ABSTRACT
Plants become photosynthetic through de-etiolation, a developmental process regulated by red/far-red light-absorbing phytochromes and blue/ultraviolet A light-absorbing cryptochromes. Genetic screens have identified in the last decade many far-red light signaling mutants and several red and blue light signaling mutants, suggesting the existence of distinct red, far-red, or blue light signaling pathways downstream of phytochromes and cryptochromes. However, genetic screens have also identified mutants with defective de-etiolation responses under multiple wavelengths. Thus, the optimal de-etiolation responses of a plant depend on coordination among the different light signaling pathways. This review intends to discuss several recently identified signaling components that have a potential role to integrate red, far-red, and blue light signalings. This review also highlights the recent discoveries on proteolytic degradation in the desensitization of light signal transmission, and the tight connection of light signaling with photoperiodic flowering and circadian rhythm. Studies on the controlling mechanisms of de-etiolation, photoperiodic flowering, and circadian rhythm have been the fascinating topics in Arabidopsis research. The knowledge obtained from Arabidopsis can be readily applied to food crops and ornamental species, and can be contributed to our general understanding of signal perception and transduction in all organisms.
Similar content being viewed by others
PLANT DE-ETIOLATION IS TRIGGERED BY LIGHT SIGNALS
Light is arguably the most important resource for plants, and plants have evolved an array of photosensory pigments enabling them to develop optimally in a broad range of ambient light conditions. The photoreceptors include red and far-red-absorbing phytochromes and UV-A/blue light-absorbing crytochromes and phototropins 1, 2, 3. Among the photoreceptors, phytochromes and cryptochromes regulate seedling de-etiolation responses, photoperiodic flowering, and circadian rhythm, whereas phototropins function to improve the efficiency of photosynthesis by regulating phototropic response, stomatal opening, and chloroplast relocation movement. The de-etiolation responses include the inhibition of hypocotyl elongation, the opening of cotyledons and hypocotyl hooks, and the development of chloroplasts.
PHYTOCHROMES AND CRYPTOCHROMES
Phytochromes exist in dimeric forms with a covalently attached linear tetrapyrrole chromophore near their N-termini and two putative dimerization sites in their C-termini 2. An important property of phytochromes is their capacity to undergo photoconversion between the red light-absorbing (Pr) form and the far-red light-absorbing (Pfr) form upon sequential absorption of the red or far-red light photons. In Arabidopsis, phytochrome gene family has five members, PHYA, PHYB, PHYC, PHYD, and PHYE 4. The specificity for the light perception is confined to the N-terminus 5. phyA is the sole photoreceptor to mediate far-red light de-etiolation responses and phyB is the major photorecptor to mediate red light de-etiolation responses. As a result, PHYA null mutants have reduced responses to continuous far-red light but have the same phenotype as wild-type in continuous red light 2. In contrast, PHYB null mutants are defective in red light-mediated de-etiolation but are normal as wild type in continuous far-red light 2. Recent studies show that phyB N-terminus, when dimerized and localized to the nucleus, triggered full phyB responses with much higher photosensitivity than full-length phyB, suggesting that the C-terminus attenuates phyB activity 6. phyA is able to autophosphorylate and to phosphorylate phytochrome kinase substrate 1 (PKS1) in vitro 7, 8. Both phyA and phyB have been demonstrated to exhibit a light-dependent translocation to the nucleus 9, 10, 11. The nuclear translocation of phyB is triggered by red light, whereas the nuclear translocation of phyA is triggered by both red light and far-red light.
CRY1 mutation impairs seedling de-etiolation responses under blue/UV-A light 1. CRY1 encodes a flavoprotein with sequence similarity to photolyases, a family of flavoproteins that mediates repair of DNA damage by UV light. However, cry1 lacks photolyase activity and has a C-terminal extension not found in the photolyases 1. Dark-grown Arabidopsis seedlings carrying the C-terminal domains of either cry1 or cry2 show phenotypes that are normally associated with light-grown seedlings and are often observed for cop1 12. The signaling activity of cry1 involves a direct interaction of its C-terminus with COP1, the negative regulator of photomorphogenesis 13. cry2 is involved in the control of photoperiodic flowering in addition to its role in regulation of seedling de-etiolation responses 14. cry1 is localized to the nucleus under dark condition and is depleted from the nucleus under continuous white light condition 12, 15. In contrast, cry2 is predominantly localized to the nucleus under both dark and light conditions 12, 15. Both cry1 and cry2 also undergo a blue-light-dependent phosphorylation and the phosphorylation status is closely associated with their function and regulation 16, 17. The cry2 protein is unstable and this instability may be mediated through an interaction with COP1 18.
