Abstract
We studied the influence of extra-pair paternity on heritability estimates of morphological traits in a population of the Medium Ground Finch (Geospiza fortis) on Isla Daphne Major, Galápagos. Data from eight microsatellite loci were used to determine parentage. Six morphological traits measured on each finch were represented by two separate principal components analyses, one for the three bill measurements and one for the body size measurements. Heritabilities were calculated using weighted regressions of offspring on their parents and also offspring on their grandparents. We found that 20% of all offspring were extra-pair young but all offspring matched their mothers. Heritabilities derived from midparent–offspring regressions were all high and significantly different from zero. Removing all extra-pair young from the data set increased father–offspring regressions by an average of 21%, but mother–offspring resemblance still exceeded father–offspring resemblance by up to 42%. These results and grandparent–offspring regressions provide evidence for maternal effects, comparable in magnitude to those reported in other studies of wild birds.
Similar content being viewed by others
Introduction
Heritabilities of ecologically important traits are generally moderately high in a variety of organisms (e.g. Roff, 1997; Merilä & Sheldon, in press). This is evolutionarily important because it implies that the traits are responsive to selection. Heritabilities may even be higher than recorded, however, if undetected extra-pair mating causes a misidentification of paternity. Extra-pair paternity (EPP) occurs in a large number of socially monogamous bird species, yet the magnitude of the bias it introduces to estimates of heritabilities is generally unknown (but see Merilä et al., 1998). Alatalo et al. (1984) proposed that it could be measured indirectly as the difference in heritability estimates derived from separate father–offspring and mother–offspring regressions. Although some studies have found this to be useful (e.g. Møller & Birkhead, 1992), two studies (Hasselquist et al., 1995; Merilä et al., 1998) have yielded highly inaccurate estimates by this method.
Gauging the effects of EPPs on heritability estimates requires direct assessment of parentage from molecular genetic markers. Nevertheless, it is not straightforward for two reasons. First, the effects depend on the correlation between the trait value in the cuckolded male and the genetic father. If the trait values among them are positively correlated, for example as a result of female choice, then the effect of EPPs on h2 will be minimal. Second, the assumption of the comparative regressions method of no maternal effects may be wrong. Maternal effects are usually defined as effects of the maternal phenotype on offspring phenotype, above and beyond the contribution via nuclear genes (e.g. Kirkpatrick & Lande, 1989). Both EPPs and maternal effects can lead to higher mother–offspring than father–offspring regressions. The potential for maternal effects in birds is high because all bird species lay eggs, and the majority care for their young for an extended period of time (Price, 1998). Indeed, several maternal traits have been shown to affect offspring: maternal condition, laying date, clutch size, egg size and quality, incubation, and parental care (see Price, 1998; Lindström, 1999; for reviews).
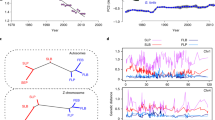
The purpose of the present study was to determine the influence of EPPs on heritability estimates of morphological traits in a population of Medium Ground Finches (Geospiza fortis) living on Isla Daphne Major in the Galápagos archipelago. This population has been the subject of a detailed, long-term study of natural selection and evolutionary response (e.g. Grant & Grant 2000 and references therein). Mother–offspring and father–offspring regressions of six morphological traits performed separately for seven cohorts of G. fortis born on Daphne Major show that mother–offspring regressions are on average higher than father–offspring regressions, but with considerable scatter (Fig. 1). This result matches those reported earlier with a smaller sample of cohorts (Boag, 1983; Grant & Grant, 1994; Grant & Grant, 2000). The present study aimed to determine to what degree these differences were attributable to (a) extra-pair paternity and (b) maternal effects. We used microsatellite DNA analyses to determine parentage and then estimated heritabilities of morphological traits based first on all data and then on the data set excluding extra-pair young (EPY). Because Fig. 1 suggested that heritability estimates from different cohorts may differ, we also investigated the possibility that pooling of data may have affected our results (cf. Hõrak & Tammaru, 1996).
A comparison of h2 estimates derived from father–offspring vs. mother–offspring regressions. h2 estimates were calculated separately for six morphological traits (body weight, tarsus length, wing length, bill depth, bill width, and bill length) in seven cohorts of Geospiza fortis (1976, 1978, 1981, 1983, 1984, 1987, and 1991). Each data point represents the two h2 estimates (father–offspring and mother–offspring) for a particular trait and cohort. Different symbols were used for the different cohorts. Only cohorts for which we had data on at least 20 families were included. The straight line corresponds to the line of equal h2 estimates. If there were no systematic difference between father–offspring and mother–offspring regressions, the points would lie scattered equally around this line.
Materials and methods
Earlier publications describe the species, its habitat, and the study methods in detail (e.g. Abbott et al., 1977; Boag & Grant, 1984a, b; Grant, 1999) and we provide only a brief summary here. The small (0.34 km2) island of Daphne Major lies in the centre of the Galápagos archipelago, Ecuador, approx. 7.5 km distant from the large island of Santa Cruz. The medium ground finch (Geospiza fortis) and populations of three other congeneric finch species that breed on Daphne were studied intensively between 1976 and 1992, and less thoroughly in succeeding years. Over the period 1976–92 the harmonic mean population size of G. fortis was 198 (Grant & Grant, 1995).
Breeding occurs in response to rain. The onset of rain varies considerably from year to year but rain tends to fall sporadically during the months of December–June (e.g. Grant et al., 2000). Most nests were located in each year in which breeding ocurred, and nestlings were banded with unique combinations of one numbered metal band and three coloured plastic bands. We identified the putative parents by observing them build the nest, incubate the eggs, and feed the nestlings. Thus, we were able to relate the measurements of the young to those of their social parents.
