only introduced for the first time by Aharoni et al.43 to increase the concentration of enzymes in droplets because more protein is produced via cell-based expression in droplets than via in vitro expression used in prior experiments. Additionally, the introduction of cells into droplets enabled studies involving whole genome mutagenesis,100 which is an interesting approach because the metabolism of the cell is highly complex and not yet fully understood. Increased yields of enzyme production can be achieved not only by improving the enzyme itself but also by improving the cellular functions of the whole microbial organism, leading to more efficient hosts for enzyme production.100
Although the method introduced by Aharoni et al.43 proved to be successful in the selection of improved variants of an enzyme, it has not been widely used because of the heterogeneity introduced by double water-in-oil-in-water emulsions prepared by two steps of homogenization. Indeed, differences in the sizes of droplets strongly affect the concentration of the enzymes, substrates and products inside droplets, thereby reducing the accuracy of the obtained results.
Employing only emulsification via microfluidics and fluorescence-activated droplet sorting as the method for screening droplets led to the full demonstration by Agresti et al.97 and Kintses et al.78 of the benefits of emulsion experiments performed in droplets. The screening of populations of 107 cells in only several hours allowed for the selection of mutants demonstrating a 10-fold increase in their catalytic rate97 and 6-fold increase in the activity and expression of the enzyme of interest.78 One outstanding example of research that would be very difficult to perform without employing the encapsulation of cells in water-in-oil emulsions is that described in a study performed by Bachmann et al.,44 in which the authors exploited compartmentalization in droplets to eliminate competition between individual cells. The serial transfer of a bacteria population into droplets of water in oil made it possible to isolate high-yield mutants that otherwise were outperformed by fast-growing variants. These results support the hypothesis that the availability of public goods has an impact on evolutionary outcomes and confirm the utility of water-in-oil emulsions as a platform for evolutionary studies that may promote new discoveries in biology.
Conclusions and perspective
In this review, we have summarized the available literature and have discussed the strong potential for the use of droplet-based microfluidics systems in single-cell studies. The development of microfluidics, specifically droplet microfluidics, has led to many interesting discoveries, and the number of published studies exploiting droplet microfluidics is steadily increasing every year.23, 24, 25, 26, 27, 28,30 Of all available technologies, droplet-based microfluidics is one of the most powerful approaches to studying the functional behavior of single cells. Although single cells can easily be studied using flow cytometry, it is typically not possible to associate secreted molecules with a particular cell. On the other hand, the dimensions of the microwells traditionally used for single-cell culture (384-well plates) are too large, causing all secreted metabolites to be highly diluted and making it impractical to detect these metabolites by conventional techniques. Droplet-based microfluidics enables the encapsulation and activation of single cells in high throughput and also ensures that soluble metabolites secreted by activated cells are confined in a relatively small volume, which allows for a large reduction in noise and thereby increases the sensitivity of the technique. One of the major obstacles that have hampered rapid progression in this field has been, for many years, the inability to perform washing steps inside droplets, which is crucial for the controlled amplification of DNA and for many other biological protocols. Several solutions to this problem have been suggested, including the use of low-melting-temperature agarose and hydrogels.59,72 The introduction of hydrogels and the washing thereof has surely improved the prospects of using droplet microfluidics;59 however, there is still need for the development of new and smart biosensors such as aptamers or Förster Resonance Energy Transfer (FRET)-based sensors that will circumvent the need to wash droplets and will allow for the dynamic monitoring of responses over time.
Furthermore, the possibility of manipulating cell-containing droplets by sorting, splitting and fusion will also enable scientists to link the functional characteristics of single cells and the effect these characteristics have on the activation of co-encapsulated cells. In this manner, scientists can study how cellular heterogeneity affects cell–cell communication and cellular decision making.
