ABSTRACT
Genetic polymorphisms in human genes can influence the risk for HIV-1 infection and disease progression, although the reported effects of these alleles have been inconsistent. This review highlights the recent discoveries on global and Chinese genetic polymorphisms and their association with HIV-1 transmission and disease progression.
Similar content being viewed by others
INTRODUCTION
HIV-1 infection results in a variety of clinical outcomes. The majority of HIV-1 infected individuals progress to AIDS within 5 to 10 years, some progress rapidly to AIDS (termed rapid progressors) while others progress to AIDS slowly (slow progressors). A small number of HIV-1 infected individuals, termed long-term non progressors (LTNP), remain clinically healthy for more than a decade after infection 1, 2, 3, 4. There are also some individuals who remain seronegative despite high risk and/or multiple exposures to HIV-1. These exposed seronegatives (ES) include infants born to HIV-1 infected mothers 5, 6, 7, commercial sex workers in epidemic areas 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, hemophiliacs who received HIV contaminated factor VIII preparations 26, 27, and sexual partners of known HIV-1 infected persons 28, 29, 30, 31, 32, 33. Understanding the mechanisms that account for slower disease progression in LTNP and the protection against HIV-1 infection observed with ES is important for the development of more potent therapeutic regimens and a vaccine. It is likely that both viral and host factors may contribute to these outcomes. Acquired immune responses, including cellular 5, 8, 9, 10, 11, 14, 15, 17, 26, 28, 34, 35, 36, 37 and humoral 10, 16, 20, 38, 39, 40, 41, 42, 43, 44 responses to HIV-1, may play an important role in protecting against and controlling HIV-1 infection. Significant studies in the past few years have also demonstrated that innate immunity including genetic polymorphisms in host genes can affect the risk for HIV-1 infection and disease progression (Tab. 1), although the effect of these alleles has been inconsistent 32, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55 (reviewed in 56). Here, we review global and Chinese studies on human genetic polymorphisms and their association with HIV-1 infection.
VARIATION IN THE CCR5 CODING REGION
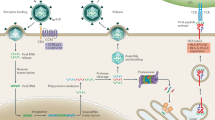
HIV-1 requires CD4 as its primary receptor and a chemokine receptor as a co-receptor to enter cells 57, 58, 59. Based on the co-receptor utilization, HIV-1 strains can be classified as “R5 tropic HIV-1” that primarily utilize C-C chemokine receptor 5 (CCR5) and “X4 tropic HIV-1” that use C-X-C chemokine receptor 4 (CXCR4) 60. R5 tropic viruses dominate in HIV-1 transmission from person to person, while X4 HIV-1 strains are frequently found during the later stages of HIV-1 infection 61. CCR5-Δ32, an allele of CCR5 that contains a 32 bp deletion, codes for a nonfunctional co-receptor and cell lines homozygous for CCR5-Δ32 are resistant to R5-virus but individuals that are CCR5-Δ32 homozygotes (people who inherited the CCR5-Δ32 from both parents) are resistant to HIV-1 infection, indicating that genotype CCR5-Δ32 is highly protective against HIV-1 infection 45, 46, 47, 48, 49, 50, 54. However, this protection is not absolute because rare individuals homozygous for CCR5-Δ32 are infected with HIV-1 strains that may utilize another co-receptor, such as CXCR4 64, 65, 66, 67, 68. Furthermore, homozygous CCR5- Δ32 is found only in 1% of the general population of Caucasians, but not in Africans, Asians or other ethnic groups 45, 46, 47, 48, 49, 50, 54, and the majority of highly exposed yet uninfected individuals have two normal CCR5 alleles (called wild-type, or wt). Similarly, we found CCR5-Δ32 homozygotes in 1.0% (7 out of 705) of HIV-1 seronegative individuals and 3.1% (3 out of 97) of ES.
CCR5-Δ32 heterozygotes (people who inherited the CCR5-Δ32 allele from one parent and a functional CCR5 allele from the other parent) are susceptible to HIV-1 infection; most studies have not supported an association of the heterozygous CCR5-Δ32 genotype with reduced HIV-1 transmission risk for adult and pediatric populations 47, 48, 49, 69, 70, 71, 72, 73. However, our recent study suggests that individuals with the combination of heterozygous CCR5-Δ32 and P1 CCR5 promoter genotype (see below) are relatively resistant to HIV-1 transmission 74. According to most reports 47, 48, 49, 55, 75, 76, 77, 78, 79, 80, 81 but not all 82, 83, individuals with heterozygous CCR5-Δ32 progress from HIV-1 infection to AIDS more slowly than persons with two normal CCR5 alleles.
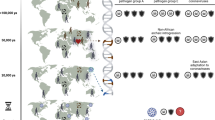
CCR5-Δ32 homozygotes make up approximately 1-3% of northern European populations; CCR5-Δ32 heterozygotes and wt individuals comprise approximately 14% and 83%, respectively, of the remainder 46, 84. The polymorphism demonstrates a decreasing north-south cline across Eurasia and is largely absent in African, Asian and Oceanic populations 84, 85, 86, interestingly this distribution may have become fixed 700 years ago in northwestern Europe 86. Indeed, studies to date indicate that CCR5-Δ32 mutant alleles were absent or infrequent in Chinese 87, 88, 89, 90, 91, 92, 93. Homozygous genotypes have not been identified in Chinese populations, while heterozygous CCR5-Δ32 is extremely rare or absent in most Chinese populations studied 87, 88, 89, 90, 91, 92, 93. There is no evidence that CCR5-Δ32 influences the HIV-1 transmission or epidemic in China.
Wang et al in collaboration with us 87 conducted a large scale investigation on human genetic polymorphisms in three cohorts of Chinese: 1) 3165 indigenous healthy subjects representing eight ethnic groups: Han (n = 1406), Uygur (n = 316), Mongolia (n = 134), Hui (n = 386), Tibetan (n = 330), Zhuang (n = 378), Dai (n = 101), and Jingbo (n =114); 2) 330 HIV-1 infected (86 subjects infected by sexual transmission and 198 subjects infected by HIV-1 contaminated blood or by sharing injection equipment; the remaining 46 subjects said nothing about HIV-1 transmission); and 3) 474 HIV-1 uninfected Han Chinese belonging to one of two HIV-1 high-risk groups: intravenous drug users (n = 215) and individuals with sexually transmitted diseases (n = 259). Heterozygous CCR5-Δ32 genotypes were found in 3 out of 1254 Han Chinese, with an allele frequency of 0.00119. Findings from this study and others 88, 89 show that CCR5-Δ32 mutants do occur in Chinese population (all are individuals with heterozygous CCR5-Δ32), and can be inherited at a very low frequency 87, 88, 89.
