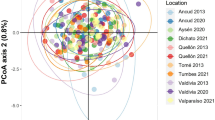
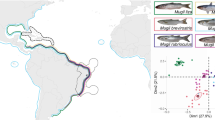
Abstract
Biochemical genetic variation at 12 allozyme loci was examined among three sympatrically occurring clam species (Astarte borealis, A. elliptica and A. striata: Bivalvia, Mollusca) from a single location off the east coast of Newfoundland, Canada. Nei's (1978) genetic distance (D) values for pairwise comparisons between the species ranged from 0.445 (between A. borealis and A. elliptica) to 0.530 (between A. borealis and A. striata) indicating speciation events ≈2.2–2.7 Myr BP. The timing of these speciation events is consistent with the proposed period of rapid speciation among the North Atlantic bivalve fauna following a significant extinction event (or events) ≈3.0 Myr BP. Allozyme variation among the three species was limited to allele frequency differences, with no absolute (fixed) differences being observed. The absence of diagnostic allelic differences in light of the large Nei's D-values is most unusual. The most plausible explanation is a form of balancing selection, whereby selection for the retention of ancestral alleles has prevented genetic diversity from increasing among the species via drift. Thirty of 36 exact tests for genetic differentiation were highly significant, indicating that the three species are genetically differentiated at these loci despite the absence of fixed allelic differences among the species. Compared with Hardy–Weinberg expectations, the allozyme loci exhibited heterozygote deficiencies which were highly statistically significant in all but one case. Two phenomena appear to account for the unusually large FIS values observed for all three species: first, inbreeding resulting in part from the extremely limited dispersal capabilities of the species, and second, a microgeographical Wahlund effect possibly operating at a scale of only tens of metres. Despite the absence of fixed allozyme differences, multivariate analyses of 14 morphometric traits and of allozyme variation at 12 loci both provided evidence of a high degree of genetic differentiation among the three species. Given the ability of the multivariate analyses of the allozyme and morphological variation to distinguish among sympatric A. borealis, A. elliptica and A. striata, it is suggested that the application of such analytical techniques on a larger scale will provide answers to long-standing questions about the uncertain taxonomy and systematics of the genus Astarte.
Similar content being viewed by others
Introduction
Species of the marine bivalve clam genus Astarte are relatively widely distributed throughout the Northern Hemisphere. Representatives of the genus occur in the east and west of the North Atlantic Ocean, throughout the Arctic Ocean, and from Japan eastward along the Alaskan and Pacific Coasts (Ockelmann, 1958; Durham & MacNeil, 1967; Lubinsky, 1980).
Despite the widespread distribution and considerable ecological importance of these clams in shallow subtidal and continental shelf soft-substrate habitats, relatively little is known of the biology and ecology of the genus. It has been reported that the sexes are usually separate, that spawning occurs late in the year (although at any one time a significant proportion of clams may have ripe gonads), and that the eggs are large (typically 150–200 μm diameter) and develop lecithotrophically (Ockelmann, 1958; Goodsell et al., 1982). Ockelmann (1958), p. 79) reported that Astarte spp. larvae are not observed in the water column at the time of spawning because ‘the eggs are adhesive by means of a mucous membrane ...’ and that a pelagic larval dispersive phase is either very short or entirely absent. Nonplanktotrophic larval development has subsequently been demonstrated for A. castanea, which possesses the adhesive encapsulated egg stage typical of the genus, and for which it was estimated that larval dispersive capability is of the order of centimetres only (Goodsell et al., 1982). It has also been reported that all species of the genus are likely to be hermaphrodites (Ockelmann, 1958) although there is no direct evidence confirming this suggestion.