Darwin’s Finches cannot be sexed reliably in the field until they are adults. Thus, we were only able to assign sexes to finches that survived to adulthood.
The families included in this data set come from six different cohorts. The representation of the different cohorts was uneven, partly because of vastly different reproduction in the different years (1987 and 1991 were El Niño years with unusually high reproduction (e.g. Grant et al., 2000)) and partly because blood sampling effort varied over time. Our data set included young from the following cohorts (sample sizes in parentheses): 1986 (1), 1987 (81), 1990 (1), 1991 (114), 1992 (9), and 1993 (18).
Extrapair paternity
Collection of blood samples began in 1987. A single drop of blood from the brachial vein was taken from eight-day-old nestlings or from adults and immatures captured in mist nets (see Petren et al., 1999 for a more detailed description of the blood sampling technique). We determined the parentage of all those G. fortis families for which we had blood samples and morphological measurements from the father, the mother, and at least one offspring, totalling 223 offspring, 77 fathers, and 76 mothers in 93 families. Parentage was determined by genotyping all these birds at eight microsatellite loci. Samples from young that mismatched their father at only a few loci, were genotyped again to exclude the possibility of genotyping error leading to a false paternity exclusion. All but one extrapair young mismatched their fathers at two or more loci. It is possible that this one young’s single locus mismatch was caused by a mutation. However, because extrapair paternities are much more likely than mutations, we considered this particular bird to be an extrapair young. The details of the genotyping techniques as well as primer sequences are given in Petren (1998). We used the following loci for our analyses here: GF1, GF2, GF3, GF7, GF8, GF9, GF11, and GF16.
Morphology
Adult and immature finches were trapped in mist nets each year, generally at the end of the dry (nonbreeding) season. Geospiza fortis are not fully grown until approximately eight weeks of age (Boag, 1984), therefore measurements were only used if the birds were older than 60 days. The following six measurements were taken from each finch: weight (in grams), wing length, tarsus length, bill length (BL), bill depth (BD), and bill width (BW; all in mm); see Grant (1999) for details of measurement techniques. Repeated measurements of the same bird were averaged. More than 65% of all measurements were made by P. R. G. Correction factors calculated from birds measured by four other, trained observers were applied to all other measurements (Grant & Grant, 1994). Repeatabilities for all traits were high (Boag, 1983; Price & Grant, 1984; Grant & Grant, 1994). Because of seasonal variation and moulting, weight and wing length had the lowest repeatabilities (0.83 and 0.79 respectively), whereas bill measurements had the highest repeatabilities (all ≥ 0.93; Grant & Grant, 1994). To achieve appropriate scaling, we used the cube-root of weight in our analyses, and all measurements were ln-transformed before analysis to stabilize variances.
Since all six morphological traits were correlated with each other (0.29 ≤ r ≤ 0.83, all P ≤ 0.0001) we used principal component analyses (PCAs) to represent the linear trends in the correlated data set along fewer, uncorrelated axes. We used two separate PCAs: one for the bill measurements only (BD, BW, BL) and one for the body size measurements (weight, tarsus, wing). In both cases, the first two principal components (PCs) were extracted from the covariance matrix. For the PCA of the bill measurements (PC-bill), PC1 and PC2 explained 84% and 11% of the total variance, respectively. For the PCA of the body measurements (PC-body), PC1 and PC2 explained 63% and 20% of the total variance, respectively. In both cases therefore the first two PCs captured most of the total variance in the traits.
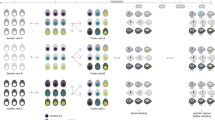
A graphical representation of the loading of each original variable onto PC1 and PC2 for the two separate PCAs is shown in Fig. 2. PC1 can be considered a measure of bill size (PC-bill, Fig. 2a) and body size (PC-body, Fig. 2b), respectively, with all variables loading positively and approximately equally. Large PC1 scores represent large birds. PC2-bill is a measure of bill shape, with bill length loading positively, and bill depth and bill width loading negatively. High PC2-bill scores indicate birds with long, narrow bills. PC2-bill primarily captures the aspect of bill shape that is also measured by the ratio of bill length to bill depth as reflected in the very high correlation between this ratio and PC2-bill (r=0.92). PC2-body is an index of body shape, with high values indicating heavy, large winged birds with short tarsi. PC1-bill and PC1-body were correlated with r=0.60 (P=0.0001) indicating that body size and bill size covary to some degree. However, less than 40% of the variance in one PC1 was explained with the other PC1, and we kept both PC1-bill and PC1-body in our analyses. PC2-bill and PC2-body were uncorrelated (r=0.05, P=0.4).
Correlations of the original traits with the first two principal components derived from two separate principal components analyses (PCA) in Geospiza fortis. (A) PCA of the three bill measurements (BD, BW, BL, see text for abbreviations). (B) PCA of the three body size measurements. Vectors indicate the direction of maximum variation for each character within the principal component plane. The mapping of each vector onto PC1 and PC2, respectively, represents the factor loading of that variable onto PC1 and PC2. Note that all correlations are positive and approximately equal for PC1, but of different sign for PC2.
One male G. fortis and his daughter were excluded from the analyses, because he represented a serious outlier. Most of his morphological measurements were three to four standard deviations above the population mean, and his body mass was eight standard deviations larger than the population mean. He was either an immigrant from the neighbouring island of St. Cruz where G. fortis are considerably larger (e.g. Grant & Grant, 1996), or a hybrid. Inclusion of this single family with a single offspring would have increased the h2 estimates by up to 13% in some cases.