Another example that highlights the importance of emulsion-based single-cell studies for solving practically impossible research questions comes from the field of directed evolution. Conventional approaches to studying evolution can be obscured by populations of cells harboring mutations that rapidly overgrow less rapidly dividing cell populations, a problem easily solved by single-cell encapsulation.44 Moreover, and not unimportant, droplet microfluidics leads to a reduction in the time and labor requirements and thus the costs involved in directed evolution experiments by decreasing their timescale from years to hours.97
Currently, the manipulation of (cell-containing) droplets requires a relatively complex setup and involves the sequential use of numerous different modules, which makes it challenging to generate robust experimental pipelines. Furthermore, droplet-based microfluidics still mostly relies on conventional ‘end point’ measurement tools, which means that the droplets generated must be compatible, for example, with respect to size and composition, with those measurement tools. Notwithstanding, technical challenges are currently being addressed and as a result are rapidly dissolving, which should enable biologists, pharmacists and immunologists to increasingly apply and implement this promising technology as a routine experimental procedure in years to come.
References
Dobell CC & van Leeuwenhoek A Antony Van Leeuwenhoek and His "Little Animals" (Brace and company, New York, USA, 1932).
Brehm-Stecher BF & Johnson EA Single-cell microbiology: Tools, technologies, and applications. Microbiol. Mol. Biol. R 68, 538–559 (2004).
Elowitz MB, Levine AJ, Siggia ED & Swain PS Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Fleming A On the antibacterial action of cultures of a penicillium, with special reference to their use in the isolation of B. influenzae. Bull World Health Organ 79, 780–790 (1929).
Itzkovitz S & van Oudenaarden A Validating transcripts with probes and imaging technology. Nat. Methods 8, S12–S19 (2011).
Shapiro HM in Practical Flow Cytometry 73–100 (John Wiley & Sons, Inc., 2005).
Newell EW, Sigal N, Bendall SC, Nolan GP & Davis MM Cytometry by time-of-flight shows combinatorial cytokine expression and virus-specific cell niches within a continuum of CD8(+) T cell phenotypes. Immunity 36, 142–152 (2012).
Bjornson ZB, Nolan GP & Fantl WJ Single-cell mass cytometry for analysis of immune system functional states. Curr. Opin. Immunol. 25, 484–494 (2013).
Beum PV, Lindorfer MA, Hall BE, George TC, Frost K, Morrissey PJ & Taylor RP Quantitative analysis of protein co-localization on B cells opsonized with rituximab and complement using the ImageStream multispectral imaging flow cytometer. J. Immunol. Methods 317, 90–99 (2006).
Lee HK, Lund JM, Ramanathan B, Mizushima N & Iwasaki A Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 315, 1398–1401 (2007).
Battich N, Stoeger T & Pelkmans L Image-based transcriptomics in thousands of single human cells at single-molecule resolution. Nat. Methods 10, 1127–1133 (2013).
Love JC, Ronan JL, Grotenbreg GM, van der Veen AG & Ploegh HL A microengraving method for rapid selection of single cells producing antigen-specific antibodies. Nat. Biotechnol. 24, 703–707 (2006).
Han Q, Bagheri N, Bradshaw EM, Hafler DA, Lauffenburger DA & Love JC Polyfunctional responses by human T cells result from sequential release of cytokines. Proc. Natl Acad. Sci. USA 109, 1607–1612 (2012).
Han Q, Bradshaw EM, Nilsson B, Hafler DA & Love JC Multidimensional analysis of the frequencies and rates of cytokine secretion from single cells by quantitative microengraving. Lab Chip 10, 1391–1400 (2010).
Chattopadhyay PK, Gierahn TM, Roederer M & Love JC Single-cell technologies for monitoring immune systems. Nat. Immunol. 15, 128–135 (2014).
El Debs B, Utharala R, Balyasnikova IV, Griffiths AD & Merten CA Functional single-cell hybridoma screening using droplet-based microfluidics. Proc. Natl Acad. Sci. USA 109, 11570–11575 (2012).
Mary P, Chen A, Chen I, Abate AR & Weitz DA On-chip background noise reduction for cell-based assays in droplets. Lab Chip 11, 2066–2070 (2011).
Novak R, Zeng Y, Shuga J, Venugopalan G, Fletcher DA, Smith MT & Mathies RA Single-cell multiplex gene detection and sequencing with microfluidically generated agarose emulsions. Angew. Chem. Int. Ed. 50, 390–395 (2011).