No published data exist that describe genetic variation for any Astarte species. The unusually limited dispersal capability of the genus provides an opportunity to examine population genetic structure and to determine how this is influenced by extremely low numbers of migrants per generation. The present paper focuses on A. borealis (Schumacher 1817), A. elliptica (Brown 1827) and A. striata (Leach 1819) (designations based upon descriptions provided by Lubinsky, 1980) which occur sympatrically off the east coast of Newfoundland, Atlantic Canada. Allozyme genetic variation at 12 loci among the three co-occurring clam species is analysed and the species-specific population genetic structure of each is determined. The degree of genetic distinctness among the three sympatric species is tested, and the evolutionary relationships investigated. Finally, the utility of multivariate analyses of both allozyme and morphometric data is assessed as a means of addressing the uncertain taxonomy of the genus (Ockelmann, 1958; Lubinsky, 1980).
Materials and methods
Study site and sample collection
Specimens were collected on 14 December 1993 using a dredge with a heavy outer nylon bag lining which covered an inner lining composed of two layers of nylon mesh, each of 10 mm mesh size. This permitted individual clams as small as 8 mm shell length to be collected. Two 10-minute tows were made in ≈170–180 m of water at a station located east of St. John's, Newfoundland (47°33§ N, 52°36§ W). All specimens were placed in individually labelled bins before being returned to the Ocean Sciences Centre. The clams were sorted by species according to descriptions provided by Lubinsky (1980), placed in labelled bags, and stored at −70°C for subsequent analysis.
Allozyme variation
Horizontal starch gel electrophoresis was carried out on digestive gland tissue dissected from each clam. This tissue was homogenized with an equal volume of distilled water and one drop of 0.4% Triton X100 detergent before being centrifuged. The clear supernatant was used as the enzyme source. The Tris-maleate gel buffer system of Koehn et al. (1984) was used for all assays. A 1% solution of agar was added to each stain and the mixture was poured over the cut gel, to form an overlay. Nine systems yielding 12 loci were assayed (Table 1). When multiple loci were observed, the slowest migrating locus was designated 1, e.g. Pgm-1 refers to the slowest Pgm locus. The slowest migrating allele at each locus was designated 1, (e.g. Lap1) with subsequent faster migratory alleles receiving subsequent numeric designations. Cross-comparisons were made among gels and among species to ensure scoring accuracy.
Analyses of the allozyme data were carried out as follows. The software package GENEPOP version 1.2 (Raymond & Rousset, 1995) was employed to test for species differentiation (a test of independence between locus-specific allele frequencies and species designation). This test therefore provides a ‘fine scale’ test of species differentiation by comparing genetic differences on a locus-by-locus basis. Conformity to Hardy–Weinberg expectations was determined using an exact test employing the Markov chain method at each locus for each of the three species. Expected numbers of heterozygotes and homozygotes were computed using Levene's correction (Raymond & Rousset, 1995). Gene frequencies and FIS estimates were calculated for each allele according to Weir & Cockerham (1984). Following the recommendations of Raymond & Rousset (1995) the standard error (SE) estimates for all calculations were kept to ≤0.01. Nei's (1978) genetic identity (I) and genetic distance (D) values were calculated for pairwise comparisons of the three species to provide an estimate of their genetic relatedness. Finally, the software package SAS (SAS Institute Inc., 1987) was employed to analyse the allozyme data by Principal Components Analysis (PCA). This procedure provides a ‘large scale’ test of genetic differentiation among the species because it considers information from all allozyme loci at the same time, and therefore contrasts with the previously described ‘fine scale’ test of independence between locus-specific allele frequencies and species designation. For every individual in the PCA, each allele at each locus was coded as zero (absent), one (present in the heterozygous condition) or two (present in the homozygous condition). PCA was carried out to determine whether the degree of genetic differentiation was sufficient to permit correct assignment of any randomly chosen clam to its species designation. If this were possible, then the three clam species should be represented by three relatively discrete clusters of points when principal component one (PC1) is plotted against PC2. To test further this assumption, nonparametric analysis of variance (ANOVA) of ranked PC1 values was carried out (all values were ranked because nonranked data were not normally distributed), and mean rank PC1 values were compared among the three clam species. If the ANOVA probability value was significant (P<0.05) the Ryan–Einot–Gabriel–Welsch multiple range test (which controls the type I error rate and does not require equal samples sizes) was employed to identify the location of significant differences among the mean rank PC1 values of the three species.