Heritability estimates
Heritabilities (h2) were calculated using regression analyses: midparent–offspring, single parent–offspring, single parent–single sex offspring (e.g. mother–daughter), and grandparent–offspring. In each analysis, measurements of offspring of the same parent(s) were averaged (see below). Midparent–offspring regressions estimate heritability directly, whereas single parent–offspring regression estimate half of the heritability, and grandparent–offspring regressions estimate one quarter of the heritability (Lynch & Walsh, 1998). Slopes and associated standard errors from the last two regressions were multiplied by two and four, respectively, to obtain h2 estimates.
The number of offspring per family (family size) varied between 1 and 9 (mean=2.5) in our data set. To minimize sampling error of the heritability estimates we therefore used weighted least-square regressions following Lynch & Walsh (1998; pp. 539–542). Each observation was weighted by the inverse of the residual sampling variances of family means about the parent–offspring regression. Thus, the weight of the ith family is:

where ni is the size of family i, and t is the intraclass correlation between sibs. B is half the regression slope for midparent regressions, and equals the regression slope for single parent or grandparent regressions, respectively. Since B is a function of the regression coefficient itself, iterative reweighting procedures were required. We used PROC NLIN in SAS (SAS, 1990) with the _WEIGHT_ statement to perform the iteratively reweighted regressions. Intraclass correlation coefficients were estimated following Sokal & Rohlf (1981); p. 216.
Assortative mating does not affect midparent–offspring regressions, but it increases the regression of offspring on single parents by a factor (1 + r), where r is the phenotypic correlation between mates (Falconer & Mackay, 1996). We found no evidence for assortative mating in our data sets (all r ≤ 0.11, P > 0.33, N=92), except for PC2-bill in the data set of young born in 1991 to parents born in 1987 (r=−0.53, P=0.006, n=25). Hence, we adjusted the heritability estimates for PC2-bill in that data set accordingly.
Different trait means and variances in males and females are a further source of bias in estimates of h2. In our data set, males were on average significantly larger than females in all morphological traits, except PC2-body (Table 1). The variances of the original morphological characters were similar in males and females, although the variance of PC2-bill in males was significantly higher than in females (Table 1). Therefore, we estimated heritabilities with the variables standardized to a mean of zero and a variance of unity for each sex separately before analysis (Lynch & Walsh, 1998). Since we were unable to determine the sex of offspring that did not survive to adulthood, we omitted those individuals from the analyses (12 birds).
In the presence of maternal effects mother–offspring regressions differ from father–offspring regressions. Specifically, let σ2Ao stand for the variance due to direct additive genetic effects, σ2Am for the variance due to maternal (indirect) additive genetic effects, σAo,Am for the covariance between direct and maternal additive genetic effects, σDo,Dm for the covariance between direct and maternal (indirect) dominance effects, σ2Em for the variance due to maternal environmental effects, σEo,Em for the covariance between direct and indirect (maternal) environmental effects, and σ2z for the phenotypic variance; then the difference between the mother–offspring and father–offspring regressions is (Lynch & Walsh, 1998; p. 708):

Thus, the difference m between mother–offspring and father–offspring regressions includes terms for the three covariances above, and for the maternal genetic and maternal environmental variances (see Lynch & Walsh, 1998, chapter 23 for details and assumptions).
The six morphological traits measured in the medium ground finch were all correlated (range: r=0.29 to r=0.83). To avoid problems of non-independence and inflated type-I error rates, we restricted our statistical testing of heritability estimates to PC1s and PC2s. However, to ensure that our results are readily comparable to other studies, we also give the h2 estimates for the original traits. Differences in heritability estimates expressed in percentage were calculated as (h2larger − h2smaller)/h2larger. All statistical tests were two-tailed.
Unless very large sample sizes are available, heritability estimates generally have large standard errors (e.g. Falconer & Mackay, 1996; Lynch & Walsh, 1998), and as a result comparisons of heritability estimates often have low statistical power. We present 95% confidence intervals with all estimates of differences between h2 estimates. Confidence intervals of differences allow an assessment of post-hoc power because values within the confidence interval represent hypotheses that are consistent (cannot be rejected) with the data.
Heritability estimates based on midparent–offspring and single parent–offspring regressions were calculated using the entire data set and re-calculated after removing all EPY. Sex-specific and grandparent–offspring regressions were only performed with the data set that excluded all EPY. Sample sizes for the various analyses vary due to the fact that (1) a few birds were mated to more than one partner during the study period and thus enter the midparent–offspring analyses more than once, while they are only represented once in the single parent–offspring regressions; and (2) because not all parents had both sons and daughters represented in the analyses.
Results
Extrapair paternity
The combined exclusion probability (Weir, 1990; p. 187) of the eight microsatellite loci was 99.96%. It is unlikely therefore that we failed to exclude a father if in fact he did not sire an offspring. Forty-four offspring from 33 different families did not match their father at one or more loci, suggesting that they were extrapair young (EPY). Thus, the frequency of EPY was 19.7% (44/223), and 35.5% (33/93) of families had at least one EPY. All offspring matched their mothers at all loci, indicating that intraspecific brood parasitism is either absent or very rare in this population. Therefore, we assume in the following that grandmothers were always correctly identified even if we did not have genotypic data.