VARIATION IN CCR5 PROMOTER REGION
Polymorphisms in the CCR5 promoter are associated with altered disease progression but not reduced transmission risk of HIV-1 infection 51, 94. Of the ten alleles in the CCR5 promoter (CCR5P1 to CCR5P10), only homozygote CCR5P1 is associated with an accelerated progression (by approximately 4 years) to AIDS 51. This acceleration of disease progression was most marked in the first 5 years of infection. McDermott et al reported that a point mutation, 59029A/G, which is linked to the CCR5P1 allele, accelerated progression to AIDS by 3.8 years 94. However, there is no association between the CCR5P1 or 59029A promoter genotypes to either increased in vivo expression of CCR5 mRNA or cell surface proteins51, 94. Thus, the mechanisms of action for the CCR5P1 and 59029A promoter remain to be defined.
There is currently little data on the polymorphisms in the CCR5 promoter in Chinese populations. A study on 96 HIV negative Chinese individuals in Taiwan indicates that only CCR5P1 and P4 haplotypes were detected, and the P1/P1, P1/P4 and P4/P4 genotype frequencies were 21.0%, 41.1% and 37.9%, respectively 95. The sequencing data confirmed the results of previous studies, showing that CCR5P1 exhibited complete linkage disequilibrium with a polymorphic allele 59029A present in the CCR5 promoter. Furthermore, fluorescence-activated cell sorter analysis revealed that, in the absence of the CCR2-64I mutation, individuals carrying CCR5P1 tended to express more surface CCR5 on monocytes and CD4+ cells. However, the association of the CCR5P1 with HIV-1 infection in Chinese populations is currently unknown.
VARIATION IN THE CCR2 CODING REGION
CCR2 is a minor HIV-1 coreceptor. The gene that codes for this chemokine receptor has a variant allele causing a Val-Ile switch at amino acid position 64 (CCR2-64I) in the first transmembrane domain of CCR2. Unlike the CCR5-Δ32 allele that inactivated the major HIV-1 co-receptor, CCR2-64I causes a conservative change in a nonexposed portion of a coreceptor of questionable physiologic relevance 68, 96, 97. CCR2-64I is common and found in 10% of Caucasians, 15% of African-Americans, 25% of Asians and 17% of Hispanics 50. Epidemiologic studies by Smith et al first demonstrated the association of CCR2-64I with delayed HIV-1 disease progression 50, which was confirmed by most subsequent studies 73, 98, 99, but not by others 82, 100, 101. CCR2-64I is not associated with reduced risk for HIV-1 infection 50. The mechanism for the association of CCR2-64I with delayed disease progression has not been elucidated despite significant attempts to do so 98, 102, 103.
The frequencies of heterozygous and homozygous CCR2-64I in Chinese were 13-35% and 1-8%, respectively, varying from study to study 87, 88, 89, 90, 91, 92, 93. There is no difference in the frequencies of both heterozygous and homozygous CCR2-64I between HIV-1 infected and HIV seronegative individuals, suggesting no association of CCR2-64I with HIV-1 transmission in Chinese. There is also generally no evidence of association of CCR2-64I with disease progression in HIV-1 infected Chinese individuals 87, 88, 89, 90, 91, 92, 93. The allelic frequency of CCR2-64I was about 20% (95% CI, 15∼30%) 87, 88, 89, 90, 91, 92, 93, which is significantly higher than that in other ethnic groups including Caucasians. Furthermore the polymorphisms of CCR2-64I in the Han Chinese population were different from those in American Caucasians 87.
CCR2-64I was common in different ethnic groups in Chinese populations 87 (f range: 16.23%–28.79%), with the lowest frequency observed in the Jingbo (f=16.23%). The Dai, Hui, and Uygur ethnic groups had CCR2-64I allele frequencies similar to that of the Han group (f range: 19.15%∼21.76%), but the Mongolia and Zhuang groups tended to have a higher frequency (f range: 23.41%∼24.63%), with the highest frequency observed in the Tibetan population (f=28.79%). However, there was no significant differences among CCR2-64I allele and genotype frequencies when the HIV-1 infected group was compared with either the HIV-1 uninfected STD, IDU, or combined at-risk group (dominant model: OR = 0.99-1.08, p>0.05; recessive model: OR = 1.22-1.41, p>0.05, respectively).
VARIATION IN SDF1
SDF1 (Stromal-Derived Factor 1, also called CXCL12) is the primary ligand for the late-stage HIV-1 receptor CXCR4. Winkler et al reported a Gly-Ala transition at position 801 (with position 1 as the A of the initiation codon) of the mRNA for SDF-1b, one of the two isoforms of the CXCR4 chemokine ligand, and demonstrated that homozygosity for this SDF1-3′A allele was associated with delayed disease progression in an analysis of 639 seroincident subjects from four HIV cohorts 104. Although this observation was confirmed by the French GRIV cohort at the limit of statistical significance (probability=0.05), other studies demonstrated contrastly that the SDF1-3′A homozygous genotype was associated with accelerated disease progression 73, 105. Ultimately, an international meta-analysis of 19 prospective cohort studies and case-control studies from Europe and Australia demonstrated that SDF1-3′A homozygotes have no decreased risk for AIDS, or death after development of AIDS 55.
The SDF1-3′A allele was also common in different Chinese ethnic groups, with the highest frequencies observed in Chinese Han, Zhuang, and Hui (f=27.76%, 25.93%, and 24.87%, respectively) 87. The SDF1-3′A allele frequency was significantly lower (p<0.05) among the Dai, Jingbo, Mongolia, Uygur, and Tibetan populations (20.30%, 17.70%, 19.10%, 20.02%, and 20.41%, respectively). There was a slight decrease in SDF1-3′A homozygotes and heterozygotes in the HIV-1 infected group compared with the healthy HIV-1 negative Han group (f=25.61%, 27.76%; p=0.038, respectively). After correction for multiple comparisons, this difference was not significant (dominant model: OR=0.84–0.95, p>0.05; recessive model: OR=1.28–1.30, p>0.05, respectively).
The above results are in general agreement with other studies in Chinese populations 88, 89, 90, 91, 92, 93. For example, we observed a similar frequency of SDF1-3′A in Shenzhen (26.9%, corresponding to 17.6∼38.2% of 95% CI) with a mixture of ethnic Chinese groups. We found a weak association of SDF1-3′A with low viral load, and no association with disease progression after HIV-1 infection 106.