Morphometric variation
Shells were assigned to one of three species (A. borealis, A. elliptica or A. striata) based upon shell descriptions provided by Lubinsky (1980). Fourteen morphometric traits, including shell length, were measured for each clam to the nearest 0.1 mm with either calipers or a stereomicroscope fitted with an ocular eyepiece (Fig. 1). Measurement error using these techniques does not contribute significantly to trait variation (Gardner, 1995). Values for each of the 13 morphometric characters were log10-transformed before being divided by log10-transformed shell length to correct for size-dependent variation. Principal components analysis of these data was carried out using the SAS software package (SAS Institute Inc., 1987). The data were examined to determine whether the morphometric characters could be employed to identify each of the three clam species, in which case they would be represented as three relatively distinct clusters of points. Multivariate analysis of variance (MANOVA) of the 13 log10-transformed and length-standardized traits was carried out to determine if significant differences in the dependent variables (morphometric traits) could be explained by variation in the independent variables (species designation). The Ryan–Einot–Gabriel–Welsch multiple range test was employed to identify the location of significant differences in mean trait values among the three species for each univariate ANOVA. After correction for multiple testing (Rice, 1989), species-specific log10-transformed length-standardized values for morphometric traits were normally distributed and independent of each other in almost all cases.
Fourteen shell traits measured for each individual clam in Astarte: (A, distance between pallial line and anterior shell margin; B, height of anterior adductor muscle scar; C, width of anterior adductor muscle scar; D, width of cardinal tooth; E, distance between cardinal tooth and pallial line; F, distance between ventral margin of anterior adductor muscle scar and ventral margin of posterior adductor muscle scar; G, height of posterior adductor muscle scar; H, width of posterior adductor muscle scar; I, distance between pallial line and ventral shell margin; J, shell height; K, shell width; L, shell length; M, distance between cardinal tooth and posterior margin of escutcheon; N, distance between cardinal tooth and anterior end of escutcheon).
Results
Allozyme variation
At most loci, the genetic variation among the three species was represented as differences in allele frequencies, the magnitude of which varied from locus to locus depending upon the species-pair under consideration (Table 2). The only monomorphic locus (defined here as exhibiting only one allele) was Idh-2 for A. elliptica: all other loci for all three species were polymorphic. The slightly lower numbers of A. elliptica and A. striata alleles per locus are probably attributable to their smaller sample sizes compared with A. borealis. Ten species-specific alleles (five for A. borealis, three for A. elliptica and two for A. striata) at seven different loci were observed. Seven of these 10 alleles may be considered to be rare in that they occurred at frequencies ≤0.05.
Exact tests of genetic differentiation indicate that despite the absence of fixed allelic differences at any of the 12 loci among the three species, most of the loci were significantly genetically differentiated between pairwise combinations of the three species. After correction for multiple testing (Rice, 1989), 30 of 36 test results provided evidence of significant genetic differentiation between species-pairs (for all significant pairwise combinations P≤0.00315). Nonsignificant differences occurred between A. borealis and A. elliptica for Idh-2, Mdh-1 and Pgm-1, between A. borealis and A. striata for Pgm-2, and between A. elliptica and A. striata for Mdh-2 and Pgm-2.
For each species, exact tests of the conformity of genotype frequencies at each locus to Hardy–Weinberg expectations indicate that all 35 polymorphic loci (i.e. all loci excluding Idh-2, which is fixed for A. elliptica) exhibited deficits of heterozygotes (Table 3). Tablewide correction for multiple testing (Rice, 1989) indicated that all results with the sole exception of Pgm-1 for A. striata were significant. FIS values for all three species were consistently very large across all loci (mean ±SD of 0.624±0.272 for A. borealis, 0.678±0.276 for A. elliptica, and 0.687±0.247 for A. striata). In all three species, the Idh-1, Idh-2 and Mdh-2 loci were actually or very nearly completely homozygous (excluding the Idh-2 locus which is absent in A. elliptica). Observed heterozygosity (Ho) values ranged from 0.000 to 0.549 (mean ±SD of 0.258±0.193) for A. borealis, 0.000–0.543 (0.194±0.173) for A. elliptica, and 0.000–0.552 (0.220±0.179) for A. striata.