LIGHT SIGNALING COMPONENTS IDENTIFIED THROUGH GENETIC SCREENS
Genetic approaches, largely based on seedling de-etiolation responses, have identified many far-red light signaling mutants, including fhy1 19, fhy3 20, spa121, fin2 22, far1 23, hfr1 or rep1 24, 25, 26, fin219 27, pat1 28, eid1 29, and laf1 30. Other mutants have been isolated for their defective red light responses, such as pef2 and pef3 31, srl1 32, gi 33, pif4 34, and srr1 35. Recently, a blue light signaling component, PP7, has been identified through a reverse genetic approach 36. Those studies together suggest the existence of distinct red, far-red, or blue light signaling pathways. Many of the signaling components reside in the nucleus but some such as FIN219 27 and PAT1 28 are cytosolic proteins. FHY1 and SRR1 exist in both the nucleus and the cytoplasm 19, 35. However, it is still unclear how the components, either cytosolic or nuclear, work together to transduce red, far-red, or blue light signals for growth and development.
LIGHT SIGNALING COMPONENTS IDENTIFIED BY BIOCHEMICAL AND INTERACTION STUDIES
Early analysis of potential components involved in phytochrome signaling has been addressed by microinjection and pharmacological techniques 37, 38, 39. In these studies, some well-known second messengers were examined for their ability to restore the light responses of a tomato aurea mutant, a chromophore biosynthesis mutant. Evidences thus obtained by these techniques suggest the involvement of cyclic GMP, G-proteins, and calcium/calmodulin in phytochrome signaling cascades. Recently, a heterotrimeric G-protein α-subunit has also been implicated in light signaling 40.
Five phytochrome interacting proteins, PIF3, PKS1, NPDK2, FyPP, and PAPP5, have been isolated through yeast two-hybrid screens 8, 41, 42, 43, 44. PIF3 is a nuclear basic helix-loop-helix protein and binds to the Pfr forms of both phyB 45 and phyA in vitro 24. Re-conversion of the Pfr form the Pr form by far-red light abruptly terminates the interaction 45. PKS1 encodes a cytoplasmic protein whereas NDPK2 is localized in both the nucleus and the cytosol 8, 41. PKS1 is detected in plants in the phosphorylated form, but red light enhances the degree of PKS1 phosphorylation. FyPP encodes the catalytic subunit of a Ser/Thr-specific protein phosphatase 2A 43. Recombinant FyPP efficiently dephosphorylated oat phytochrome A in a spectral form–dependent manner. Transgenic Arabidopsis plants with overexpressed or suppressed FyPP levels caused a delayed or accelerated flowering, respectively. PAPP5 specifically dephosphorylates biologically active Pfr-phytochromes and enhances phytochrome-mediated photoresponses 44. The dephosphorylation may enhance phytochrome stability and the binding affinity of phytochrome to NDPK2. In addition, targeted yeast two-hybrid or in vitro pull-down assays have suggested possible interactions of phyA or phyB with PIF1, PIF4, ARR4, and ELF3 34, 46, 47, 48.
INTEGRATION OF RED, FAR-RED, AND BLUE LIGHT SIGNALING
Despite the existence of distinct red, far-red, or blue light signaling pathways, genetic screens have also identified mutants with defective seedling de-etiolation responses under multiple wavelengths. Thus, the optimal performance of a plant depends on coordination among the different light signaling pathways. It has been realized that a minimal level of active phytochrome seems to be necessary for full activity of cryptochromes or phototropins 49. One classic example is the enhancement by red light of phototropic bending toward unilateral blue light. The dependence of the blue responses on active phytochromes may result from a direct interaction of both photoreceptors, and early studies indicated that cry1 and cry2 can be phosphorylated by a phytochrome A-associated kinase activity 49. The enzymatic interaction of phyA with cry1 was further confirmed in targeted yeast two-hybrid assays. The studies suggest that photoactivated phytochrome can phosphorylate a number of substrates including crys, and subsequent exposure of crys to blue light would enhance the signaling activities of crys. A functional interaction of phyB with cry2 has also been implicated in control of hypocotyl elongation, flowering time, and circadian rhythm 50. In vivo coimmunoprecipitation, colocalization, and FRET anlaysis showed a direct interaction of phyB with cry2 in nuclear speckles, suggesting a possible integration of blue and red light signaling at photoreceptor level.
Equally possible, the dependence of blue light responses on active phytochromes may occur at a common intermediate step of their signaling pathways. Fig. 1 lists all components isolated so far through forward genetics that are likely involved in integration of more than one light signaling pathways. We have identified a short hypocotyl mutant under red and blue light, hrb1 for hypersensitive to red and blue 1 51. Mutation in HRB1 also enhances the end-of-day far-red light response, inhibits leaf expansion and petiole elongation, and attenuates the expression of CAB3 and CHS genes. HRB1 is localized to the nucleus and belongs to a protein family of Drought induced 19 or Di19. HRB1 and all other family members contain a ZZ-type zinc finger domain, which in other organisms is implicated in protein-protein interactions between dystrophin and calmodulin, and between transcriptional adaptors and activators. HRB1 activity is also required for the proper expression of PIF4 under red and blue light. pif4 shows a very similar hypersensitive response as hrb1 to both red and blue light and is epistatic to hrb1 in control of light-regulated gene expression responses. Thus, HRB1 and PIF4 together may define points where red light signaling and blue light signaling intersect. Recently, a gain-of-function mutant, sob1-D for suppressor of phytochrome B-4 dominant, has been reported and its phenotype is caused by the overexpression of a nuclear Dof transcription factor, OBF4 Binding Protein 3 or OBP3 52. Antisense lines with reduced OBP3 expression are less responsive to red light in the inhibition of hypocotyl elongation, and also have larger cotyledons under red and blue light, suggesting a role of OBP3 in both phyB and cry1 signaling.