VARIATION IN RANTES
RANTES (regulated on activation normal T cell expressed and secreted) is one of the natural ligands for the chemokine receptor CCR5 and potently suppresses in vitro replication of the R5 strains of HIV-1 107, which use CCR5 as a coreceptor. Two single nucleotide polymorphisms (SNP), -403G/A and -28C/G, in the promoter region of RANTES were initially identified by Liu et al in Japan 52. The -403A-28G haplotype was shown to be associated with delayed disease progression in HIV-1 infected Japanese, but exerts no influence on the incidence of HIV-1 infection 52. In European-Americans, the compound genotype -403G/A -28C/C was reported to be resistant to AIDS progression in one study 53, but not in another 54. These RANTES polymorphisms have no effect on HIV-1 infection and disease progression in African-Americans 54. Most recently, An et al have found that 3 SNPs (-403A in the promoter, In1.1C in the first intron, and 3′222C in the 3′ untranslated region) are associated with increased frequency of HIV-1 infection, and that the In1.1C allele or haplotypes display a strong association with rapid progression to AIDS among HIV-1 infected African-Americans and European-Americans 108. These and other RANTES SNPs may also influence the varied epidemiology of HIV-1 infection throughout the world 54, 108.
There is relatively little information describing variation in the RANTES gene and the association with HIV-1 infection in Chinese populations 109, 110. Liu et al 109 identified 6 genotypes of RANTES promoter -403 and -28 in the Han Chinese group. RANTES genotypes AC/AG, AC/GC, AG/GC, GC/GC were associated with reduced susceptibility to HIV-1 infection. However, there was no significant difference in the allele frequencies between people living with HIV-1 and HIV negative individuals. There were significant differences of RANTES In1.1C between HIV-1 infected and healthy individuals in males, suggesting that the In1.1C-bearing genotypes could increase susceptibility to HIV-1 infection. No such significance was found in females. A study by Zhao et al of 1082 Chinese blood donors from northern and southern China and 249 HIV patients from southern China indicated that Chinese AIDS patients, compared to seronegative adults, had a significantly higher frequency of the -403G allele and haplotype I, -403G/-28C (p<0.05), and a lower frequency of the -403A/A genotype (p<0.01). Symptomatic patients had a higher frequency of the -28G allele and a lower frequency of the -28C/C genotype (p<0.01). These results suggest that -403G may be associated with increased susceptibility to HIV infection, while -28G may be associated with advanced disease progression. The impact of these SNPs on HIV infection appears to be unique in Chinese, while a large scale study would be warranted to verify these findings.
VARIATION IN HLA
The HLA (human leukocyte antigen) region includes 128 expressed genes, of which, about 43 genes are associated with human immunity 111. HLA class I (A, B and C) and II genes (IDR, DQ and DP) have considerable allele variation between individuals and populations, which provides a broad range for individual recognition of viral agents to which they have been exposed in the past, as well as those to which they have not 112. Because different HLA alleles specify cell-surface molecules with specific motif recognition sites for infectious agents 113, differential HIV-1 peptide motif recognition can influence both the time interval from infection to AIDS 114 and the kinetics of HIV-1 adaptive escape from immune surveillance in an infected individual 115. For example, Carrington et al demonstrated the association of heterozygosity of HLA alleles B35 and Cw4 with accelerated disease progression 116. Two HLA alleles, HLA-B27 and HLA-B57 are associated with a delayed progression to AIDS 117, 118. Activating the killer immunoglobulin-like receptors (KIRs) allele KIR3DS1, in combination with HLA alleles (HLA-Bw4) that encode molecules with isoleucine at position 80 (HLA-B Bw4-80Ile), was associated with delayed progression to AIDS 119.
Xu et al 120 determined the distribution of HLA-B alleles in 106 healthy HIV negative and 73 HIV positive Chinese Yi ethnic individuals and its association with HIV infection. The frequency of alleles B*07, B*35, and B*46 were increased in HIV-1 positive subjects, whereas the alleles B*55, B*44 and B*78 were absent in the HIV infected persons studied. The B*46 allele was present in a significantly higher gene frequency among HIV-1 positive individuals (P=0.02, OR=3.32, 95% CI=1.13-9.78) compared with control subjects, suggesting that HLA-B*46 may be associated with its increased susceptibility to HIV-1 infections.
VARIATION in DC-SIGN AND DC-SIGNR CODING and promoter REGION
Because dendritic cells (DCs) are among the first cells encountered by HIV-1 during sexual transmission and DCs migrate from mucosal sites to the secondary lymphoid organs upon capturing antigen 121, 122, it has been proposed that HIV-1 uses DCs as carriers to gain entry into lymph nodes and subsequently infect CD4+ T cells 123. DC-SIGN (Dendritic cell-specific intercellular adhesion molecule-3-grabbing nonintegrin) on DCs, originally described as a C-type (calcium-dependent) lectin, is able to capture HIV-1, HIV-2 and SIV 124, and retains the attached virus in an infectious state for days and then transmits the virus to CD4 and co-receptor positive cells 123. DC-SIGNR (DC-SIGN related) shows similar functions to DC-SIGN for capturing HIV-1 and enhancing HIV-1 infection of T cells 125, 126. Studies including ours have shown that mRNA encoding DC-SIGN and DC-SIGNR is present in DCs 127, 128, 129, though the DC-SIGNR transcripts are largely alternatively spliced isoforms 127.
Both DC-SIGN and DC-SIGNR, clustered on chromosome 19 125, 128, are organized into three domains: an N-terminal cytoplasmic region, a neck region containing seven repeats of a 23 amino acid sequence, and a C-terminal domain with homology to C-type lectins 128. We assessed whether polymorphisms in the DC-SIGN and DC-SIGNR repeat region could affect individual HIV-1 susceptibility and subsequent disease progression by analyzing DC-SIGN and DC-SIGNR repeat polymorphisms in diverse cohorts of ES, HIV-1 seropositive (HIV-1+), long-term non-progressors (LTNP), and HIV-1 seronegative (HIV-1-) individuals. We identified novel variants in DC-SIGN repeat region and observed that heterozygous (7/6 and 7/8) DC-SIGN reduced the risk of HIV-1 infection (3.2% in ES, 0.0% in HIV-1+, P = 0.011) 32. Of the 835 individuals we tested, all 8 individuals with DC-SIGN repeat region variations were from HIV-1- individuals, of whom 3 were in ES. Compared with HIV-1+ individuals, a higher prevalence of DC-SIGN variations in the repeat region was observed among ES individuals, suggesting an association of DC-SIGN variation with resistance to HIV-1 infection in ES 32. We further assessed polymorphisms in the DC-SIGNR repeat region in diverse cohorts of multiply exposed seronegative or high-risk seronegative, HIV-1 infected and HIV-1 seronegative individuals from Seattle and the Multicenter AIDS Cohort Study (MACS) cohorts 130. Our results suggest that individuals with a 7/7 genotype in the DC-SIGNR repeat region are associated with an increased risk for HIV-1 infection (P=0.0015). However, these effects were much stronger in the Seattle cohort (P=0.0014) than in the MACS cohort (P=0.1890). Individuals with a 7/5 genotype in the DC-SIGNR repeat region are more frequently in the multiply exposed seronegative or high-risk seronegative cohorts in the Seattle-MACS combined cohort (P=0.029) or in the Seattle cohort only (P=0.027). Most recently, we identified the “resistant” variants of DC-SIGN and DC-SIGNR heterozygous 7/5 in Chinese populations, and are further determining their association with HIV-1 infection in China (Zhu et al, personnel communication).