Nei's (1978) I and D-values between species-pairs are presented in Table 4. The D-values ranged from 0.445 to 0.530 and indicated that the most genetically differentiated species-pair was A. borealis and A. striata, whereas the most genetically similar pair was A. borealis and A. elliptica. The three D-values were relatively similar, however, and indicate that all three pairwise species combinations are approximately equally related.
PCA of the allozyme data revealed the existence of three discrete (nonoverlapping) clusters of points when the second principal component (PC2) was plotted as a function of PC1 (Fig. 2). Clusters were consistent with the species designations according to Lubinsky (1980). PC1 explained ≈9% of the total variance in the data set, and 50% of the variance was explained by the first 12 principal components. Nonparametric analysis of variance (ANOVA) indicated that each species was significantly different from each of the others in mean rank PC1 values (Table 5). The ANOVA model explained ≈60% of the variance in the data set (R2=0.586).
Morphometric variation
PCA of the morphometric data revealed three relatively discrete, but slightly overlapping, clusters when PC2 was plotted as a function of PC1 (Fig. 3). Clusters were consistent with the species designations according to Lubinsky (1980). PC1 explained ≈50% of the variance in the data set, the first five principal components explained ≈84%, and the first 10 principal components explained >97% of the variance. MANOVA revealed the existence of highly significant differences among species for variation in all morphometric traits (Table 6). After correction for multiple testing (Rice, 1989) significant variation was also observed among the three species for 12 of the 13 traits (P<0.0013 in all cases). Species-specific differences explained from 10% to 81% of the variation in each individual trait. Only differences in log10-transformed length-standardized shell height were not significant.
Discussion
From the equation t=5×106D it is possible to estimate the time of divergence between species-pairs (Nei, 1987). For A. borealis and A. elliptica this is 2.25 Myr, for A. borealis and A. striata 2.65 Myr, and for A. elliptica and A. striata 2.45 Myr. The similarity of these estimates suggests that the three species arose at much the same time from an ancestral stock or from the oldest of the three species, which is likely to be A. borealis (Lubinsky, 1980). The timing of this multiple speciation event as derived from the allozyme data is consistent with biogeographical knowledge. Durham & MacNeil (1967) and Vermeij (1991) indicate that the genus Astarte is of North Atlantic origin with a fossil record extending back to the Oligocene (≈25–35 Myr BP). Briggs (1995) notes that at about 3.0 Myr BP most of the boreal species of the cold-temperate Arctic Ocean fauna were eliminated by a major cooling event and the modern fauna began to develop. The effects of this cooling event were also felt much further south, in the Atlantic Ocean, in the southern North Sea, and in the Mediterranean (Raffi et al., 1986; Stanley, 1986). These major extinction events set the stage for the evolution of a new boreal (cold-temperate) North Atlantic fauna. The biochemical genetics data for this genus are therefore consistent with the hypothesis that the period immediately following the major extinction episode for bivalve species through most the northern Atlantic Ocean was followed by a period of rapid speciation. The most plausible explanation for this burst of speciation is the ability of surviving genera/species to take advantage of newly vacant niches.