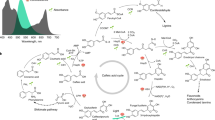
Integrative controls of de-etiolation, photoperiodic flowering, and circadian regulation. Phytochromes and cryptochromes perceive red, far-red, or blue light signals, and several components downstream of the photoreceptors integrate red and blue (PIF4, OBP3, and HRB1), red and far-red (COG1, RFI2, and PFT1), and far-red and blue (SUB1 and HFR1) light signaling for de-etiolation. Among them, HRB1 and RFI2 are under circadian-regulation similar to most clock or flowering genes, whereas HRB1, RFI2, and PFT1 play a role in photoperiodic control of CO or FTexpression. Phytochromes and cryptochromes are also involved in the resetting of the circadian clock, and ELF3 mediates between photoreceptors and the circadian clock. LHY/CCA1 and TOC1/ELF4 form a negative feedback loop within the circadian oscillator, and PRR7, ZTL, and FKF1 function closely with the circadian oscillator. Besides their circadian or flowering function, SRR1, GI, ELF3, ELF4, FKF1, and ZTL also act in red light-mediated de-etiolation, whereas TOC1 and PRR7 act in red/far-red light-mediated de-etiolation. Downstream of the clock, GI mediates the regulation of CO expression, a key gene in photoperiodic flowering that regulates the expression of FT, an integrator of several flowering pathways. Mutation in TOC1, CCA1, or LHY causes early flowering and mutation in FKF1 or ZTL causes late flowering. Arrows represent positive effects and perpendicular lines represent negative effects.
SUB1, a cytoplasmic calcium-binding protein, plays a role to integrate blue and far-red light signaling 53. Mutation in SUB1 causes a hypersensitive hypocotyl growth response and much enhanced CHS and CHI expression under relatively low fluence rates of blue and far-red light. Genetic analysis indicates that SUB1 functions downstream of crys and modulates phyA-mediated far-red light responses. SUB1 localizes in the nuclear periphery region surrounding the nucleus, and may regulate light responses by suppressing light-dependent accumulation of HY5 protein. HFR1, a bHLH transcription factor, was initially isolated based on a defect in a subset of phyA-mediated far-red light responses in hfr1 mutant 24. Recently, hfr1 has been shown to have reduced de-etiolation responses, including hypocotyl elogation, cotyledon expansion, and anthocyanin accumulation, under high fluence rates of blue light 54. Genetic analysis indicates that HFR1 function in cry1 signaling pathway since cry1 is the major photoreceptor responsible for de-etiolation under strong blue light. Although SUB1 and HFR1 function in both blue and far-red light signaling pathways, they may use quite different mechanisms to integrate phyA and cry signaling.
Six other mutants show defective de-etiolation responses under both red and far-red light, including cog1, pef1, psi2, pft1, prr7, and rfi2 31, 55, 56, 57, 58, Chen and Ni, unpublished. Both cog1 and psi2 show hypersensitive hypocotyl growth response to red and far-red light, and COG1 and PSI2 thus encode negative components of phyA and phyB signaling 55, 56. The cog1-D mutation is caused by activation of a Dof domain-containing transcription factor, and transgenic lines expressing antisense COG1 results in a hypersensitive response to red and far-red light. In contrast, pef1, prr7, and rfi2 show a hyposensitive hypocotyl growth response to both red and far-red light, suggesting a positive role of these components in phyA and phyB signaling 31, 58. Chen and Ni, unpublished. PRR7 encodes a PSEUDO-RESPONSE REGULATOR 58. RFI2 encodes a nuclear protein with a C3H2C3-type zinc finger or RING-domain known to mediate protein-protein interactions [Chen and Ni, unpublished]. Interestingly, pft1 was hypo-responsive to far-red light but hyper-responsive to red light 57. PFT1 may therefore function at a phytochrome signaling node where antagonistic interactions between phyA and phyB occur.
Another Arabidopsis mutant, rfi1-1 for red and far-red insensitive 1-1, shows a long hypocotyl under red, far-red, and blue light [Kang and Ni, unpublished]. The long hypocotyl phenotype is caused by an overaccumulation of RFI1 transcript, and is recapitulated by overexpression of RFI1 in transgenic Arabidopsis. However, rfi1-2, a knockout allele of RFI1, exhibits a short hypocotyl phenotype only under blue light. Thus, RFI1 functions specifically in blue light signaling but overexpression of RFI1 expands its activity to modulate phytochrome signaling. Studies on both rfi1-1 and rfi1-2 demonstrated that RFI1 acts either positively or negatively. The positive action of RFI1 signaling may involve HFR1, a basic helix-loop-helix protein, based on the opposite effects of rfi1-1 and rfi1-2 mutations on HFR1 expression and the overlapping phenotypes of rfi1-2 and hfr1 under blue light. RFI1 localizes to the cytosol, and contains an N-terminal SPX and a C-terminal EXS domain found in members of the SYG1 protein family from fungi, C. elegans, fly, mammals, and Arabidopsis.