Marcus JS, Anderson WF & Quake SR Microfluidic single-cell mRNA isolation and analysis. Anal. Chem. 78, 3084–3089 (2006).
Moignard V, Macaulay IC, Swiers G, Buettner F, Schütte J, Calero-Nieto FJ, Kinston S, Joshi A, Hannah R, Theis FJ, Jacobsen SE, de Bruijn MF & Göttgens B . Characterization of transcriptional networks in blood stem and progenitor cells using high-throughput single-cell gene expression analysis. Nat. Cell Biol. 15, 363–372 (2013).
Brennecke P, Anders S, Kim JK, Kolodziejczyk AA, Zhang XW, Proserpio V, Baying B, Benes V, Teichmann SA, Marioni JC & Heisler MG . Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Nossal GJV & Lederberg J Antibody production by single cells. Nature 181, 1419–1420 (1958).
Theberge AB, Courtois F, Schaerli Y, Fischlechner M, Abell C, Hollfelder F & Huck WTS Microdroplets in microfluidics: an evolving platform for discoveries in chemistry and biology. Angew. Chem. Int. Ed. 49, 5846–5868 (2010).
Huebner A, Srisa-Art M, Holt D, Abell C, Hollfelder F, deMello AJ & Edel JB Quantitative detection of protein expression in single cells using droplet microfluidics. Chem. Commun. 1218–1220 (2007).
Baret JC, Miller OJ, Taly V, Ryckelynck M, El-Harrak A, Frenz L, Rick C, Samuels ML, Hutchison JB, Agresti JJ, Link DR, Weitz DA & Griffiths AD Fluorescence-activated droplet sorting (FADS): efficient microfluidic cell sorting based on enzymatic activity. Lab Chip 9, 1850–1858 (2009).
Huebner A, Olguin LF, Bratton D, Whyte G, Huck WTS, de Mello AJ, Edel JB, Abell C & Hollfelder F . Development of quantitative cell-based enzyme assays in microdroplets. Anal. Chem. 80, 3890–3896 (2008).
Courtois F, Olguin LF, Whyte G, Theberge AB, Huck WTS, Hollfelder F & Abell C Controlling the retention of small molecules in emulsion microdroplets for use in cell-based assays. Anal. Chem. 81, 3008–3016 (2009).
Mazutis L, Gilbert J, Ung WL, Weitz DA, Griffiths AD & Heyman JA Single-cell analysis and sorting using droplet-based microfluidics. Nat. Protoc. 8, 870–891 (2013).
Miller OJ, Harrak AE, Mangeat T, Baret JC, Frenz L, Debs BE, Mayot E, Samuels ML, Rooney EK, Dieu P, Galvan M, Link DR & Griffiths AD High-resolution dose–response screening using droplet-based microfluidics. Proc. Natl Acad. Sci. USA 109, 378–383 (2012).
Eastburn DJ, Sciambi A & Abate AR Ultrahigh-throughput mammalian single-cell reverse-transcriptase polymerase chain reaction in microfluidic drops. Anal. Chem. 85, 8016–8021 (2013).
Frenz L, Blank K, Brouzes E & Griffiths AD Reliable microfluidic on-chip incubation of droplets in delay-lines. Lab Chip 9, 1344–1348 (2009).
Leung K, Zahn H, Leaver T, Konwar KM, Hanson NW, Page AP, Lo CC, Chain PS, Hallam SJ & Hansen CL . A programmable droplet-based microfluidic device applied to multiparameter analysis of single microbes and microbial communities. Proc. Natl Acad. Sci. USA 109, 7665–7670 (2012).
Courtois F, Olguin LF, Whyte G, Bratton D, Huck WTS, Abell C & Hollfelder F An integrated device for monitoring time-dependent in vitro expression from single genes in picolitre droplets. Chembiochem 9, 439–446 (2008).
Lee M, Collins JW, Aubrecht DM, Sperling RA, Solomon L, Ha JW, Yi GR, Weitz DA & Manoharan VN Synchronized reinjection and coalescence of droplets in microfluidics. Lab Chip 14, 509–513 (2014).