The absence of diagnostic alleles among three sympatrically occurring congeneric bivalve species which exhibit large genetic distances has not previously been reported. There are three possible explanations. First, the shared alleles might result from interspecific hybridization (be it ongoing or an historical episode). Such hybridization would need to result in extensive introgression to erode species-specific allelic differences, whilst, at the same time, maintaining significant allele frequency differences among the respective genomes. However, the three species are distantly related (see Nei's D and I-values, Table 4) which reduces the likelihood of hybridization, and the PCA plot of the allozyme data (Fig. 2) is inconsistent with the suggestion of extensive hybridization. Three clearly defined species-specific clusters of points are apparent with no evidence of the genetic intermediacy (identifiable as individual points intermediate between the species-specific clusters of points) which one expects in the event of hybridization. Other explanations (see later) seem much more plausible, and for the above reasons we do not favour the hybridization hypothesis. The second possible explanation is one of retained ancestral polymorphism. The three Astarte species might exhibit considerable allelic similarity as a result of their descent from a common ancestor which was characterized by such allelic diversity. However, it is unlikely that three species which are so distantly related would retain the observed degree of ancestral polymorphism under a neutral-drift scenario. Such a situation would be expected to increase genetic differences among the species rather than to preserve them, and ultimately to result in the establishment of absolute (=diagnostic) differences among the species. This, however, has not happened. The third possible explanation involves a form of balancing selection at each locus. The considerable genetic similarity among the three species in terms of shared alleles may have arisen as a result of selection within each species for the same alleles across all loci (at least, across those loci sampled). Balancing selection therefore prevents or reduces the establishment of increased genetic divergence, and might occur among newly established sibling species which exhibit considerable genetic similarity immediately after the speciation event(s). The true cause of the absence of fixed allelic differences among the three species might, of course, result from two or more of the above explanations. Further examination of other allozyme loci, of mitochondrial DNA, of other populations, or of different species within the genus should help to resolve the problem.
Tests of conformity to Hardy–Weinberg expectations indicate that all loci (except Pgm-1 of A. striata) of all three species exhibit significant heterozygote deficiencies. Such findings are in agreement with a large body of literature which indicates that marine bivalve populations are usually characterized by heterozygote deficiencies (reviewed by Zouros, 1987; Zouros & Foltz, 1987). However, the magnitude of the species-specific heterozygote deficiencies at each locus and their occurrence at so many loci at the same time is unusual. Based upon knowledge of the reproductive biology of the genus Astarte and the nature of the habitat from which it was collected, two separate factors are most likely to have contributed to the very high FIS values across all loci. The first factor is the mode of reproduction of the genus (refer to details in the Introduction) which results in an extremely low dispersal capability (Goodsell et al., 1982), coupled with the possibility of hermaphroditism (Ockelmann, 1958). Inbreeding, most probably via self-fertilization, will result in significant deficits of heterozygote genotypes. It is also likely to result in a population genetic structure that contains a number of individuals with identical composite multilocus genotypes. Inbreeding, resulting from small population size and the absence of migrants between populations, will result in very high FIS values across all loci, but will not tend to result in a population genetic structure containing many individuals with the same composite multiple genotypes. Examination of the data set indicates that for all three species every single multilocus composite genotype was unique (n=175 for A. borealis, n=35 for A. elliptica and n=29 for A. striata), indicating that selfing as an explanation for the high FIS values is unlikely, whereas biparental inbreeding is much more likely. Whether one or more of the species is hermaphroditic, and how this would contribute to population genetic structure, remains to be established.