Effects of extrapair paternity on heritability estimates
For the entire data set including EPY, the heritability estimates derived from midparent–offspring regressions of all morphological traits and of PC1s and PC2s ranged from 0.43 (tarsus, Table 2) to 0.83 (PC2-bill, Table 3). All midparent–offspring heritabilities were statistically different from zero. With the exception of wing length and both PC2s, the mother–offspring heritability estimates were higher than those derived from father–offspring regressions, often considerably higher (Tables 2 and 3). For PC1-bill and PC1-body these differences were significant and amounted to 41% (95% CI: 8% to 73%, t148=2.47, P=0.014), and 59% (95% CI: 18% to 100%, t148=2.86, P=0.005), respectively. For PC2-bill and PC2-body father–offspring resemblance exceeded that of mothers and their offspring by 13% (95% CI: −29% to 54%) and 22% (95% CI: −33% to 77%) but not significantly so (all P > 0.43).
Given the EPP rate of ∼20%, we would expect father–offspring heritabilities to increase by a similar amount after excluding EPY, whereas mother–offspring resemblance should not change (Alatalo et al., 1989). Excluding all EPY increased father–offspring resemblance for all traits by an average of 25%. For PC1-bill the increase amounted to 29% (95%CI: −8% to 66%) and for PC1-body it was 34% (95% CI: −28% to 96%; Table 3). For the two shape factors the figures were 11% (95% CI: −24% to 46%) for PC2-bill and 10% (95% CI: −37% to 58%) for PC2-body. On average, these increases (21%) were quite close to the expected 20%. Note, however, the large confidence intervals around the changes in h2, and that none of them was statistically significant (all P > 0.12).
As expected, mother–offspring resemblances were relatively unaffected by exclusion of EPY, and no significant differences were observed. Heritabilities of the original traits ostensibly declined, though only by 6% on average. The two PC1 traits declined on average by 7% and the two PC2 traits declined on average by 1% (Table 3).
We were able to identify the genetic father unequivocally for 16 of the EPY in our sample. This allows us to test our prediction that the difference in h2 estimated from father–offspring regressions with and without EPY is a function of the correlation between the trait values of the genetic fathers and the cuckolded males. A comparison of the four PC traits of the genetic fathers and the corresponding cuckolded males revealed a significant negative correlation between the body size measures of the two ‘fathers’ of an EPY (PC1-body: r=−0.52, P=0.04, n=16), and a nonsignificant correlation of r=−0.3 (P=0.27, n=16) for bill size. Both shape factors were uncorrelated between the two males (r=0.01 and r=−0.04 for PC2-bill and PC2-body, respectively; all P > 0.88). As predicted, we find a very strong negative correlation between the correlation coefficients of the trait values of the males and the percentage difference in h2 estimates (r=−0.96, P=0.04, n=4). Therefore, the difference in father–offspring resemblance caused by EPPs is more pronounced when the genetic fathers and the cuckolded males are on average morphologically different than when their morphologies are uncorrelated.
In summary, we found that for most traits mother–offspring regressions exceeded father–offspring regressions because of EPYs. The magnitude of the effect of EPYs on h2 estimates was a function of the morphological resemblance between the cuckolded male and the genetic father.
Effects of pooling years on heritability estimates
To investigate the potentially confounding effect of pooling data from different cohorts on the h2 estimates, we repeated the analyses using only data from young born in one particular cohort to parents born in another cohort. If cohorts are heterogeneous, then a single-cohort analysis should give different results from the overall analysis. We had sufficient samples sizes for only one such cohort analysis: young born in 1991 to parents born in 1987. The EPY rate in this restricted data set was almost identical to the entire data set: 19% of young and 32% of families. Heritability estimates derived from this unpooled data set (Table 4) gave results that were qualitatively similar to the ones from the entire data set. The most important difference is that here, unlike in the combined data, mother–offspring regressions significantly exceeded father–offspring regressions by 41% for PC1-bill after removing all EPY (95% CI: 6% to 76%, t48=2.35, P = 0.023). This difference provides evidence of some heterogeneity among cohorts in parent-specific regressions.
Maternal effects
After removing all EPY from the analyses, mother–offspring resemblance still exceeded father–offspring resemblance at PC1-bill and PC1-body by 19% and 23%, respectively (Table 3). The opposite held for the PC2s: father–offspring regressions exceeded mother–offspring regressions by 35% (bill shape) and 21% (body shape). None of these differences are statistically significant (all P > 0.33 for PC1s, and all P > 0.07 for PC2s). However, as shown above, mother–offspring regressions for PC1-bill significantly exceeded father–offspring regressions by 41% in the single-cohort analysis (Table 4). Thus, there is statistical evidence for maternal effects after EPY are removed in the single-cohort data set.
Maternal effects are expected to lead to higher maternal grandparent–offspring than paternal grandparent–offspring regressions (Lynch & Walsh, 1998; p. 691). We only had sufficient sample sizes for the grandmother–offspring regressions. The two grandmother–offspring regressions gave similar results for all variables excepting PC1-body (Table 5). Both paternal and maternal grandmother–offspring regressions resulted in significant heritability estimates for PC1-bill, PC2-bill and PC2-body (Table 5). h2 of PC1-body based on the paternal grandmother–offspring regressions, however, is not statistically different from zero. The difference in h2 for PC1-body between maternal and paternal grandmother regressions was very large (94%, 95% CI: 66% to 123%) and highly significant (t79=6.5, P < 0.0001). Thus, the grandparent regressions suggest the presence of maternal effects in body size, and the single parent–offspring regressions in the single-cohort data set suggest the presence of maternal effects in bill size.
