Abstract
The olive tree (Olea europaea) has successfully invaded several regions in Australia and Pacific islands. Two olive subspecies (subspp. europaea and cuspidata) were first introduced in these areas during the nineteenth century. In the present study, we determine the origin of invasive olives and investigate the importance of historical effects on the genetic diversity of populations. Four invasive populations from Australia and Hawaii were characterized using eight nuclear DNA microsatellites, plastid DNA markers as well as ITS-1 sequences. Based on these data, their genetic similarity with native populations was investigated, and it was determined that East Australian and Hawaiian populations (subsp. cuspidata) have originated from southern Africa while South Australian populations (subsp. europaea) have mostly derived from western or central Mediterranean cultivars. Invasive populations of subsp. cuspidata showed significant loss of genetic diversity in comparison to a putative source population, and a recent bottleneck was evidenced in Hawaii. Conversely, invasive populations of subsp. europaea did not display significant loss of genetic diversity in comparison to a native Mediterranean population. Different histories of invasion were inferred for these two taxa with multiple cultivars introduced restoring gene diversity for europaea and a single successful founder event and sequential introductions to East Australia and then Hawaii for cuspidata. Furthermore, one hybrid (cuspidata × europaea) was identified in East Australia. The importance of hybridizations in the future evolution of the olive invasiveness remains to be investigated.
Similar content being viewed by others
Introduction
Humans have contributed to the breaching of natural dispersal barriers of many organisms as they transported them around the globe (Mooney and Cleland, 2001). Plants have been introduced into new areas either accidentally or voluntarily (for example, as crops or ornamentals, as well as for erosion control, forage or timber; Baker, 1974). In their novel environments, some plants establish, spread and become invasive with potential detrimental consequences on the new ecosystems they invade (Sakai et al., 2001; Keane and Crawley, 2002). Invasive species have become a major conservation issue. They are now regarded as a significant component of global environmental change (Elton, 1958; D’Antonio and Vitousek, 1992; Lodge, 1993; Vitousek et al., 1996, 1997a, 1997b) and one of the most important drivers of biodiversity loss, second only to habitat destruction (Walker and Steffen, 1997; Wilcove et al., 1998; Sala et al., 2000). Indeed, invaders compete with local species for limited resources, alter ecosystem processes and increase disturbance (Hobbs and Mooney, 1986; Vitousek, 1990; Williamson, 1996). Intentional and accidental introduction of invasive plants into new ranges has been an ever-increasing phenomenon with the enhancement of global trade and transport (D’Antonio and Vitousek, 1992; Levine and D’Antonio, 2003). As a consequence, multiple introductions of the same species have often occurred (Kolbe et al., 2004; Genton et al., 2005; Facon et al., 2006) and hybridizations between two previously isolated taxa or populations may also act as a stimulus for the evolution of invasiveness (Abbott et al., 2003; Ellstrand and Schierenbeck, 2006).
The olive tree (Olea europaea L.; Oleaceae) is frequently considered as a pest where it has been introduced. It is a highly invasive tree in the Pacific islands and in parts of East and South Australia (West, 2002; Starr et al., 2003; Cooke et al., 2005; Bass et al., 2006). The native wild olive relatives are distributed from South Africa to South Asia, in Saharan mountains, Macaronesia and Mediterranean countries (Green, 2002). Two subspecies display a large continental distribution: subsp. europaea in the Mediterranean basin and subsp. cuspidata from Austral Africa to China. These taxa are diploids, allogamous and anemophilous, and their seeds are dispersed by birds (Spennemann and Allen, 2000; Green, 2002). Subspecies europaea was domesticated about 6000 years ago in the Near East and in the Iberian Peninsula (Zohary and Hopf, 2000; Besnard et al., 2001b; Terral et al., 2004). Humans have then largely contributed to its dissemination in the Mediterranean region and introduced cultivated forms into new areas (for example, Australia, California, Chile and South Africa). Subspecies cuspidata is not of a great economic importance, but its wood is sometimes used to make furniture in South Africa and it is cultivated as an ornamental tree in China. In addition, it has been utilized as a rootstock and hedge plant (Spennemann and Allen, 2000; Starr et al., 2003).
In Australia, cultivated olives were first brought to Sydney in 1800 (Spennemann and Allen, 2000) and to Adelaide in 1836 (Guerin et al., 2003). After a decline of the Australian olive industry in the early 1900s, an increasing interest in olive products occurred during the late 1940s and 1950s (Bass et al., 2006). Before the 1920s, naturalized forms established in and invaded South Australia (Adelaide) across a wide range of habitats, predominantly within the 400–600 mm median annual rainfall range (Spennemann and Allen, 2000; Mekuria et al., 2002; Bass et al., 2006). Such naturalized populations are now found in other parts of Australia (Western Australia, Victoria, New South Wales and southeast Queensland; Bass et al., 2006) and in northern New Zealand (Heenan et al., 1999). Moreover, in the 1990s, Australian olive industry was revitalized leading to a considerable increase in the number of olive groves. It is possible that new feral forms will escape from these areas in the coming decades (Spennemann and Allen, 2000; Bass et al., 2006). Today, there are over 100 known varieties of olives in Australia (Sweeney and Davies, 1998). Furthermore, in the 1810s, O. europaea subsp. cuspidata was introduced to Australia and naturalized populations were first recorded at the Norfolk Island and in the Sydney region (Cumberland plain woodland) from the nineteenth century (Spennemann and Allen, 2000). It has also been reported in the Hawaii Archipelago since the 1960s (Starr et al., 2003).