Matin et al examined 1,611 European-American individuals at risk for parental (n=713) or mucosal (n=898) infection for genetic polymorphisms in DC-SIGN promoter region 131. It was found that individuals at risk for parentally acquired infection who had -336C were more susceptible to infection than were persons with -336T (odd ratio=1.87, p= 0.001). However, this association was not observed in those at risk for mucosally acquired infection.
CONCLUSION
Results from us and others indicate that homozygous CCR5-Δ32, the combination of heterozygous CCR5-Δ32 and CCR5-59029A, and the DC-SIGN and DC-SIGNR repeat polymorphisms affect HIV-1 transmission (Tab. 1). However, homozygous CCR5-Δ32 has not been identified and heterozygous CCR5-Δ32 is extremely rare in Chinese populations, indicating no or little effect of CCR5-Δ32 on HIV-1 transmission and epidemic in China. Genetic polymorphisms that have been shown to influence HIV-1 transmission are relatively rare and only account for the resistance of a small proportion of ES individuals to infection 32, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, underscoring the need for more research.
There are relatively more genetic polymorphisms in human genes that may alter disease progression of HIV-1 infection (Tab. 1). However, the associations of genetic variants with HIV-1 disease progression in Chinese have not been well established. Only RANTES -403A and -28G, and SDF1-3′A alleles have been shown to have a weak influence on the disease progression in HIV-1 infected Chinese.
The relatively negative results from studies to date on Chinese populations may not necessarily indicate that the genetic polymorphisms have little effect on HIV-1 infection in Chinese populations. Instead, more efforts should be made to establish good study cohorts including longitudinally followed “resistant” ES, LTNP and primary HIV-1 infection. Further investigation of the association of these polymorphisms with HIV-1 infection with well established study cohorts, and the identification of new polymorphisms that may influence infection and disease progression is warranted. In addition, some less common polymorphisms identified such as in CCR5 132, 133, 134, 135, 136 and CXCR4 137, 138 might be included in large scale studies with well-established cohorts.
References
Buchbinder SP, Katz MH, Hessol NA, O'Malley PM, Holmberg SD . Long-term HIV-1 infection without immunologic progression. AIDS 1994; 8:1123–8.
Cao Y, Qin L, Zhang L, Safrit J, Ho DD . Virologic, immunologic characterization of long-term survivors of HIV-1 infection. N Engl J Med 1995; 332:201–8.
Pantaleo G, Menzo S, Vaccarezza M, et al. Studies in subjects with long-term nonprogressive HIV infection. New Eng J Med 1995; 332:209–16.
Deacon NJ, Tsykin A, Solomon A, et al. Genomic structure of an attenuated quasi species of HIV-1 from a blood transfusion donor and recipients. Science 1995; 270:988–91.
Rowland-Jones SL, Nixon DF, Aldhous MC, et al. HIV-specific cytotoxic T-cell activity in an HIV-exposed but uninfected infant. Lancet 1993; 341:860–861.
De Maria A, Cirillo C, Moretta L . Occurrence of human immunodeficiency virus type 1 (HIV-1)-specific cytolytic T cell activity in apparently uninfected children born to HIV-1-infected mothers. J Infect Dis 1994; 170:1296–9.
Mayaux MJ, Blanche S, Rouzioux C, et al. Maternal factors associated with perinatal HIV-1 transmission: the French Cohort Study: 7 years of follow-up observation. J Acquir Immune Defic Syndr Hum Retrovirol 1995; 8:188–94.
Langlade-Demoyen P, Ngo-Giang-Huong N, Ferchal F, Oksenhendler E . Human immunodeficiency virus (HIV) nef-specific cytotoxic T lymphocytes in noninfected heterosexual contact of HIV-infected patients. J Clin Invest 1994; 93:1293–7.
Rowland-Jones SL, Sutton J, Ariyoshi K, et al. HIV-specific cytotoxic T-cells in HIV-exposed but uninfected Gambian women. Nat Med 1995; 1:59–64.
Mazzoli S, Trabattoni D, Lo Caputo S, et al. HIV-specific mucosal and cellular immunity in HIV-seronegative partners of HIV-seropositive individuals. Nat Med 1997; 3:1250–7.
Rowland-Jones SL, Dong T, Fowke KR, et al. Cytotoxic T cell responses to multipleconserved HIV epitopes in HIV- resistant prostitutes in Nairobi. J Clin Invest 1998; 102:1758–65.
Rowland-Jones SL, Dong T, Dorrell L, et al. Broadly cross-reactive HIV-specific cytotoxic T-lymphocytes in highly- exposed persistently seronegative donors. Immunol Lett 1999; 66:9–14.
Kaul R, Trabattoni D, Bwayo JJ, et al. HIV-1-specific mucosal IgA in a cohort of HIV-1-resistant Kenyan sex workers. Aids 1999; 13:23–9.
Kaul R, Plummer FA, Kimani J, et al. HIV-1-specific mucosal CD8+ lymphocyte responses in the cervix of HIV-1- resistant prostitutes in Nairobi. J Immunol 2000; 164:1602–11.
Kaul R, Rowland-Jones SL, Kimani J, et al. Late seroconversion in HIV-resistant Nairobi prostitutes despite pre- existing HIV-specific CD8+ responses. J Clin Invest 2001; 107:341–9.
Kaul R, Plummer F, Clerici M, Bomsel M, Lopalco L, Broliden K . Mucosal IgA in exposed, uninfected subjects: evidence for a role in protection against HIV infection. Aids 2001; 15:431–2.
Kaul R, Dong T, Plummer FA, et al. CD8(+) lymphocytes respond to different HIV epitopes in seronegative and infected subjects. J Clin Invest 2001; 107:1303–10.
MacDonald KS, Fowke KR, Kimani J, et al. Influence of HLA supertypes on susceptibility and resistance to human immunodeficiency virus type 1 infection. J Infect Dis 2000; 181:1581–9.
Dorrell L, Hessell AJ, Wang M, et al. Absence of specific mucosal antibody responses in HIV-exposed uninfected sex workers from the Gambia. Aids 2000; 14:1117–22.
Devito C, Hinkula J, Kaul R, et al. Mucosal and plasma IgA from HIV-exposed seronegative individuals neutralize a primary HIV-1 isolate. Aids 2000; 14:1917–20.
Sriwanthana B, Hodge T, Mastro TD, et al. HIV-specific cytotoxic T lymphocytes, HLA-A11, and chemokine-related factors may act synergistically to determine HIV resistance in CCR5 delta32-negative female sex workers in Chiang Rai, northern Thailand. AIDS Res Hum Retroviruses 2001; 17:719–34.