Other causes of higher mother–offspring resemblance
Sex-linked genes or sex-limited expression of genes are another potential cause of differences in heritability estimates (e.g. Roff, 1997). To investigate this possibility, we calculated the sex-specific heritabilities, i.e. father–son, father–daughter, mother–son, and mother–daughter regressions (Table 6). All father–son, father–daughter, and mother–daughter regressions yielded significant estimates of heritabilities for PC1 and PC2. In mother–son comparisons, however, bill shape and body shape did not show significant heritable variation. For PC1-bill, the mother–daughter regression yielded an estimate of h2 that was 36% (95% CI: 5% to 67%) higher than the father–son regression, a difference that was statistically significant (t102=2.3, P=0.024). Similarly, for PC2-body, the mother–son comparison was 60% (95% CI: 15% to 104%) lower than the mother–daughter regression (t101=2.6, P=0.01). None of the other comparisons yielded significant differences in h2 estimates. Overall therefore there is evidence for differences in sex-specific heritabilities in two of the traits in our data set.
Discussion
The main results of this study are, first heritabilities of the measured and synthetic traits are very high, second a 20% EPP frequency causes a substantial underestimate of heritabilities from father–offspring regressions, and third there is equivocal evidence for maternal effects from comparisons of single parent–offspring and grandparent–offspring regressions.
Heritabilities of morphological traits
The heritability estimates based on midparent–offspring regressions with the entire data set were in general agreement with estimates derived previously from this population (e.g. Boag & Grant, 1978; Boag, 1983; Grant & Grant, 1994, 2000). Removing all extra-pair young increased the h2 estimates of bill size and shape to 0.85 and 0.88, respectively (Table 3). Therefore, our data suggest that more than 85% of the phenotypic variation in these morphological traits is attributable to additive genetic variation. This is a distinctly larger proportion than found in many other studies (e.g. Boag & van Noordwijk, 1987; Merilä & Sheldon, in press), but differences in sample sizes and study design make comparisons across studies difficult. Higher h2 estimates can result from either increased additive genetic variation or decreased phenotypic variance. Darwin’s Finches have been a focus of evolutionary biologists precisely because they exhibit large levels of phenotypic variance (e.g. Grant, 1999). Therefore, low phenotypic variances are not responsible for the relatively high h2 estimates. Relatively high levels of additive genetic variation have been attributed to rare but persistent introgressive hybridization (Grant & Grant, 2000). Since this is a non-experimental study we cannot rule out the possibility that our heritability estimates are inflated by common environment effects. However, regressions of cuckolded fathers on their extrapair young were all small and nonsignificant (all h2 < 0.2, all P > 0.15), indicating little or no inflation from common environmental effects (see also Grant & Grant, 2000).
Body size and shape had heritabilities of 0.56 and 0.45, respectively (Table 3). These values are 33% (95% CI: −12% to 78%, t156=1.46, P > 0.14) and 49% (95% CI: 1% to 96%, t156=2.02, P=0.045) lower than the corresponding h2 estimates for bill size and shape. The reasons for the lower h2 estimates are to be found in the lower repeatabilities of tarsus length, wing length, and body mass as compared to the bill measurements, and in the lower effects of introgressive hybridization on the body size traits (Grant & Grant, 1994). The lower repeatabilities are due both to inherently higher measurement errors (tarsus length) and higher variation within an individual over time (wing and body mass).
Heritability estimates are based on several assumptions, which, if not met, may bias the resulting estimates considerably. All standard texts in quantitative genetics (e.g. Falconer & Mackay, 1996; Lynch & Walsh, 1998) discuss the various assumptions involved. We will emphazise just one of them here: heritability estimates are specific to the population and the environment in which they were estimated. Consequently, some studies have found heritability estimates to be substantially different in different environments (e.g. Gebhardt-Henrich & van Noordwijk, 1991; Smith & Wettermark, 1995; Merilä, 1997; Merilä & Fry, 1998; Kunz & Ekman 2000) although this is not a universal outcome (see Merilä & Sheldon (in press) for a detailed discussion). When environmental conditions vary among years, parents will have experienced different growth conditions from those of their offspring. Under these circumstances, pooling data from more than one year can lead to substantial variation in heritability estimates, with a downward bias being approximately twice as likely as an upward bias (cf. Hõrak & Tammaru, 1996). By comparing the results obtained from our pooled data set (Table 3) with the results from the single-cohort analysis (Table 4), we found evidence for weak heterogeneity among cohorts in h2 estimates derived from singleparent–offspring regressions, but midparent–offspring regressions are remarkably similar.
Effects of misidentified paternity
In an earlier study of the 1978 cohort, Boag (1983) assumed paternity was incorrectly identified when females changed mates. By excluding those affected families he obtained a substantial increase in h2 estimates. Our study shows the potential magnitude of the effects of misidentified paternity. As expected, known EPPs resulted in smaller heritability estimates from father–offspring regressions than from mother–offspring comparisons, significantly so for beak size and body size (PC1-bill and PC1-body; Table 3). Removing all EPY from the data set consequently led to an increase in heritability estimates, by up to 33%. A few points are noteworthy. First, although not significantly so, EPPs can have an effect on estimates of heritability even from midparent–offspring regressions (Tables 2 and 3): excluding all EPY increased h2 derived from midparent–offspring regression by 21% (95% CI: −21% to 64%) for PC1-bill. Second, the results obtained from different, uncorrelated traits can vary substantially: EPPs caused mother–offspring and father–offspring resemblance in PC1-body to differ by 59% but father–offspring resemblance exceeded mother–offspring resemblance for bill and body shape. Similarly, removing all EPYs resulted in a 33% increase in father–offspring comparisons for body size, but only a 10% increase for body shape. However, the average of all four PC traits (21%) is remarkably close to the expected 20%. Third, although the average effect of EPYs on heritability estimates was quite close to the expected effect, the difference between mother–offspring and father–offspring resemblance for bill and body size calculated from the entire data set would have overestimated the true EPP rate by more than a factor of two. Many avian studies that have employed heritability analyses to estimate EPP rates used nestling tarsus length in their analyses (e.g. Alatalo et al., 1984, 1989; Lifjeld & Slagsvold, 1989; Norris & Blakey, 1989; Hasselquist et al., 1995) because it is virtually fully grown before fledging. In our population, using tarsus length would have led to an estimate of EPP rates of nearly 58%, almost three times the actual rate. Thus, our findings lend further support to the view that differences in heritabilities are not a reliable means of estimating EPP rates (e.g. Hasselquist et al., 1995; Merilä et al., 1998).