A second factor contributing to the very high FIS values appears to be the Wahlund effect. Potentially this can involve the pooling of subpopulations such as cohorts which are characterized by different allele frequencies (population subdivision on a temporal scale), or might involve the pooling of geographical subpopulations which are characterized by differing allele frequencies (spatial population subdivision). The data support the existence of some form of intraspecific Wahlund effect, because the usual pattern of locus-specific response as a result of the Wahlund effect is for loci to exhibit different degrees of heterozygote deficiency according to the magnitude of the differences in allele frequencies between subpopulations at each locus (Richardson et al., 1986). This was indeed observed for all three species (FIS values ranged from 0.274 to 1.000 for A. borealis, from 0.018 to 1.000 for A. elliptica, and from 0.239 to 1.000 for A. striata) and therefore the Wahlund effect appears to be a significant factor in contributing to the observed heterozygote deficiencies. As a simple test of cohort population subdivision (genotypic heterogeneity among different age classes) the data for each species were divided into two subsets based upon shell size (clam size and age are assumed to be strongly positively correlated) to give species-specific allozyme information for small (presumptive young) and large (presumptive old) clams (data for this test are not formally presented). Tests of genotypic differentiation at each locus were carried out between small and large clams for each of the three species. After species-specific correction for multiple testing (Rice, 1989), the Pgm-2 locus of A. borealis and the Pgi locus of A. elliptica were observed to exhibit significant genotypic differentiation: all other loci exhibited genotypic homogeneity for all three species. From this somewhat crude test there is no evidence of presumptive age-dependent genotypic differentiation for any of the species. Any further testing of cohort population subdivision will require detailed information about the age structure of each species, which is presently not available. It therefore appears that a small-scale spatial Wahlund effect is in operation, and that clams from different populations, perhaps only a few metres apart, were pooled during each 10 min trawl sample. It remains to be established on what geographical scale population differentiation exists, but given the very low dispersal capability of the genus (Goodsell et al., 1982) it may be of the order of tens of centimetres to a maximum of a few metres, coincident with the scale of suitable habitat type (sand/mud) among boulder/cobble outcrops.
There is confusion regarding the taxonomic and systematic relationships of members of the genus Astarte (Ockelmann, 1958; Lubinsky, 1980). Despite the absence of absolute allelic differences at the loci examined, exact tests of genetic differentiation indicate that all but six of 36 locus comparisons are significantly differentiated between species-pairs. However, because the 12 loci surveyed constitute such a small portion of the genome of each species it is possible that further examination of allozyme variation will identify one or more loci which do exhibit fixed allelic differences among these species. Estimates of Nei's I and D-values (Nei, 1978) also indicate that the three taxa are sufficiently differentiated to be considered as separate species. For example, Hoagland (1984) has suggested that D-values of ≥0.3 among marine molluscs are sufficient to regard the taxa as distinct species. Such an estimate is consistent with other published D-values between ‘good’ species (e.g. Ayala, 1975; Nei, 1987). In other cases of questionable taxonomy and uncertain systematic affinity, morphometric and/or allozyme variation have been successfully analysed by multivariate statistical techniques to permit the identification of co-occurring and even hybridizing species (reviewed by Gardner, 1997). In the present study, PCA of allozyme and morphometric data both indicate that three separate species coexist. The PCA of the allozyme data differentiates the three species more clearly than the PCA of the morphometric data (i.e. the three clusters of species-specific points were more distinct in the former analysis) which probably reflects the greater utility of genetic data over morphometric data for species discrimination, as observed by other authors (e.g. McDonald et al., 1991; Gardner, 1996). These analyses strongly indicate that the three Astarte species maintain a high degree of genetic distinctness, and should be regarded as distinct or ‘good’ species (as suggested by Lubinsky, 1980), despite the fact that there are no absolute (fixed) differences in allele distributions among them. Further genetic and morphometric analyses of specimens from other locations throughout the wide range of this genus are required before a clearer understanding of the taxonomy and evolutionary relationships among its species can be gained.
References
Ayala, F. J. (1975). Genetic differentiation during the speciation process. Evol Biol, 8: 1–78.
Briggs, J. C. (1995). Global Biogeography. Elsevier, Amsterdam.
Durham, J. W. and Macneil, F. S. (1967). Cenozoic migrations of marine invertebrates through the Bering Strait region. In: Hopkins, D. M. (ed.) The Bering Land Bridge, pp. 326–349. Stanford University Press, Stanford, CA.
Gardner, J. P. A. (1995). Developmental stability is not disrupted by extensive hybridization and introgression among populations of the marine bivalve molluscs Mytilus edulis (L.) and M. galloprovincialis (Lmk.) from south-west England. Biol J Linn Soc, 54: 71–86.
Gardner, J. P. A. (1996). The Mytilus edulis species complex in southwest England: the extent of hybridization and introgression and their effects upon interlocus associations and morphometric variation. Mar Biol, 125: 385–399.