LIGHT SIGNALING IS MODULATED BY PROTEIN DEGRADATION
Mutants in a different class exhibit a light-grown phenotype even when grown in darkness, known as the cop/det/fus family with total of 11 loci 59. Many of the proteins are involved in control of the stability of some key light signaling components 59, 60. One of them, COP1, encodes an E3 ubiquitin ligase containing a Ring-finger domain, followed by a coiled-coil domain and a WD40 domain. In darkness, COP1 interacts directly with HY5, a bZIP transcription factor involved in photomorphogenesis, and targets it to proteasome-mediated degradation 59. The proteasome-mediated degradation of HY5 appears to involve SPA1, a negative regulator of phyA signaling that also contains a coiled-coil region and a WD40 domain 61. Indeed, COP1 and SPA1 interact through their coiled-coil regions, and the coiled-coil domain of SPA1 is able to enhance the E3 ligase activity of COP1 towards LAF1, a transcription factor involved in phyA signaling 60. COP1 also targets HYH, a novel bZIP protein predominantly involved in blue light regulation of development, for dark-specific degradation 62.
The role of COP1 in phyA degradation has also been documented in the desensitization of plant light signal transmission and the termination of signaling through proteolytic down-regulation ensures that responses to a single stimulatory event are not perpetuated indefinitely 63. In addition, a controlled degradation of the basic-loop-helix transcription factor PIF3 also appears to be a major regulatory step in light signaling 64. Recently, COP1 has been shown an ubiquitin ligase activity toward another basic-loop-helix transcription factor, HFR1, whereas light enhances HFR1 protein stabilization and accumulation 65, 66, 67. A N-terminal domain, conserved among several basic-loop-helix class proteins involved in light signaling, interacts with the COP1 WD40 domain and is identified as a determinant of HFR1 stability. Therefore, COP1 in darkness might repress light responses by targeting a subset of positive regulators for destruction. On exposure to light, depletion of COP1 in the nucleus allows the positive regulators to accumulate and to promote downstream photomorphogenic responses.
INTEGRATION OF LIGHT SIGNALING WITH PHOTOPERIODIC FLOWERING
Arabidopsis is a facultative long-day plant for which flower initiation is accelerated under long-day photoperiod but delayed under short-day photoperiod 68. Among the phytochrome photoreceptors, phyB mediates a red light inhibition of flowering under both long-day and short-day conditions. phyA promotes flowering possibly through two independent mechanisms: suppressing phyB function or promoting flowering independent of phyB 69, 70. cry2-deficient Arabidopsis plants have a delayed flowering phenotype in response to extended photoperiods 14. The regulation of flowering time by phytochromes is partially through their regulation on the expression of CONSTANS (CO), a key component in the photoperiodic flowering pathway. The abundance of CO mRNA was reduced in phyA mutant but was increased in phyB mutant 57, 71, 72. In addition, the expression of CO is also enhanced under long-day photoperiod and subject to circadian regulation 73. Recent studies suggest that CO protein also appears under posttranscriptional regulation by light signals, and different photoreceptors have distinct roles to modulate CO activity 74. Crys and phyA stabilize CO protein under blue light and far-red light, whereas phyB promotes the degradation of CO under red light to generate a daily rhythm in CO abundance. However, the signaling events involved in photoreceptor regulation on key flowering genes remain largely unknown.
Two classes of mutants with a reduced response to day length have been isolated: those that flower later than wild-type plants under long days but are unaffected under short days or, alternatively, early-flowering mutants under short days. Many flowering mutants, although initially isolated for their flowering phenotypes, also exhibit certain defects in their seedling de-etiolation responses (Fig. 1). GI is required for maintaining circadian amplitude and appropriate period length 77. Mutation in GI causes a late flowering phenotype under inductive long-day condition and also a long hypocotyl phenotype under red light 33, 78. Another mutant, pft1, shows a mild hypocotyl phenotype, but displays a strong late-flowering phenotype under long-day condition 57. Mutation in PFT1 also completely suppresses the early-flowering phenotype of phyB, suggesting that PFT1 mainly functions to regulate flowering downstream of phyB in a photoperiod-independent pathway. Mutation in ELF4 causes an early flowering phenotype under short-day photoperiod, whereas mutation in ELF3 results in an early flowering phenotype under both long-day and short-day photoperiods 75, 79. Both elf3 and elf4 have a longer hypocotyl under red light, and cause a general disruption of circadian rhythms 47, 75, 76, 79. For example, elf4 shows attenuated expression of CCA1, a gene that may function as a central oscillator 75. ELF4 thus functions either in a circadian oscillator or by conferring accuracy and persistence on a circadian oscillator, whereas ELF3 is probably a circadian clock input pathway component.