The prediction underlying the use of heritabilities to estimate EPP rates, namely that an EPP rate of, for example, 20% will lead to a difference in h2 estimates of 20% (Alatalo et al., 1984) depends on the assumption that the trait values of the genetic fathers and those of the cuckolded males are uncorrelated. This may not be correct. If they are positively or negatively correlated, as a result of random sampling or female choice, then the effects of EPPs on h2 will be either less or more pronounced than expected. Our results illustrate this point. The difference caused by EPPs in h2 estimated from father–offspring regressions was tightly correlated with the correlations between the morphology of the genetic fathers and the cuckolded males. The more the two males differed on average (negative correlation), the larger the difference in h2. In fact, more than 92% of the variation between the four PC traits in differences in father–offspring resemblance caused by EPPs is accounted for by the degree to which the trait values are correlated between genetic fathers and cuckolded males. The effects of EPPs on h2 estimates therefore are not only a function of EPP rates, but also of the (dis-) similarity of trait values of the genetic fathers and the cuckolded males.
Finally, heritability estimates are notoriously inaccurate unless large sample sizes are available (e.g. Falconer & Mackay, 1996; Lynch & Walsh, 1998). Since sample sizes from natural populations, even from long-term studies such as the present one, are generally at best moderate, low statistical power is almost inevitable. In our data this is reflected in the large confidence intervals for the observed differences between h2 estimates. For example, the 95% confidence interval for the difference in estimates of h2 for PC1-body between father–offspring and mother–offspring regressions ranged from 18 to 100%. This means that differences in h2 as small as 18% and as large as 100% are consistent with our data and statistically indistinguishable. The fact that EPPs led to a statistically significant difference between mother–offspring and father–offspring regressions for both PC1s underlines the fact that EPPs can have a large effect on heritability estimates.
Maternal effects
Studies in the past have led to the view that maternal effects on structural size in birds diminish throughout life and are no longer detectable when adult size is reached (e.g. Price, 1998). Nevertheless, there appears to be mounting evidence for enduring maternal effects on some size traits in birds. Higher mother–offspring than father–offspring resemblances have recently been reported for a structural size trait, tarsus length, in barnacle geese (Branta leucopsis, Larsson & Forslund, 1992), pied flycatchers (Ficedula hypoleuca, Potti & Merino, 1994), great reed warblers (Acrocephalus arundinaceus, Hasselquist et al., 1995) and collared flycatchers (Ficedula albicollis, Merilä et al., 1998).
There are three indications of maternal effects in our data. First, apparent, though nonsignificant, differences between mother–offspring and father–offspring resemblances remain in the pooled data set after removal of EPY. Second, mother–offspring regressions in the single-cohort analysis (Table 4) give significantly higher h2 estimates for bill size after EPY were removed than do the father–offspring regressions. Third, maternal grandmother–offspring regressions for PC1-body were significantly higher than paternal grandmother–offspring regressions. Taken at face value, these results indicate maternal effects approximately equal in magnitude to those in the studies cited above. For example, we calculate the difference between mother–offspring and father–offspring regressions (m, eqn 2) in tarsus length of birds reported by Merilä et al. (1998), Potti & Merino, 1994), and Hasselquist et al. (1995) to be 0.08, 0.155, and 0.295, respectively. In our study, values of m for the combined data set are 0.09 for bill size and 0.08 for body size. In the single-cohort data the corresponding values of m are 0.28 and 0.11.
The strongest evidence of maternal effects is provided by the single-cohort analysis for bill size (Table 4). The evidence from grandparent regressions involves a different trait (body size), and is difficult to reconcile with the fact that h2 estimates for body size show little difference between mother–offspring and father–offspring regressions (see Lynch & Walsh, 1998; p. 691). There is no grandparental evidence of maternal effects in bill size, unless the differences between mother–offspring and father–offspring regressions were caused primarily by positive covariances between direct dominance and maternal dominance effects (σDo,Dm, compare eqn 2 and table 23.1 in Lynch & Walsh (1998)). This is unlikely because Grant & Grant (1994) found evidence for only weak dominance effects in G. fortis and these contributed little to phenotypic variation.
In spite of these indications of maternal effects, we consider three alternative explanations for the differences between mother–offspring and father–offspring regressions: sex-linkage or limitation, ecological processes, and statistical biases. Sex-linked inheritance or sex-limited expression of genes might affect body and bill size, as reported for chickens (e.g. Chambers, 1990; Barbato & Vasilatos-Younken, 1991). However, sex-specific heritabilites in our data set (Table 6) clearly do not support X-linked inheritance. Under X-linked inheritance we would expect the sex-specific regressions to be ordered in the following way: XX parent/XY offspring = XY parent/XX offspring ≥ XX parent/XX offspring > XY parent/XY offspring (Mather & Jinks, 1982). In birds, where females are the heterogametic sex, we would therefore expect the mother–daughter regression to be the lowest. In G. fortis, however, mother–daughter regressions are the highest for three of the four PC traits (Table 6). Our observations do not match two expectations with Y-linked inheritance (a) father–daughter regressions should be lower than father–son regressions (Table 6), and (b) intraclass correlations (t) should be lower among sons than among daughters, since the latter share the same Y-chromosome (data not shown). Nevertheless, the patterns of sex-specific heritabilities in our data are sufficiently pronounced to merit further study (see also Merilä & Gustafsson, 1993).