Chabert M, Dorfman KD & Viovy JL Droplet fusion by alternating current (AC) field electrocoalescence in microchannels. Electrophoresis 26, 3706–3715 (2005).
Abate AR, Hung T, Mary P, Agresti JJ & Weitz DA High-throughput injection with microfluidics using picoinjectors. Proc. Natl Acad. Sci. USA. 107, 19163–19166 (2010).
Link DR, Anna SL, Weitz DA & Stone HA Geometrically mediated breakup of drops in microfluidic devices. Phys. Rev. Lett. 92, 054503 (2004).
Joensson HN, Uhlen M & Svahn HA Droplet size based separation by deterministic lateral displacement-separating droplets by cell-induced shrinking. Lab Chip 11, 1305–1310 (2011).
Chabert M & Viovy J-L Microfluidic high-throughput encapsulation and hydrodynamic self-sorting of single cells. Proc. Natl Acad. Sci. USA. 105, 3191–3196 (2008).
Song H & Ismagilov RF Millisecond kinetics on a microfluidic chip using nanoliters of reagents. J. Am. Chem. Soc. 125, 14613–14619 (2003).
Dolega ME, Jakiela S, Razew M, Rakszewska A, Cybulski O & Garstecki P . Iterative operations on microdroplets and continuous monitoring of processes within them; determination of solubility diagrams of proteins. Lab Chip 12, 4022–4025 (2012).
Han Z, Li W, Huang Y & Zheng B Measuring rapid enzymatic kinetics by electrochemical method in droplet-based microfluidic devices with pneumatic valves. Anal. Chem. 81, 5840–5845 (2009).
Aharoni A, Amitai G, Bernath K, Magdassi S & Tawfik DS High-throughput screening of enzyme libraries: thiolactonases evolved by fluorescence-activated sorting of single cells in emulsion compartments. Chem. Biol. 12, 1281–1289 (2005).
Bachmann H, Fischlechner M, Rabbers I, Barfa N, Branco Dos Santos F, Molenaar D & Teusink B Availability of public goods shapes the evolution of competing metabolic strategies. Proc. Natl Acad. Sci. USA 110, 14302–14307 (2013).
Baret JC, Beck Y, Billas-Massobrio I, Moras D & Griffiths AD Quantitative cell-based reporter gene assays using droplet-based microfluidics. Chem. Biol. 17, 528–536 (2010).
Baret JC A remote syringe for cells, beads and particle injection in microfluidic channels, <http://blogs.rsc.org/chipsandtips/2009/08/20/a-remote-syringe-for-cells-beads-and-particle-injection-in-microfluidic-channels/> (2009). Accessed 27 March 2014.
Cooper R & Lee L Preventing suspension settling during injection, <http://blogs.rsc.org/chipsandtips/2007/08/21/preventing-suspension-settling-during-injection/> (2007). Accessed 27 March 2014.
Edd JF, Di Carlo D, Humphry KJ, Koster S, Irimia D, Weitz DA & Toner M . Controlled encapsulation of single-cells into monodisperse picolitre drops. Lab Chip 8, 1262–1264 (2008).
Kumaresan P, Yang CJ, Cronier SA, Blazej RG & Mathies RA High-throughput single copy DNA amplification and cell analysis in engineered nanoliter droplets. Anal. Chem. 80, 3522–3529 (2008).
Link DR, Grasland-Mongrain E, Duri A, Sarrazin F, Cheng Z, Cristobal G, Marquez M & Weitz DA Electric control of droplets in microfluidic devices. Angew. Chem. Int. Ed. 45, 2556–2560 (2006).
Ahn K, Kerbage C, Hunt TP, Westervelt RM, Link DR & Weitz DA Dielectrophoretic manipulation of drops for high-speed microfluidic sorting devices. Appl. Phys. Lett. 88, 024104 (2006).