rfi2, a red/far-red long hypocotyl mutant identified in our laboratory, flowers early under both long-day and short-day photoperiods similar to phyB-9 but in contrast to phyA-211 (Chen and Ni, unpublished). The early flowering phenotype is accountable by an enhanced expression of CO and FT, genes that promotes floral transition. Further genetic analysis indicated an epistasis of co-2 to rfi2-1. RFI2 also showed a long-day photoperiod-enhanced expression and a free-running circadian rhythm similar to CO. RFI2 acts differently from ELF3 and ELF4 since the circadian regulation and outputs are not affected in rfi2-1, suggesting a function of RFI2 under the circadian regulation. RFI2 therefore reveals a previously unidentified step that integrates phytochrome and circadian signals in control of CO expression and photoperiodic flowering. In contrast, hrb1, a red/blue short hypocotyl mutant identified in our laboratory, flowers late under both long-day and short-day conditions [Kang and Ni, unpublished]. Overexpression of HRB1 creates an early flowering phenotype under short-day condition. The flowering phenotypes are caused by an altered expression of FT in hrb1 or HRB1 overexpression lines. Like many other flowering genes, HRB1 expression is also regulated by the circadian clock.
INTEGRATION OF LIGHT SIGNALING WITH CIRCADIAN REGULATION
In the simplest model, the circadian clock is composed of three main components, input pathways that perceive and transmit the environmental information to reset a central oscillator, the central oscillator that generates a 24 h period, and output pathways that generate physiological rhythms 80. LHY, CCA1 and TOC1 consist of part of the central mechanism that generates circadian rhythms in plants. LHY and CCA1 were proposed to act along with TOC1 in a transcriptional feedback loop in which TOC1, which is expressed only in the evening, promotes the expression of LHY/CCA1 at dawn, and in turn LHY/CCA1 represses the expression of TOC1 81. Recent research has identified another player, ELF4, that could act together with TOC1 to induce the expression of LHY/CCA1 75.
Light is one of the environmental cues to entrain the circadian clock. Phytochromes, cryptochromes, and their signaling components therefore function in the input pathways. ELF3 appears to mediate between the photoreceptors and the circadian clock 47. SRR1 also functions to integrate light signals to control circadian responses since srr1 is altered not only in phyB-controlled hypocotyl elongation but also in multiple outputs of the circadian clock 35. prr7 was initially isolated by its long hypocotyls under red and far-red light 58. The absence of PRR7 or PSEUDO-RESPNSE REGULATOR 7 also causes a coordinated 3 to 6 hr shift in the phasing of the oscillatory expression of CCA1, LHY, and TOC1. PRR7 belongs to a small gene family called TOC1/APRR1 that includes TOC1 or TIMING-OF-CAB1. The proteins in this family lack the conserved phospho-accepting Asp of the bacterial response regulators.
On the other hand, many of the clock function-associated genes, such as CCA1, LHY, TOC1, FKF1, and ZTL, are also involved in light-induced de-etiolation and photoperiodic flowering responses (Fig. 1). These genes were initially isolated from a number of recessive mutations that alter the free-running period of the Arabidopsis circadian clock 80, 82, 83, 84, 85. Since phytochrome and cryptochrome signaling is required for proper circadian oscillations, modulations of the de-etiolation process by some clock function-associated components are not unexpected. Mutations in TOC1, LHY and CCA1 genes cause early flowering under short days, and defective hypocotyl elongation responses 80, 82, 83. TOC1 has been particularly documented for its involvement in red and far-red light control of hypocotyl elongation and red light-mediated regulation of CCA1 and LHY expression during early seedling development 80. FKF1 and ZTL encode proteins containing a N-terminal PAS domain, an F-box, and six C-terminal kelch repeats. fkf1 and ztl are hypersensitive to red light in their hypocotyl elongation responses and flower late 84, 85. A simple view to define a signaling component in a particular pathway, light signaling or photoperiodic flowering or circadian rhythm, is apparently biased. This review has described a few pieces in the puzzle of interactions among light signaling, photoperiodic flowering, and circadian rhythm. The underlying mechanisms and the molecular and biochemical events involved remain to be elucidated in the years to come.
References
Cashmore AR . Cryptochromes: enabling plants and animals to determine circadian time. Cell 2003; 114:537–43.
Quail PH . Phytochrome photosensory signaling network. Nat Rev Mol Cell Biol 2002; 3:85–93.
Wada M, Kagawa T, Sato Y . Chloroplast movement. Annu Rev Plant Biol 2003; 54:455–68.
Clack T, Mathews S, Sharrock RA . The phytochrome apoprotein family in Arabidopsis is encoded by five genes: the sequences and expression of PHYD and PHYE. Plant Mol Biol 1994; 25:413–27.
Wanger D, Fairchild CD, Kuhn RM, Quail PH . Chromophore-bearing NH2-terminal domains of phytochromes A and B determine their photosensory specificity and differential light lability. Proc Natl Acad Sci U S A 1996; 92:4011–5.
Matsushita T, Mochizuki N, Nagatani A . Dimers of the N-terminal domain of phytochrome B are functional in the nucleus. Nature 2003; 424:571–4.
Yeh K, Lagarias C . Eukaryotic phytochromes: light-regulated serine/threonine protein kinases with histidine kinase ancestry. Proc Natl Acad Sci U S A 1998; 95:13976–81.
Fankhauser C, Yeh AC, Lagarias JC, et al. PKS1, a substrate phosphorylated by phytochrome that modulates light signaling in Arabidopsis. Science 1999; 284:1539–41.