It is unlikely that ecological and measurement biases, such as different ages at measurement combined with subsequent growth, would have led to the observed differences between mother–offspring and father–offspring regressions. First, almost all medium ground finches included in the present analyses were measured at the same time in their first year of life; only four males were measured at an older age. Second, although we found evidence for linear change in adult G. fortis for bill size with age and a pattern of first increase and then decline for bill shape, there was no difference between the sexes (ANCOVA, PC1-bill: F=0.7, P > 0.4; PC2-bill: F=0.1, P > 0.7), nor were sex differences found for body size or shape (ANCOVA, all F < 0.87, P > 0.35). Since this is a nonexperimental study, higher environmental correlations between mother–daughter and father–daughter than mother–son and father–son might have inflated our estimates of heritability. But this too is unlikely. Regressions of extrapair young on their social fathers were all of small magnitude (range: r=−0.36 to r=0.20) and not significant (all P > 0.15), making it unlikely that strong environmental correlations were responsible for the observed pattern.
If selection occurred between the age at measurement and the age of reproduction, we would expect the variances of the parents to be different from those of the offspring. We found that the variances in PC1 and PC2 were often higher among the parents than among the offspring, albeit not statistically significantly so (Levene’s tests, all P > 0.1). Distributions that exhibit substantial skewness or kurtosis may also bias heritability estimates in ways that are not understood. Fortunately these potential biases were generally minimal in our data set, although skewness and kurtosis were pronounced in the sample of grandparents.
We conclude that there may be some maternal effects on morphological traits of adults. A more comprehensive study with greater statistical power and greater control of potentially confounding variables is needed.
Estimating h2 in the field when paternity is uncertain
It is often impractical to assess paternity by molecular methods. In such instances, where misidentified paternity is suspected, what is the best method for estimating heritabilities? One obvious candidate is mother–offspring regression. This has two disadvantages: potential inflation by maternal effects of unknown magnitude and high standard errors. Midparent–offspring regressions suffer these disadvantages to a lesser degree, so unless EPP is very frequent this form of regression analysis is probably the best to use. Full-sib and maternal half-sib analyses are both susceptible to inflation by maternal effects. Only paternal half-sib analyses are not affected by maternal effects, but they require molecular determination of parentage. We conclude that all three standard methods of regression and correlation should be undertaken and reported, with greatest weight being given in most circumstances to midparent–offspring regression.
References
Abbott, I., Abbott, L. K. and Grant, P. R. (1977). Comparative ecology of Galapagos Ground Finches (Geospiza Gould): evaluation of the importance of floristic diversity and interspecific competition. Ecol Monogr, 47: 151–184.
Alatalo, R. V., Gustafsson, L. and Lundberg, A. (1984). High frequency of cuckoldry in pied and collared flycatchers. Oikos, 42: 41–47.
Alatalo, R. V., Gustafsson, L. and Lundberg, A. (1989). Extra-pair paternity and heritability estimates of tarsus length in pied and collared flycatchers. Oikos, 56: 54–58.
Barbato, G. F. and Vasilatos-Younken, R. (1991). Sex-linked and maternal effects on growth in chickens. Poultry Sci, 70: 709–718.
Boag, P. T. (1983). The heritability of external morphology in Darwins ground finches (Geospiza) on Isla Daphne Major, Galapagos (Ecuador). Evolution, 37: 877–894.
Boag, P. T. (1984). Growth and allometry of external morphology in Darwins finches (Geospiza) on Isla Daphne Major, Galapagos (Ecuador). J Zool Lond, 204: 413–442.
Boag, P. T. and Grant, P. R. (1978). Heritability of external morphology in Darwin’s finches. Nature, 274: 793–794.
Boag, P. T. and Grant, P. R. (1984a). The classical case of character release: Darwins finches (Geospiza) on Isla Daphne Major, Galapagos. Biol J Linn Soc, 22: 243–287.
Boag, P. T. and Grant, P. R. (1984b). Darwin’s Finches (Geospiza) on Isla Daphne Major, Galapagos: breeding and feeding ecology in a climatically variable environment. Ecol Monogr, 54: 463–489.
Boag, P. T. and van Noordwijk, A. J. (1987). Quantitative genetics. In: Cooke, F. and Buckley, P. A. (eds) Avian Genetics. Academic Press, London.
Chambers, J. R. (1990). Genetics of growth and meat production in chickens. In: Crawford, R. D. (ed.) Poultry Breeding and Genetics, pp. 599–643. Elsevier, Amsterdam.
Falconer, D. S. and Mackay, T. F. C. (1996). Introduction to Quantitative Genetics, 4th edn. Longman, Essex.
Gebhardt-Henrich, S. G. and van Noordwijk, A. J. (1991). Nestling growth in the Great Tit: I. Heritability estimates under different environmental conditions. J Evol Biol, 4: 341–362.
Grant, P. R. (1999). Ecology and Evolution of Darwin’s Finches, 2nd edn., Princeton University Press, Princeton, NJ.
Grant, P. R. and Grant, B. R. (1994). Phenotypic and genetic effects of hybridization in Darwin’s finches. Evolution, 48: 297–316.
Grant, P. R. and Grant, B. R. (1995). Predicting microevolutionary responses to directional selection on heritable variation. Evolution, 49: 241–251.