Fowke KR, Kaul R, Rosenthal KL, et al. HIV-1-specific cellular immune responses among HIV-1-resistant sex workers. Immunol Cell Biol 2000; 78:586–95.
Rowland-Jones SL, McMichael A . Immune responses in HIV-exposed seronegatives: have they repelled the virus? Curr Opin Immunol 1995; 7:448–55.
Rowland-Jones SL, Pinheiro S, Kaul R, et al. How important is the 'quality' of the cytotoxic T lymphocyte (CTL) response in protection against HIV infection? Immunol Lett 2001; 79:15–20.
Plummer FA, Ball TB, Kimani J, Fowke KR . Resistance to HIV-1 infection among highly exposed sex workers in Nairobi: what mediates protection and why does it develop? Immunol Lett 1999; 66:27–34.
Gibbons J, Cory JM, Hewlett IK, Epstein JS, Eyster ME . Silent infections with human immunodeficiency virus type 1 are highly unlikely in multitransfused seronegative hemophiliacs. Blood 1990; 76:1624–1626.
Lederman MM, Jackson JB, Kroner BL, et al. Human immunodeficiency virus (HIV) type 1 infection status and in vitro susceptibility to HIV infection among high-risk HIV-1-seronegative hemophiliacs. J Infect Dis 1995; 172:228–31.
Goh WC, Markee J, Akridge RE, et al. Protection against human immunodeficiency virus type 1 infection in persons with repeated exposure: evidence for T cell immunity in the absence of inherited CCR5 coreceptor defects. J Infect Dis 1999; 179:548–57.
Akridge R, Hladik F, Markee J, et al. Cellular immunity and target cell susceptibility in persons with repeated HIV-1 exposure. Immunol Lett 1999; 66:15–9.
Schmechel SC, Russell N, Hladik F, et al. Immune defence against HIV-1 infection in HIV-1-exposed seronegative persons. Immunol Lett 2001; 79:21–7.
Hladik F, Desbien A, Lang J, et al. Most highly exposed seronegative men lack HIV-1-specific, IFN-{gamma}-secreting T cells. J Immunol 2003; 171:2671–83.
Liu H, Hwangbo Y, Holte S, et al. Analysis of genetic polymorphisms in CCR5, CCR2, stromal cell-derived factor-1, RANTES, and dendritic cell-specific intercellular adhesion molecule-3-grabbing nonintegrin in seronegative individuals repeatedly exposed to HIV-1. J Infect Dis 2004; 190:1055–8.
Imagawa DT, Lee MH, Wolinsky SM, et al. Human immunodeficiency virus type 1 infection in homosexual men who remain seronegative for prolonged periods. N Engl J Med 1989; 320:1458–62.
Pinto LA, Sullivan J, Berzofsky JA, et al. ENV-specific cytotoxic T lymphocyte responses in HIV seronegative health care workers occupationally exposed to HIV-contaminated body fluids. J Clin Invest 1995; 96:867–76.
Clerici M, Giorgi J, Chou C, et al. Cell-mediated immune response to human immunodeficiency virus (HIV) type 1 in seronegative homosexual men with recent sexual exposure to HIV-1. J Infect Dis 1992; 165:1012–9.
Bernard NF, Yannakis CM, Lee JS, Tsoukas CM . Human immunodeficiency virus (HIV)-specific cytotoxic T lymphocyte activity in HIV-exposed seronegative persons. J Infect Dis 1999; 179:538–47.
Makedonas G, Bruneau J, Lin H, Sekaly RP, Lamothe F, Bernard NF . HIV-specific CD8 T-cell activity in uninfected injection drug users is associated with maintenance of seronegativity. Aids 2002; 16:1595–602.
Mazzoli S, Lopalco L, Salvi A, et al. Human immunodeficiency virus (HIV)-specific IgA nd HIV neutralizing activity in the serum of exposed seronegative partners of HIV-seropositive persons. J Infect Dis 1999; 180:871–5.
Lopalco L, Pastori C, Cosma A, et al. Anti-cell antibodies in exposed seronegative individuals with HIV type 1-neutralizing activity. AIDS Res Hum Retroviruses 2000; 16:109–115.
Devito C, Broliden K, Kaul R, et al. Mucosal and plasma IgA from HIV-1-exposed uninfected individuals inhibit HIV-1 transcytosis across human epithelial cells. J Immunol 2000; 165:5170–6.
Brown L, Souberbielle BE, Marriott JB, et al. The conserved carboxy terminal region of HIV-1 gp120 is recognized by seronegative HIV-exposed people. Aids 1999; 13:2515–21.
Beretta A, Weiss SH, Rappocciolo G, et al. Human immunodeficiency virus type 1 (HIV-1)-seronegative injection drug users at risk for HIV exposure have antibodies to HLA class I antigens and T cells specific for HIV envelope. J Infect Dis 1996; 173:472–6.
Pinto LA, Sharpe S, Cohen DI, Shearer GM . Alloantigen-stimulated anti-HIV activity. Blood 1998; 92:3346–54.
Skurnick JH, Palumbo P, DeVico A, et al. Correlates of nontransmission in US women at high risk of human immunodeficiency virus type 1 infection through sexual exposure. J Infect Dis 2002; 185:428–38.
Liu R, Paxton WA, Choe S, et al. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 1996; 86:367–77.
Samson M, Libert F, Doranz BJ, et al. Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature 1996; 382:722–25.
Dean M, Carrington M, Winkler C, et al. Genetic restriction of HIV-1 infection and progression to AIDS by a deletion allele of the CKR5 structural gene. Hemophilia Growth and Development Study, Multicenter AIDS Cohort Study, Multicenter Hemophilia Cohort Study, San Francisco City Cohort, ALIVE Study. Science 1996; 273:1856–62.
Huang Y, Paxton WA, Wolinsky SM, et al. The role of a mutant CCR5 allele in HIV-1 transmission and disease progression. Nat Med 1996; 2:1240–3.
Michael NL, Chang G, Louie LG, et al. The role of viral phenotype and CCR-5 gene defects in HIV-1 transmission and disease progression. Nat Med 1997; 3:338–40.
Smith MW, Dean M, Carrington M, et al. Contrasting genetic influence of CCR2 and CCR5 variants on HIV-1 infection and disease progression. Hemophilia Growth and Development Study (HGDS), Multicenter AIDS Cohort Study (MACS), Multicenter Hemophilia Cohort Study (MHCS), San Francisco City Cohort (SFCC), ALIVE Study. Science 1997; 277:959–65.
Martin MP, Dean M, Smith MW, et al. Genetic acceleration of AIDS progression by a promoter variant of CCR5. Science 1998; 282:1907–11.