Baroud CN, Delville J-P, Gallaire F & Wunenburger R Thermocapillary valve for droplet production and sorting. Phys. Rev. E. Stat. Nonlin. Soft. Matter Phys. 75, 046302 (2007).
Kemna EWM, Segerink LI, Wolbers F, Vermes I & van den Berg A Label-free, high-throughput, electrical detection of cells in droplets. Analyst. 138, 4585–4592 (2013).
Niu X, Zhang M, Peng S, Wen W & Sheng P Real-time detection, control, and sorting of microfluidic droplets. Biomicrofluidics 1, 044101 (2007).
Hu X, Bessette PH, Qian J, Meinhart CD, Daugherty Patrick S & Soh HT . Marker-specific sorting of rare cells using dielectrophoresis. Proc. Natl Acad. Sci. USA 102, 15757–15761 (2005).
Chen Y, Hsu C & Chen C Manipulation of microdroplets by using dielectrophoresis. IEEE EMBC Short Papers No. 3207 (2013).
Boitard L, Cottinet D, Kleinschmitt C, Bremond N, Baudry J, Yvert G & Bibette J Monitoring single-cell bioenergetics via the coarsening of emulsion droplets. Proc. Natl Acad. Sci. USA 109, 7181–7186 (2012).
Clausell-Tormos J, Lieber D, Baret JC, El-Harrak A, Miller OJ, Frenz L, Blouwolff J, Humphry KJ, Koster S, Duan H, Holtze C, Weitz DA, Griffiths AD & Merten CA . Droplet-based microfluidic platforms for the encapsulation and screening of Mammalian cells and multicellular organisms. Chem. Biol. 15, 427–437 (2008).
Chokkalingam V, Tel J, Wimmers F, Liu X, Semenov S, Thiele J, Figdor CG & Huck WTS Probing cellular heterogeneity in cytokine-secreting immune cells using droplet-based microfluidics. Lab Chip 13, 4740–4744 (2013).
Brouzes E, Medkova M, Savenelli N, Marran D, Twardowski M, Hutchison JB, Rothberg JM, Link DR, Perrimon N & Samuels ML Droplet microfluidic technology for single-cell high-throughput screening. Proc. Natl Acad. Sci. USA 106, 14195–14200 (2009).
Martinez CJ, Kim JW, Ye CW, Ortiz I, Rowat AC, Marquez M & Weitz D A microfluidic approach to encapsulate living cells in uniform alginate hydrogel microparticles. Macromol. Biosci. 12, 946–951 (2012).
Boedicker JQ, Li L, Kline TR & Ismagilov RF Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8, 1265–1272 (2008).
Pan J, Stephenson AL, Kazamia E, Huck WTS, Dennis JS, Smith AG & Abell C Quantitative tracking of the growth of individual algal cells in microdroplet compartments. Integr. Biol. 3, 1043–1051 (2011).
Jakiela S, Kaminski TS, Cybulski O, Weibel DB & Garstecki P Bacterial Growth and Adaptation in Microdroplet Chemostats. Angew. Chem. Int. Ed. 52, 8908–8911 (2013).
Hufnagel H, Huebner A, Gulch C, Guse K, Abell C & Hollfelder F An integrated cell culture lab on a chip: modular microdevices for cultivation of mammalian cells and delivery into microfluidic microdroplets. Lab Chip 9, 1576–1582 (2009).
Tan WH & Takeuchi S Monodisperse alginate hydrogel microbeads for cell encapsulation. Adv. Mater. 19, 2696–2701 (2007).
Kumachev A, Greener J, Tumarkin E, Eiser E, Zandstra PW & Kumacheva E High-throughput generation of hydrogel microbeads with varying elasticity for cell encapsulation. Biomaterials 32, 1477–1483 (2011).
Tsuda Y, Morimoto Y & Takeuchi S Monodisperse cell-encapsulating peptide microgel beads for 3D cell culture. Langmuir 26, 2645–2649 (2009).
Li CY, Wood DK, Hsu CM & Bhatia SN DNA-templated assembly of droplet-derived PEG microtissues. Lab Chip 11, 2967–2975 (2011).