Grant, P. R. and Grant, B. R. (1996). Finch communities in a climatically fluctuating environment. In: Cody, M. L. and Smallwood, J. A. (eds) Long-Term Studies of Vertebrate Communities, pp. 343–390. Academic Press, San Diego.
Grant, P. R. and Grant, B. R. (2000). Quantitative genetic variation in populations of Darwin’s Finches. In: Mousseau, T. A., Sinervo, B. and Endler, J. A. (eds) Adaptive Genetic Variation in the Wild, pp. 3–40. Oxford University Press, Oxford.
Grant, P. R., Grant, B. R., Keller, L. F. and Petren, K. (2000). Effects of El Niño events on Darwin’s finch reproduction. Ecology, 81: 2442–2457.
Hasselquist, D., Bensch, S. and Von, S. T. (1995). Estimating cuckoldry in birds: The heritability method and DNA fingerprinting give different results. Oikos, 72: 173–178.
Hõrak, P. and Tammaru, T. (1996). Between-year variation in breeding conditions biases heritability estimates for body size in birds. Ardea, 84: 127–135.
Kirkpatrick, M. and Lande, R. (1989). The evolution of maternal characters. Evolution, 43: 485–503.
Kunz, C. and Ekman, J. (2000). Genetic and environmental components of growth in nestling blue tit (Parus caeruleus). J Evol Biol, 13: 199–212.
Larsson, K. and Forslund, P. (1992). Genetic and social inheritance of body and egg size in the barnacle goose (Branta leucopsis). Evolution, 46: 235–244.
Lifjeld, J. T. and Slagsvold, T. (1989). How frequent is cuckoldry in pied flycatchers Ficedula hypoleuca? Problems with the use of heritability estimates of tarsus length. Oikos, 54: 205–210.
Lindström, J. (1999). Early development and fitness in birds and mammals. Trends Evol Ecol, 14: 343–347.
Lynch, M. and Walsh, B. (1998). Genetics and Analysis of Quantitative Traits. Sinauer Associates, Sunderland, MA.
Mather, K. and Jinks, J. L. (1982). Biometrical Genetics, 3rd edn. Chapman & Hall, London.
Merilä, J. (1997). Expression of genetic variation in body size of the collared flycatcher under different environmental conditions. Evolution, 51: 526–536.
Merilä, J. and Fry, J. D. (1998). Genetic variation and causes of genotype–environment interaction in the body size of blue tit (Parus caeruleus). Genetics, 148: 1233–1244.
Merilä, J. and Gustafsson, L. (1993). Inheritance of size and shape in a natural population of collared flycatchers, Ficedula albicollis. J Evol Biol, 6: 375–395.
Merilä, J. and Sheldon, B. C. in pressAvian quantitative genetics. Current Ornithology, 16.
Merilä, J., Sheldon, B. C. and Ellegren, H. (1998). Quantitative genetics of sexual size dimorphism in the collared flycatcher, Ficedula albicollis. Evolution, 52: 870–876.
Møller, A. P. and Birkhead, T. R. (1992). Validation of the heritability method to estimate extra-pair paternity in birds. Oikos, 64: 485–488.
Norris, K. J. and Blakey, J. K. (1989). Evidence for cuckoldry in the great tit Parus major. Ibis, 131: 436–441.
Petren, K. (1998). Microsatellite primers from Geospiza fortis and cross-species amplification in Darwin’s Finches. Mol Ecol, 7: 1782–1784.
Petren, K., Grant, R. B. and Grant, P. R. (1999). Low extrapair paternity in the cactus finch (Geospiza scandens). Auk, 116: 252–256.
Potti, J. and Merino, S. (1994). Heritability estimates and maternal effects on tarsus length in pied flycatchers, Ficedula hypoleuca. Oecologia, 100: 331–338.
Price, T. D. (1998). Maternal and paternal effects in birds: effects on offspring fitness. In: Mousseau, T. A. and Fox, C. W. (eds) Maternal Effects as Adaptations, pp. 202–226. Oxford University Press, New York.
Price, T. D. and Grant, P. R. (1984). Life history traits and natural selection for small body size in a population of Darwin’s finches. Evolution, 38: 483–494.
Roff, D. A. (1997). Evolutionary Quantitative Genetics. Chapman & Hall, New York.
SAS INSTITUTE INC (1990). SAS/STAT User’s Guide, version 6, 4th edn. SAS Institute, Cary, NC.
Smith, H. G. and Wettermark, K. J. (1995). Heritability of nestling growth in cross-fostered European starlings Sturnus vulgaris. Genetics, 141: 657–665.
Sokal, R. R. and Rohlf, F. J. (1981). Biometry, 2nd edn., W. H. Freeman, New York.
Weir, B. S. (1990). Genetic Data Analysis. Sinauer Associates, Sunderland, MA.
Acknowledgements
We thank E. Dyreson, A. van Noordwijk, T. Price, and K. Rodríguez-Clark for helpful discussions, J. Merilä and B. Sheldon for providing an unpublished manuscript, and two anonymous reviewers for helpful comments. The Charles Darwin Research Station (Galápagos) and the Galápagos National Parks Service gave crucial logistical support in the field. Numerous workers in the field helped measure finches. J. Coombs-Hahn provided valuable help in the laboratory. Financial support was provided by the Swiss Academy of Sciences, the Swiss National Science Foundation, and the Roche Research Foundation (to LFK), the US National Science Foundation (to PRG and BRG), and the NSF-Sloan Foundation (to KP).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Keller, L., Grant, P., Grant, B. et al. Heritability of morphological traits in Darwin’s Finches: misidentified paternity and maternal effects. Heredity 87, 325–336 (2001). https://doi.org/10.1046/j.1365-2540.2001.00900.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00900.x