Liu H, Chao D, Nakayama EE, et al. Polymorphism in RANTES chemokine promoter affects HIV-1 disease progression. Proc Natl Acad Sci U S A 1999; 96:4581–5.
McDermott DH, Beecroft MJ, Kleeberger CA, et al. Chemokine RANTES promoter polymorphism affects risk of both HIV infection and disease progression in the Multicenter AIDS Cohort Study. Aids 2000; 14:2671–8.
Gonzalez E, Dhanda R, Bamshad M, et al. Global survey of genetic variation in CCR5, RANTES, and MIP-1alpha : Impact on the epidemiology of the HIV-1 pandemic. Proc Natl Acad Sci U S A 2001; 98:5199–204.
Ioannidis JP, Rosenberg PS, Goedert JJ, et al. Effects of CCR5-Delta32, CCR2–64I, and SDF-1 3′A alleles on HIV-1 disease progression: An international meta-analysis of individual-patient data. Ann Intern Med 2001; 135:782–95.
O'Brien SJ, Moore JP . The effect of genetic variation in chemokines and their receptors on HIV transmission and progression to AIDS. Immunol Rev 2000; 177:99–111.
Feng Y, Broder CC, Kennedy PE, Berger EA . HIV-1 entry cofactor: functional cDNA cloning of a seven-transmembrane, G protein-coupled receptor [see comments]. Science 1996; 272:872–7.
Deng H, Liu R, Ellmeier W, et al. Identification of a major co-receptor for primary isolates of HIV-1 [see comments]. Nature 1996; 381:661–6.
Dragic T, Litwin V, Allaway GP, et al. HIV-1 entry into CD4+ cells is mediated by the chemokine receptor CC-CKR-5. Nature 1996; 381:667–73.
Berger E, Doms R, Fenyö E, et al. A new classification for HIV-1. Nature 1998;391:240.
Connor RI, Sheridan KE, Ceradini D, Choe S, Landau NR . Change in coreceptor use correlates with disease progression in HIV-1-infected individuals. J Exp Med 1997; 185:621–8.
Connor RI, Paxton WA, Sheridan KE, Koup RA . Macrophages and CD4+ T lymphocytes from two multiply exposed, uninfected individuals resist infection with primary non-syncytium-inducing isolates of human immunodeficiency virus type 1. J Virol 1996; 70:8758–64.
Paxton WA, Martin SR, Tse D, et al. Relative resistance to HIV-1 infection of CD4 lymphocytes from persons who remain uninfected despite multiple high-risk sexual exposure. Nat Med 1996; 2:412–7.
Biti R, Ffrench R, Young J, et al. HIV-1 infection in an individual homozygous for the CCR5 deletion allele. Nat Med 1997; 3:252–3.
O'Brien TR, Winkler C, Dean M, et al. HIV-1 infection in a man homozygous for CCR5 delta 32. Lancet 1997; 349:1219–24.
Theodorou I, Meyer L, Magierowska M, Katlama C, Rouzioux C . HIV-1 infection in an individual homozygous for CCR5 delta 32. Seroco Study Group. Lancet 1997; 349:1219–20.
Balotta C, Bagnarelli P, Violin M, et al. Homozygous delta 32 deletion of the CCR-5 chemokine receptor gene in an HIV-1-infected patient. AIDS 1997; 11:F67–71.
Michael NL, Nelson JA, KewalRamani VN, et al. Exclusive and persistent use of the entry coreceptor CXCR4 by human immunodeficiency virus type 1 from a subject homozygous for CCR5 delta32. J Virol 1998; 72:6040–7.
Edelstein RE, Arcuino LA, Hughes JP, et al. Risk of mother-to-infant transmission of HIV-1 is not reduced in CCR5/delta32ccr5 heterozygotes. J Acquir Immune Defic Syndr Hum Retrovirol 1997; 16:243–6.
Malo A, Rommel F, Bogner J, et al. Lack of protection from HIV infection by the mutant HIV coreceptor CCR5 in intravenously HIV infected hemophilia patients. Immunobiology 1998; 198:485–8.
Misrahi M, Teglas JP, N'Go N, et al. CCR5 chemokine receptor variant in HIV-1 mother-to-child transmission and disease progression in children. French Pediatric HIV Infection Study Group Jama 1998; 279:277–80.
Shearer WT, Kalish LA, Zimmerman PA . CCR5 HIV-1 vertical transmission. Women and Infants Transmission Study Group. J Acquir Immune Defic Syndr Hum Retrovirol 1998; 17:180–1.
Mummidi S, Ahuja SS, Gonzalez E, et al. Genealogy of the CCR5 locus and chemokine system gene variants associated with altered rates of HIV-1 disease progression. Nat Med 1998; 4:786–93.
Hladik F, Liu H, Speelmon E, et al. Combined effect of CCR5-Delta32 heterozygosity and the CCR5 promoter polymorphism -2459 A/G on CCR5 expression and resistance to human immunodeficiency virus type 1 transmission. J Virol 2005; 79:11677–84.
Katzenstein TL, Eugen-Olsen J, Hofmann B, et al. HIV-infected individuals with the CCR delta32/CCR5 genotype have lower HIV RNA levels and higher CD4 cell counts in the early years of the infection than do patients with the wild type. Copenhagen AIDS Cohort Study Group. J Acquir Immune Defic Syndr Hum Retrovirol 1997; 16:10–4.
Zimmerman PA, Buckler-White A, Alkhatib G, et al. Inherited resistance to HIV-1 conferred by an inactivating mutation in CC chemokine receptor 5: studies in populations with contrasting clinical phenotypes, defined racial background, and quantified risk. Mol Med 1997; 3:23–36.
de Roda Husman AM, Koot M, Cornelissen M, et al. Association between CCR5 genotype and the clinical course of HIV-1 infection. Ann Intern Med 1997; 127:882–90.
Eugen-Olsen J, Iversen AK, Garred P, et al. Heterozygosity for a deletion in the CKR-5 gene leads to prolonged AIDS-free survival and slower CD4 T-cell decline in a cohort of HIV-seropositive individuals. AIDS 1997; 11:305–310.
Stewart GJ, Ashton LJ, Biti RA, et al. Increased frequency of CCR-5 delta 32 heterozygotes among long-term non- progressors with HIV-1 infection. The Australian Long-Term Non- Progressor Study Group. AIDS 1997; 11:1833–8.
Paxton WA, Kang S and Koup RA . The HIV type 1 coreceptor CCR5 and its role in viral transmission and disease progression. AIDS Res Hum Retroviruses 1998; 14 Suppl 1:S89–92.
Rappaport J, Cho YY, Hendel H, Schwartz EJ, Schachter F, Zagury JF . 32 bp CCR-5 gene deletion and resistance to fast progression in HIV-1 infected heterozygotes [letter]. Lancet 1997; 349:922–3.