Rossow T, Heyman JA, Ehrlicher AJ, Langhoff A, Weitz DA, Haag R & Seiffert S Controlled synthesis of cell-laden microgels by radical-free gelation in droplet microfluidics. J. Am. Chem. Soc. 134, 4983–4989 (2012).
Ma S, Natoli M, Liu X, Neubauer MP, Watt FM, Fery A & Huck WTS Monodisperse collagen-gelatin beads as potential platforms for 3D cell culturing. J. Mater. Chem. B 1, 5128–5136 (2013).
Chan HF, Zhang Y, Ho Y-P, Chiu Y-L, Jung Y & Leong KW Rapid formation of multicellular spheroids in double-emulsion droplets with controllable microenvironment. Sci. Rep. 3, 3462 (2013).
Chokkalingam V, Ma Y, Thiele J, Tel J, Schalk W & Huck WTS An electro-coalescence chip for effective emulsion breaking in droplet microfluidics. Lab Chip 14, 2398–2402 (2014).
Tumarkin E, Tzadu L, Csaszar E, Seo M, Zhang H, Lee A, Peerani R, Purpura K, Zandstra PW & Kumacheva E High-throughput combinatorial cell co-culture using microfluidics. Integr. Biol. 3, 653–662 (2011).
Brown RB & Audet J Current techniques for single-cell lysis. J. R. Soc. Interface 5, S131–S138 (2008).
He M, Edgar JS, Jeffries GDM, Lorenz RM, Shelby JP & Chiu DT Selective encapsulation of single cells and subcellular organelles into picoliter- and femtoliter-volume droplets. Anal. Chem. 77, 1539–1544 (2005).
Rau KR, Guerra A, Vogel A & Venugopalan V Investigation of laser-induced cell lysis using time-resolved imaging. Appl. Phys. Lett. 84, 2940–2942 (2004).
Kintses B, Hein C, Mohamed MF, Fischlechner M, Courtois F, Leine C & Hollfelder F Picoliter cell lysate assays in microfluidic droplet compartments for directed enzyme evolution. Chem. Biol. 19, 1001–1009 (2012).
Najah M, Griffiths AD & Ryckelynck M Teaching single-cell digital analysis using droplet-based microfluidics. Anal. Chem. 84, 1202–1209 (2012).
Kim T & Eberwine J Mammalian cell transfection: the present and the future. Anal. Bioanal. Chem. 397, 3173–3178 (2010).
Zhan Y, Wang J, Bao N & Lu C Electroporation of cells in microfluidic droplets. Anal. Chem. 81, 2027–2031 (2009).
Chen F, Zhan Y, Geng T, Lian H, Xu P & Lu C Chemical transfection of cells in picoliter aqueous droplets in fluorocarbon oil. Anal. Chem. 83, 8816–8820 (2011).
Eun YJ, Utada AS, Copeland MF, Takeuchi S & Weibel DB Encapsulating bacteria in agarose microparticles using microfluidics for high-throughput cell analysis and isolation. ACS Chem. Biol. 6, 260–266 (2011).
Joensson HN, Samuels ML, Brouzes ER, Medkova M, Uhlen M, Link DR & Andersson-Svahn H Detection and analysis of low-abundance cell-surface biomarkers using enzymatic amplification in microfluidic droplets. Angew. Chem. Int. Ed. 48, 2518–2521 (2009).
Hashimshony T, Wagner F, Sher N & Yanai I CEL-Seq: single-cell RNA-seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012).
Guo MT, Rotem A, Heyman JA & Weitz DA Droplet microfluidics for high-throughput biological assays. Lab Chip 12, 2146–2155 (2012).
Plessy C, Desbois L, Fujii T & Carninci P Population transcriptomics with single-cell resolution: A new field made possible by microfluidics. Bioessays 35, 131–140 (2013).
Miller MC, Doyle GV & Terstappen LW Significance of circulating tumor cells detected by the cellsearch system in patients with metastatic breast colorectal and prostate cancer. J. Oncol. 2010, 617421 (2010).