Sakamoto K, Nagatani A . Nuclear localization activity of phytochrome B. Plant J 1996; 10:859–68.
Kircher S, Kozma-Bognar L, Kim L, et al. Light quality-dependent nuclear import of the plant photoreceptors phytochrome A and B. Plant Cell 1999; 11:1445–56.
Yamaguchi R, Nakamura M, Mochizuki N, Kay SA, Nagatani A . Light-dependent translocation of a phytochrome B-GFP fusion protein to the nucleus in transgenic Arabidopsis. J Cell Biol 1999; 145:437–45.
Yang H, Wu Y, Tang R, et al. The C-termini of Arabidopsis cryptochrome mediate a constitutive light response. Cell 2000; 103:815–27.
Yang H, Tang R, Cashmore AR . The signaling mechanism of Arabidopsis CRY1 involves direct interaction with COP1. Plant Cell 2001; 13:2573–87.
Guo H, Yang H, Mockler T, Lin C . Regulation of flowering time by Arabidopsis photoreceptors. Science 1998; 279:1360–3.
Guo H, Duong H, Ma N, Lin C . The Arabidopsis blue light receptor cryptochrome 2 is a nuclear protein regulated by a blue light-dependent post-translational mechanism. Plant J 1999; 19:279–87.
Shalitin D, Yang H, Mockler TC, et al. Regulation of Arabidopsis cryptochrome 2 by blue-light-dependent phosphorylation. Nature 2002; 417:763–7.
Shalitin D, Yu X, Maymon M, Mockler T, Lin C . Blue light–dependent in vivo and in vitro phosphorylation of Arabidopsis cryptochrome 1. Plant Cell 2003; 15:2421–9.
Wang H, Ma LG, Li JM, Zhao HY, Deng XW . Direct interaction of Arabidopsis cryptochromes with COP1 in light control development. Science 2001; 294:154–8.
Desnos T, Puente P, Whitelam GC, Harberd NP . FHY1: a phytochrome A-specific signal transducer. Genes Dev 2001; 15:2980–90.
Wang H, Deng XW . Arabidopsis FHY3 defines a key phytochrome A signaling component directly interacting with its homologous partner FAR1. EMBO J 2002; 21:1339–49.
Hoecker U, Tepperman JM, Quail PH . SPA1, a WD-repeat protein specific to phytochrome A signal transduction. Science 1999; 284:496–9.
Soh MS, Hong SH, Hanzawa H, Furuya M, Nam HG . Genetic identification of FIN2, a far red light-specific signaling component of Arabidopsis thaliana. Plant J 1998; 16:411–9.
Hudson M, Ringli C, Boylan MT, Quail PH . The FAR1 locus encodes a novel nuclear protein specific to phytochrome A signaling. Genes Dev 1999; 13:2017–27.
Fairchild CD, Schumaker MA, Quail PH . HFR1 encodes an atypical bHLH protein that acts in phytochrome A signal transduction. Genes Dev 2000; 14:2377–91.
Fankhauser C, Chory J . RSF1, an Arabidopsis locus implicated in phytochrome A signaling. Plant Physiol 2000; 124:39–46.
Soh MS, Kim YM, Han SJ, Song PS . REP1, a basic helix-loop-helix protein, is required for a branch pathway of phytochrome A signaling in Arabidopsis. Plant Cell 2000; 12:2061–73.
Hsieh HL, Okamoto H, Wang M, et al. FIN219, an auxin-regulated gene, defines a link between phytochrome A and the downstream regulator COP1 in light control of Arabidopsis development. Genes Dev 2002; 14:1958–70.
Bolle C, Koncz C, Chua NH . PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction. Genes Dev 2000; 14:1269–78.
Dieterle M, Zhou YC, Schafer E, Funk M, Kretsch T . EID1, an F-box protein involved in phytochrome A-specific light signaling. Genes Dev 2001; 15:939–44.
Ballesteros ML, Bolle C, Lois LM, et al. LAF1, a myb transcription factor activator for phytochrome A signaling. Genes Dev 2001; 15:2613–25.
Ahmad M, Cashmore AR . The pef mutants of Arabidopsis thaliana define lesions early in the phytochrome signaling pathway. Plant J 1996; 10:1103–10.
Huq E, Kang Y, Halliday KJ, Qin M, Quail PH . SRL1: a new locus specific to the phyB-signaling pathway in Arabidopsis. Plant J 2002; 4:461–70.
Huq E, Tepperman JM, Quail PH . GIGANTEA is a nuclear protein involved in phytochrome signaling in Arabidopsis. Proc Natl Acad Sci U S A 2000; 97:9789–94.
Huq E, Quail1 PH . PIF4, a phytochrome-interacting bHLH factor, functions as a negative regulator of phytochrome B signaling in Arabidopsis. EMBO J 2002; 21:2441–50.
Staiger D, Allenbach L, Salathia N, et al. The Arabidopsis SRR1 gene mediates phyB signaling and is required for normal circadian clock function. Genes Dev 2003; 17:256–68.