Schinkel J, Langendam MW, Coutinho RA, Krol A, Brouwer M, Schuitemaker H . No evidence for an effect of the CCR5 delta32/+ and CCR2b 64I/+ mutations on human immunodeficiency virus (HIV)-1 disease progression among HIV-1-infected injecting drug users. J Infect Dis 1999; 179:825–31.
Wilkinson DA, Operskalski EA, Busch MP, Mosley JW, Koup RA . A 32-bp deletion within the CCR5 locus protects against transmission of parenterally acquired human immunodeficiency virus but does not affect progression to AIDS-defining illness. J Infect Dis 1998; 178:1163–6.
Martinson JJ, Chapman NH, Rees DC, Liu YT, Clegg JB . Global distribution of the CCR5 gene 32-basepair deletion. Nat Genet 1997; 16:100–3.
Libert F, Cochaux P, Beckman G, et al. The deltaccr5 mutation conferring protection against HIV-1 in Caucasian populations has a single and recent origin in Northeastern Europe. Hum Mol Genet 1998; 7:399–406.
Stephens JC, Reich DE, Goldstein DB, et al. Dating the origin of the CCR5-ð32 AIDS-resistance allele by the coalescence of haplotypes. Am J Hum Genet 1998; 62:1507–15.
Wang FS, Hong WG, Cao Y, et al. Population survey of CCR5 delta32, CCR5 m303, CCR2b 64I, and SDF1 3′A allele frequencies in indigenous Chinese healthy individuals, and in HIV-1-infected and HIV-1-uninfected individuals in HIV-1 risk groups. J Acquir Immune Defic Syndr 2003; 32:124–30.
Feng T, Ni A, Yang G, Galvin SR, Hoffman IF, Cohen MS . Distribution of the CCR5 gene 32-base pair deletion and CCR5 expression in Chinese minorities. J Acquir Immune Defic Syndr 2003; 32:131–4.
Hong W, Wang F, Jin L, et al. Genotyping of HIV resistant alleles in indigenous Tibetan ethnic group of China: low frequency of CCR5triangle32 and high frequency of CCR2b-64I alleles. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 2001; 18:268–71.
Zhang C, Fu S, Xue Y, et al. Distribution of the CCR5 gene 32-basepair deletion in 11 Chinese populations. Anthropol Anz 2002; 60:267–71.
Jiang JD, Wang Y, Wang ZZ, et al. Low frequency of the ccr5delta32 HIV-resistance allele in mainland China: identification of the first case of ccr5delta32 mutation in the Chinese population. Scand J Infect Dis 1999; 31:345–8.
Deng XL, Hong KX, Chen J-p, et al. Genetic polymorphisms of human immunodeficiency virus coreceptor CCR5 delta 32 and CCR2-64I alleles in Chinese Yi Ethnic group in Sichun. Chinese J Epidemiology 2004; 25:1050–3.
Wang XH, Feng TJ, Wang FS, et al. The effect of CCR delta32, CCR5m303, CCR2-64I and SDF1-3′A gene polymorphism to the prognosis of Chinese HIV-1 carriers. Chinese J Experimental Clinical Virology 2005; 19.
McDermott DH, Zimmerman PA, Guignard F, Kleeberger CA, Leitman SF, Murphy PM . CCR5 promoter polymorphism and HIV-1 disease progression. Multicenter AIDS Cohort Study (MACS). Lancet 1998; 352:866–70.
Li C, Lu SC, Hsieh PS, et al. Distribution of human chemokine (C-X3-C) receptor 1 (CX3CR1) gene polymorphisms and haplotypes of the CC chemokine receptor 5 (CCR5) promoter in Chinese people, and the effects of CCR5 haplotypes on CCR5 expression. Int J Immunogenet 2005; 32:99–106.
Zhang YJ, Moore JP . Will multiple coreceptors need to be targeted by inhibitors of human immunodeficiency virus type 1 entry? J Virol 1999; 73:3443–8.
Zhang YJ, Dragic T, Cao Y, et al. Use of coreceptors other than CCR5 by non-syncytium-inducing adult and pediatric isolates of human immunodeficiency virus type 1 is rare in vitro. J Virol 1998; 72:9337–44.
Kostrikis LG, Huang Y, Moore JP, et al. A chemokine receptor CCR2 allele delays HIV-1 disease progression and is associated with a CCR5 promoter mutation. Nat Med 1998; 4:350–3.
Rizzardi GP, Morawetz RA, Vicenzi E, et al. CCR2 polymorphism and HIV disease. Swiss HIV Cohort. Nat Med 1998; 4:252–3.
Michael NL, Louie LG, Rohrbaugh AL, et al. The role of CCR5 and CCR2 polymorphisms in HIV-1 transmission and disease progression. Nat Med 1997; 3:1160–2.
Eugen-Olsen J, Iversen AK, Benfield TL, Koppelhus U, Garred P . Chemokine receptor CCR2b 64I polymorphism and its relation to CD4 T-cell counts and disease progression in a Danish cohort of HIV-infected individuals. Copenhagen AIDS cohort. J Acquir Immune Defic Syndr Hum Retrovirol 1998; 18:110–6.
Mariani R, Wong S, Mulder LC, et al. CCR2-64I polymorphism is not associated with altered CCR5 expression or coreceptor function. J Virol 1999; 73:2450–9.
Lee B, Doranz BJ, Rana S, et al. Influence of the CCR2-V64I polymorphism on human immunodeficiency virus type 1 coreceptor activity and on chemokine receptor function of CCR2b, CCR3, CCR5, and CXCR4. J Virol 1998; 72:7450–8.
Winkler C, Modi W, Smith MW, et al. Genetic restriction of AIDS pathogenesis by an SDF-1 chemokine gene variant. Science 1998; 279:389–93.
van Rij RP, Broersen S, Goudsmit J, Coutinho RA, Schuitemaker H . The role of a stromal cell-derived factor-1 chemokine gene variant in the clinical course of HIV-1 infection. Aids 1998; 12:F85–90.
Wang H, Li B, Xu L, et al. Genetic analysis of human immunodeficiency virus type 1 strains from AIDS patients in Shenzhen. Chin J Lab Med 2005; 28.
Cocchi F, DeVico AL, Garzino-Demo A, Arya SK, Gallo RC, Lusso P . Identification of RANTES, MIP-1 alpha, and MIP-1 beta as the major HIV- suppressive factors produced by CD8+ T cells. Science 1995; 270:1811–1815.
An P, Nelson GW, Wang L, et al. Modulating influence on HIV/AIDS by interacting RANTES gene variants. Proc Natl Acad Sci U S A 2002; 99:10002–7.
Liu XL, Wang FS, Jin L, Liu MX, Xu DZ . Preliminary study on the association of chemokine RANTES gene polymorphisms with HIV-1 infection in Chinese Han population. Zhonghua Liu Xing Bing Xue Za Zhi 2003; 24:971–5.