Gardner, J. P. A. (1997). Hybridization in the sea. Adv Mar Biol, 31: 1–78.
Goodsell, J. G., Lutz, R. A., Castagna, M. and Kraeuter, J. (1982). Nonplanktotrophic larval development of two species of continental shelf bivalves. J Shellfish Res, 2: 91
Hoagland, K. E. (1984). Use of molecular genetics to distinguish species of the gastropod genus Crepidula (Prosobranchia: Calyptraeidae). Malacologia, 25: 607–628.
Koehn, R. K., Hall, J. G., Innes, D. J. and Zera, A. J. (1984). Genetic differentiation of Mytilus edulis in eastern North America. Mar Biol, 79: 117–126.
Lubinsky, I. (1980). Marine bivalve molluscs of the Canadian central and eastern Arctic: faunal composition and zoogeography. Can Bull Fish Aquat Sci, 207: 1–111.
McDonald, J. H., Seed, R. and Koehn, R. K. (1991). Allozymes and morphometric characters of three species of Mytilus in the Northern and Southern Hemispheres. Mar Biol, 111: 323–333.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
Nei, M. (1987). Molecular Evolutionary Genetics. Columbia University Press, New York.
Ockelmann, W. K. (1958). The zoology of East Greenland: marine Lamellibranchiata. Medd Grønl, 122: 1–256.
Raffi, S., Stanley, S. M. and Marasti, R. (1986). Biogeographic patterns and Plio-Pleistocene extinction of Bivalvia in the Mediterranean and southern North Sea. Paleobiology, 11: 368–388.
Raymond, R. and Rousset, F. (1995). Genepop (v. 1.2). A population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Rice, W. R. (1989). Analyzing tables of statistical tests. Evolution, 43: 223–225.
Richardson, B. J., Baverstock, P. R. and Adams, M. (1986). Allozyme Electrophoresis. A Handbook for Animal Systematics and Population Studies. Academic Press, Sydney.
Sas Institute, INC. (1987). SAS/STAT guide for personal computers. Release 603. Cary, NC.
Stanley, S. M. (1986). Anatomy of a regional mass extinction: Plio-Pleistocene decimation of the western Atlantic bivalve fauna. Palaios, 1: 17–36.
Vermeij, G. J. (1991). Anatomy of an invasion: the trans-Arctic interchange. Paleobiology, 17: 281–307.
Weir, B. S. and Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38: 1358–1370.
Zouros, E. (1987). On the relation between heterozygosity and heterosis: an evaluation of the evidence from marine mollusks. Isozymes Curr. Top Biol Med Res, 15: 255–279.
Zouros, E. and Foltz, D. W. (1987). The use of allelic isozyme variation for the study of heterosis. Isozymes Curr. Top Biol Med Res, 13: 1–59.
Acknowledgements
We thank the master and crew of the C.S.S. Marinus for assistance with collection of specimens. Assistance in the laboratory was provided by Sunil Dhargalkar, Rebekah Eyles and Melanie Saunders. We thank Prof. Charles Daugherty and Dr David Innes for helpful discussion and comments on an earlier version of the manuscript. This research was supported by a Natural Science and Engineering Research Council of Canada (NSERC) Research Grant to R.J.T. J.P.A.G. was supported by a postdoctoral fellowship funded by NSERC (Joint Global Ocean Flux Study programme).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gardner, J., Thompson, R. High levels of shared allozyme polymorphism among strongly differentiated congeneric clams of the genus Astarte (Bivalvia: Mollusca). Heredity 82, 89–99 (1999). https://doi.org/10.1038/sj.hdy.6884590
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6884590
Keywords
This article is cited by
-
Morphological characteristics and genetic differentiation of Lutraria maxima in coast waters off southeast China
Journal of Oceanology and Limnology (2021)
-
Population genetic variation in the New Zealand greenshell mussel: locus-dependent conflicting signals of weak structure and high gene flow balanced against pronounced structure and high self-recruitment
Marine Biology (2013)