Mφller SG, Kim YS, Kunkel T, Chua NH . PP7 is a positive regulator of blue light signaling in Arabidopsis. Plant Cell 2003; 15:1111–9.
Neuhaus G, Bowler C, Hiratsuka K, Yamagata H, Chua NH . Phytochrome-regulated repression of gene expression requires calcium and cGMP. EMBO J 1997; 16:2554–64.
Romero LC, Lam E . Guanine nucleotife binding protein involvement in early steps of phytochrome-regulated gene expression. Proc Natl Acad Sci U S A 1993; 90:1465–9.
Bowler C, Neuhaus G, Yamagata H, Chua NH . Cyclic GMP and calcium mediate phytochrome phototransduction. Cell 1994; 77:73–81.
Okamoto H, Matsui M, Deng, XW . Overexpression of the heterotrimeric G-proteinα-subunit enhances phytochrome-mediated inhibition of hypocotyl elongation in Arabidopsis. Plant Cell 2001; 13:1639–51.
Choi G, Yi H, Lee J, et al. Phytochrome signalling is mediated through nucleoside diphosphate kinase 2. Nature 1999; 401:610–3.
Ni M, Tepperman JM, Quail PH . PIF3, a phytochrome-interacting factor necessary for normal photoinduced signal transduction, is a novel basic helix-loop-helix protein. Cell 1998; 95:657–67.
Kim DH, Kang JG, Yanga SS, et al. A phytochrome-associated protein phosphatase 2A modulates light signals in flowering time control in Arabidopsis. Plant Cell 2002; 14:3043–56.
Ryu JS, Kim J, Kunkel T, et al. Phytochrome-specific type 5 phosphatase controls light signal flux by enhancing phytochrome stability and affinity for a signal tranducer. Cell 2005; 120:395–406.
Ni M, Tepperman JM, Quail PH . Binding of phytochrome B to its nuclear signaling partner PIF3 is reversibly induced by light. Nature 1999; 400:781–4.
Huq E, Al-Sady B, Hudson M, et al. Phytochrome-interacting factor 1 is a critical bHLH regulator of chlorophyll biosynthesis. Science 2004; 305:1937–41.
Liu XL, Covington MF, Fankhauser C, Chory J, Wagner DR . ELF3 encodes a circadian clock-regulated nuclear protein that functions in an Arabidopsis phyB signal transduction pathway. Plant Cell 2001; 13:1293–304.
Sweere U, Eichenberg K, Lohrmann J, et al. Interaction of the response regulator ARR4 with phytochrome B in modulating red light signaling. Science 2001; 294:1108–11.
Ahmad M, Jarrilo JA, Smirnova O, Cashmore AR . The CRY1 blue light photoreceptor of Arabidopsis interacts with phytochrome A in vitro. Mol Cell 1998; 1:939–48.
Mas P, Devlin PF, Panda S, Kay SA . Functional interaction of phytochrome B and cryptochrome 2. Nature 2000; 408:207–11.
Kang XJ, Chong J, Ni M . HYPERSENSITIVE TO RED AND BLUE 1, a ZZ-Type Zinc Finger Protein, regulates phytochrome B-mediated red and cryptochrome-mediated blue light responses. Plant Cell 2005; 17:822–35.
Ward JM, Cufr CA, Denzel MA, Neff MM . The Dof transcription factor OBP3 modulates phytochrome and cryptochrome signaling in Arabidopsis. Plant Cell 2005; 17:475–85.
Guo H, Mockler T, Duong H, Lin C . SUB1, an Arabidopsis Ca2+-binding protein involved in cryptochrome and phytochrome coaction. Science 2001; 291:487–90.
Duek PD, Fankhauser C . HFR1, a putative bHLH transcription factor, mediates both phytochrome A and cryptochrome signaling. Plant J 2003; 34:827–36
Park DH, Lim PO, Kim JS . The Arabidopsis COG1 gene encodes a Dof domain transcription factor and negatively regulates phytochrome signaling. Plant J 2003; 34:161–71.
Genoud T, Millar AJ, Nishizawa N, et al. An Arabidopsis mutant hypersensitive to red and far-red light signals. Plant Cell 1998; 10:889–904.
Cerdán PD, Chory J . Regulation of flowering time by light quality. Nature 2003; 423:881–5.
Kaczorowski KA, Quail PH . Arabidopsis PSEUDO-RESPONSE REGULATOR7 is a signaling intermediate in phytochrome-regulated seedling deetiolation and phasing of the circadian clock. Plant Cell 2003; 15:2654–65.
Osterlund MT, Hardtke CS, Wei N, Deng XW . Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 2000; 405:462–6.
Seo HS, Yang JY, Ishikawa M, et al. LAF1 ubiquitination by COP1 controls photomorphogenesis and is stimulated by SPA1. Nature 2003; 423:995–9.
Saijo Y, Sullivan JA, Wang H, et al. The COP1-SPA1 interaction defines a critical step in phytochrome A-mediated regulation of HY5 activity. Genes Dev 2003; 17:2642–7.
Holm M, Ma LG, Qu LJ, Deng XW . Two interacting bZIP proteins are direct targets of COP1-mediated control of light-dependent gene expression in Arabidopsis. Genes Dev 2002; 16:1247–59.