Stahlberg A, Kubista M & Aman P Single-cell gene-expression profiling and its potential diagnostic applications. Expert Rev. Mol. Diagn. 11, 735–740 (2011).
Zeng Y, Novak R, Shuga J, Smith MT & Mathies RA High-performance single cell genetic analysis using microfluidic emulsion generator arrays. Anal. Chem. 82, 3183–3190 (2010).
Mary P, Dauphinot L, Bois N, Potier MC, Studer V & Tabeling P Analysis of gene expression at the single-cell level using microdroplet-based microfluidic technology. Biomicrofluidics 5, 24109 (2011).
Zhang HF, Jenkins G, Zou Y, Zhu Z & Yang CJ Massively parallel single-molecule and single-cell emulsion reverse transcription polymerase chain reaction using agarose droplet microfluidics. Anal. Chem. 84, 3599–3606 (2012).
Geng T, Novak R & Mathies RA Single-cell forensic short tandem repeat typing within microfluidic droplets. Anal. Chem. 86, 703–712 (2013).
Konry T, Dominguez-Villar M, Baecher-Allan C, Hafler DA & Yarmush ML Droplet-based microfluidic platforms for single T cell secretion analysis of IL-10 cytokine. Biosens. Bioelectron. 26, 2707–2710 (2011).
Zinchenko A, Devenish SRA, Kintses B, Colin P-Y, Fischlechner M & Hollfelder F One in a million: flow cytometric sorting of single cell-lysate assays in monodisperse picolitre double emulsion droplets for directed evolution. Anal. Chem. 86, 2526–2533 (2014).
Juul S, Ho YP, Koch J, Andersen FF, Stougaard M, Leong KW & Knudsen BR Detection of single enzymatic events in rare or single cells using microfluidics. ACS Nano. 5, 8305–8310 (2011).
Agresti JJ, Antipov E, Abate AR, Ahn K, Rowat AC, Baret JC, Marquez M, Klibanov AM, Griffiths AD & Weitz DA Ultrahigh-throughput screening in drop-based microfluidics for directed evolution. Proc. Natl Acad. Sci. USA 107, 4004–4009 (2010).
Griffiths AD, Williams SC, Hartley O, Tomlinson IM, Waterhouse P, Crosby WL, Kontermann RE, Jones PT, Low NM & Allison TJ Isolation of high affinity human antibodies directly from large synthetic repertoires. EMBO J. 13, 3245–3260 (1994).
Tawfik DS & Griffiths AD Man-made cell-like compartments for molecular evolution. Nat. Biotechnol. 16, 652–656 (1998).
Sjostrom SL, Bai Y, Huang M, Liu Z, Nielsen J, Joensson HN & Andersson Svahn H High-throughput screening for industrial enzyme production hosts by droplet microfluidics. Lab Chip 14, 806–813 (2014).
Acknowledgements
Work in the Huck group is supported by a European Research Council (ERC) Advanced Grant (246812 Intercom), a VICI grant and a VENI grant (86313024 to JTel) of the Netherlands Organization for Scientific Research (NWO) and by funding from the Ministry of Education, Culture and Science (Gravity program 024.001.035).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Rakszewska, A., Tel, J., Chokkalingam, V. et al. One drop at a time: toward droplet microfluidics as a versatile tool for single-cell analysis. NPG Asia Mater 6, e133 (2014). https://doi.org/10.1038/am.2014.86
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/am.2014.86
This article is cited by
-
Regulating the Oil-Water Interface to Construct Double Emulsions: Current Understanding and Their Biomedical Applications
Chemical Research in Chinese Universities (2022)
-
Microvalves for integrated selective droplet generation, splitting and merging on a chip
Microfluidics and Nanofluidics (2021)
-
Simulation studies on picolitre volume droplets generation and trapping in T-junction microchannels
SN Applied Sciences (2020)
-
Dendronized fluorosurfactant for highly stable water-in-fluorinated oil emulsions with minimal inter-droplet transfer of small molecules
Nature Communications (2019)
-
Drop formation in microfluidic cross-junction: jetting to dripping to jetting transition
Microfluidics and Nanofluidics (2019)