Zhao XY, Lee SS, Wong KH, et al. Effects of single nucleotide polymorphisms in the RANTES promoter region in healthy and HIV-infected indigenous Chinese. Eur J Immunogenet 2004; 31:179–83.
Aguado B, Campbell RD . Characterization of a human MHC class III region gene product with S-thioesterase activity. Biochem J 1999; 341(Pt 3):679–89.
Parham P, Ohta T . Population biology of antigen presentation by MHC class I molecules. Science 1996; 272:67–74.
Barber LD, Gillece-Castro B, Percival L, Li X, Clayberger C, Parham P . Overlap in the repertoires of peptides bound in vivo by a group of related class I HLA-B allotypes. Curr Biol 1995; 5:179–90.
Gao X, Nelson GW, Karacki P, et al. Effect of a single amino acid change in MHC class I molecules on the rate of progression to AIDS. N Engl J Med 2001; 344:1668–75.
Moore CB, John M, James IR, Christiansen FT, Witt CS, Mallal SA . Evidence of HIV-1 adaptation to HLA-restricted immune responses at a population level. Science 2002; 296:1439–43.
Carrington M, Nelson GW, Martin MP, et al. HLA and HIV-1: heterozygote advantage and B*35-Cw*04 disadvantage [see comments]. Science 1999; 283:1748–1752.
Carrington M, O'Brien SJ . The influence of HLA genotype on AIDS. Annu Rev Med 2003; 54:535–51.
O'Brien SJ, Gao X and Carrington M . HLA, AIDS: a cautionary tale. Trends Mol Med 2001; 7:379–81.
Martin MP, Gao X, Lee JH, et al. Epistatic interaction between KIR3DS1 and HLA-B delays the progression to AIDS. Nat Genet 2002; 31:429–34.
Xu MY, Hong KX, Deng XL, et al. Association of HLA-B alleles with human immunodeficiency virus type 1 infection in the Yi ethnic group in Sichuan province. Biomed Environ Sci 2004; 17:203–8.
Stahl-Hennig C, Steinman RM, Tenner-Racz K, et al. Rapid infection of oral mucosal-associated lymphoid tissue with simian immunodeficiency virus. Science 1999; 285:1261–5.
Barratt-Boyes S, Watkins S, Finn O . In vivo migration of dendritic cells differentiated in vitro: a chimpanzee model. J Immunol 1997; 158:4543–7.
Geijtenbeek TB, Kwon DS, Torensma R, et al. DC-SIGN, a dendritic cell-specific HIV-1-binding protein that enhances trans-infection of T cells. Cell 2000; 100:587–97.
Pohlmann S, Baribaud F, Lee B, et al. DC-SIGN Interactions with Human Immunodeficiency Virus Type 1 and 2 and Simian Immunodeficiency Virus. J Virol 2001; 75:4664–72.
Bashirova AA, Geijtenbeek TBH, van Duijnhoven GCF, et al. A Dendritic Cell-specific Intercellular Adhesion Molecule 3-grabbing Nonintegrin (DC-SIGN)-related Protein Is Highly Expressed on Human Liver Sinusoidal Endothelial Cells and Promotes HIV-1 Infection. J Exp Med 2001; 193:671–78.
Pohlmann S, Soilleux EJ, Baribaud F, et al. DC-SIGNR, a DC-SIGN homologue expressed in endothelial cells, binds to human and simian immunodeficiency viruses and activates infection in trans. Proc Natl Acad Sci U S A 2001; 98:2670–5.
Liu H, Hladik F, Andrus T, et al. Most DC-SIGNR transcripts at mucosal HIV transmission sites are alternatively spliced isoforms. Eur J Hum Genet 2005; 13:707–15.
Soilleux EJ, Barten R, Trowsdale J . Cutting Edge: DC-SIGN; a Related Gene, DC-SIGNR; and CD23 Form a Cluster on 19p13. J Immunol 2000; 165:2937–42.
Mummidi S, Catano G, Lam L, et al. Extensive Repertoire of Membrane-bound and Soluble Dendritic Cell-specific ICAM-3-grabbing Nonintegrin 1 (DC-SIGN1) and DC-SIGN2 Isoforms. INTER-INDIVIDUAL VARIATION IN EXPRESSION OF DC-SIGN TRANSCRIPTS. J Biol Chem 2001; 276:33196–212.
Liu HL, Carrington M, Wang CW, et al. Effects of Genetic Polymorphisms in the DC-SIGNR Repeat Region onHIV-1 Susceptibility. Journal of Infectious Disease 2005.
Martin MP, Lederman MM, Hutcheson HB, et al. Association of DC-SIGN promoter polymorphism with increased risk for parenteral, but not mucosal, acquisition of human immunodeficiency virus type 1 infection. J Virol 2004; 78:14053–6.
Quillent C, Oberlin E, Braun J, et al. HIV-1-resistance phenotype conferred by combination of two separate inherited mutations of CCR5 gene. Lancet 1998; 351:14–8.
Carrington M, Kissner T, Gerrard B, Ivanov S, O'Brien SJ, Dean M . Novel alleles of the chemokine-receptor gene CCR5. Am J Hum Genet 1997; 61:1261–7.
Ansari-Lari MA, Liu XM, Metzker ML, Rut AR, Gibbs RA . The extent of genetic variation in the CCR5 gene. Nat Genet 1997; 16:221–2.
Ma L, Dudoit Y, Tran T, et al. Biochemical and HIV-1 coreceptor properties of K26R, a new CCR5 Variant in China's Sichuan population. J Acquir Immune Defic Syndr 2005; 39:38–43.
Zhao XY, Lee SS, Wong KH, et al. Functional analysis of naturally occurring mutations in the open reading frame of CCR5 in HIV-infected Chinese patients and healthy controls. J Acquir Immune Defic Syndr 2005; 38:509–17.
Martin MP, Carrington M, Dean M, et al. CXCR4 polymorphisms and HIV-1 pathogenesis. J Acquir Immune Defic Syndr Hum Retrovirol 1998; 19:430.
Wegner SA, Ehrenberg PK, Chang G, Dayhoff DE, Sleeker AL, Michael NL . Genomic organization and functional characterization of the chemokine receptor CXCR4, a major entry co-receptor for human immunodeficiency virus type 1. J Biol Chem 1998; 273:4754–60.
Acknowledgements
We thank information and discussion from Drs. Xiao Hui WANG, Chun Hui WANG and Huan Liang LIU. This work was supported by Public Health Service grants AI 45402, AI 49109, P30 AI-27757-18 (S3), and AI 056994 (to T. Zhu).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
ZHU, T., FENG, T., XIAO, X. et al. Global human genetics of HIV-1 infection and China. Cell Res 15, 833–842 (2005). https://doi.org/10.1038/sj.cr.7290355
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290355