Seo HS, Watanabe E, Tokutomi S, Nagatani A, Chua NH . Photoreceptor ubiquitination by COP1 E3 ligase desensitizes phytochrome A signaling. Genes Dev 2004; 18:617–22.
Bauer D, Viczián A, Kircher S, et al. Constitutive photomorphogenesis 1 and multiple photoreceptors control degradation of phytochrome interacting factor 3, a transcription factor required for light signaling in Arabidopsis. Plant Cell 2004; 16:1433–45.
Duek PD, Mireille V, Elmer MV, van Oosten VR, Fankhauser C . The degradation of HFR1, a putative bHLH class transcription factor involved in light signaling, is regulated by phosphorylation and requires COP1. Curr Biol 2004; 14:2296–301.
Jang IC, Yang JY, Seo HS, Chua NH . HFR1 is targeted by COP1 E3 ligase for post-translational proteolysis during phytochrome A signaling. Genes Dev 2005; 19:593–602.
Yang J, Lin R, Sullivan J, et al. Light regulates COP1-mediated degradation of HFR1, a transcription factor essential for light signaling in Arabidopsis. Plant Cell 2005; 17:804–21.
Searle I, Coupland G . Induction of flowering by seasonal changes in photoperiod. EMBO J 2004; 23:1217–22.
Lin C . Photoreceptors and regulation of flowering time. Plant Physiol 2000; 123:39–50.
Mockler T, Yang HY, Yu XH, et al. Regulation of photoperiodic flowering by Arabidopsis phototeceptor. Proc Natl Acad Sci U S A 2003; 100:2140–5.
Tepperman JM, Zhu T, Chang HS, Wang X, Quail PH . Multiple transcription-factor genes are early targets of phytochrome A signaling. Proc Natl Acad Sci U S A 2001; 98:9437–42.
Putterill J, Robson F, Lee K, Simon R, Coupland G . The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to Zinc finger transcription factor. Cell 1995; 80:847–57.
Suárez-López P, Wheatley K, Robson F, et al. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 2001; 410:1116–20.
Valverde F, Mouradov A, Soppe W, et al. Photoreceptor regulation of CONSTANS protein in photoperiodic flowering. Science 2004; 303:1003–6.
Doyle MR, Davis SJ, Bastow RM, et al. The ELF4 gene controls circadian rhythms and flowering time in Arabidopsis thaliana. Nature 2002; 419:74–7.
Khanna R, Kikis EA, Quail PH . EARLY FLOWERING 4 functions in phytochrome B-regulated seedling de-etiolation. Plant Physiol 2003; 133:1530–38.
Park DH, Somers DE, Kim YS, et al. Control of circadian rhythms and photoperiodic flowering by the Arabidopsis GIGANTEA gene. Science 1999; 285:1579–82.
Fowler S, Lee K, Onouchi H, et al. GIGANTEA: a circadian clock-controlled gene that regulates photoperiodic flowering in Arabidopsis and encodes a protein with several possible membrane-spanning domains. EMBO J 1999; 18:4679–88.
Covington MF, Panda S, Liu XL, et al. ELF3 modulates resetting of the circadian clock in Arabidopsis. Plant Cell 2001; 3:1305–15.
Mas P, Alabadi D, Yanovsky MJ, Oyama T, Kay SA . Dual role of TOC1 in the control of circadian and photomorphogenic responses in Arabidopsis. Plant Cell 2003; 15:223–36.
Alabadi D, Oyama T, Yanovsky MJ, et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 2001; 293:880–3.
Wang ZY, Tobin E . Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 1998; 93:1207–17.
Schaffer R, Ramsay N, Samach A, et al. The late elongated hypocotyl mutation of Arabidopsis disrupts circadian rhythms and the photoperiodic control of flowering. Cell 1998; 93:1219–29.
Somers DE, Schultz TF, Milnamow M, Kay SA . ZEITLUPE encodes a novel clock-associated PAS protein from Arabidopsis. Cell 2000; 101:319–29.
Nelson DC, Lasswell J, Rogg LE, Cohen MA, Bartel B . FKF1, a clock-controlled gene that regulates the transition to flowering in Arabidopsis. Cell 2000; 101:331–40.
Acknowledgements
Our work on HRB1, RFI1, and RFI2 is supported in part by University of Minnesota start-up and Grant-in-Aid funds (to MinNI) and by grant from the National Research Initiative of the USDA Cooperative State Research, Education and Extension Service (2004-35304-14939 to Min NI).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
NI, M. Integration of light signaling with photoperiodic flowering and circadian rhythm. Cell Res 15, 559–566 (2005). https://doi.org/10.1038/sj.cr.7290325
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290325
Keywords
This article is cited by
-
Hierarchical graphical model reveals HFR1 bridging circadian rhythm and flower development in Arabidopsis thaliana
npj Systems Biology and Applications (2019)
-
Developmental transcriptome analysis of floral transition in Rosa odorata var. gigantea
Plant Molecular Biology (2018)
-
Comparative de novo transcriptome analysis of male and female Sea buckthorn
3 Biotech (2018)