Humans have mainly contributed to intercontinental displacements of this species while active bird dispersion of olive fruits (particularly by the Common Starling) has allowed local dissemination and putatively relatively long distance migrations (Rey and Alcántara, 2000; Spennemann and Allen, 2000; Bass et al., 2006). Invasive populations of both subspecies are now representing a serious threat for the indigenous floras in woodlands of southern and Eastern Australia, Norfolk Island, Northern New Zealand and Hawaii as they can form the dominant vegetation and rapidly shade out native species (Green, 1994; Csurhes and Edwards, 1998; Wagner et al., 1999; West, 2002; Cooke et al., 2005; Bass et al., 2006). Furthermore, the proportion of endemic species is high in these areas and understanding the dynamics and impact of invasions is an important step for the conservation of the concerned ecosystems.
The phylogeographic approach has already provided very pertinent insights to determine the source of introduction of some invasive species (Amsellem et al., 2000; Milne and Abbott, 2004). Besnard et al. (2007) have recently reconstructed a detailed phylogeography of the O. europaea complex in its native range based on DNA polymorphism from the nuclear (bi-parentally inherited) and plastid (maternally inherited) genomes. This study provided powerful data to test the origins of naturalized olive populations. Furthermore, Besnard et al. (2007) identified hybridization and reticulation among the olive subspecies. Consequently, subspecies europaea and cuspidata are likely to hybridize or introgress when in sympatry (Besnard et al., 2001c; Rubio de Casas et al., 2006). This phenomenon could alleviate the loss of genetic diversity due to bottlenecks arising from small initial founder populations during colonization events (Husband and Barrett, 1991; Lee, 2002) and contribute to the ecological success of colonizing populations. However, genomic incompatibilities may affect the fitness of early generations of hybrids (Rieseberg et al., 1996).
The origins of invasive olive populations and their mutual relationship have not been documented yet. Here we address this issue using molecular data from both native and invasive trees. The importance of genetic diversity loss during the invasions of Australia and the Pacific Islands was assessed. Possible hybridization between distinct invasive subspecies was also tested. The resulting data were considered to compare the invasion process from either cultivated (subsp. europaea) or wild olives (subsp. cuspidata).
Materials and methods
Plant material
Four invasive populations of O. europaea were sampled in Australia (Campbelltown, 34°07S, 150°49E; Brownhill Creek, 34°59S, 138°37E and Lonsdale, 35°06S, 138°29E) and Hawaii (Maui, 20°51N, 156°20W). Between 26 and 30 individuals per population were included in the study (Table 1). Samples of randomly chosen adult trees were collected along a transect of about 200 m. According to their geographical origin (Wagner et al., 1999; Spennemann and Allen, 2000) and morphological traits (for example, undersurface color of leaves, fruit size), these populations were considered to be either related to subsp. cuspidata (Campbelltown and Maui) or subsp. europaea (Brownhill Creek and Lonsdale; Adelaide region). In the Campbelltown and Maui populations the sampled trees are assumed to be relatively young since these areas were recently (less than 50 years) invaded by the olive tree. In the Adelaide region, the two olive groves are separated by only 16 km. The Brownhill Creek population is located near an ancient olive orchard. Some old feral trees attest that this population was established early, probably from seeds of the first orchards planted in the late 1800s/early 1900s (Bass et al., 2006). The Lonsdale population is probably more recent and may have resulted from seeds dispersed at relatively long distances as there are few planted olive trees nearby.
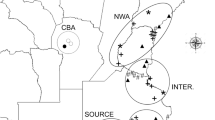
Additionally, 17 populations representative of the entire native range of the subspecies europaea and cuspidata were sampled (Figure 1; Table 1). A minimum of four individuals were collected from each population, excepted for Amalundu (Zimbabwe; one individual). Several of these populations were previously characterized with random amplified polypmorphic DNA (RAPD) markers and for plastid DNA (ptDNA) polymorphism (Besnard et al., 2001c, 2002, 2007). Only Gué de Constantine, Mallorca, Serra de Arrabida and Guangzhou were not analyzed with RAPDs. In addition, fifteen cultivars from Italy (Frantoio), France (Olivière, Cailletier), Spain (Arbequina, Cornicabra, Hojiblanca, Lechin de Sevilla, Manzanilla and Manzanilla de Sevilla), Portugal (Galega), Algeria (Taksrit), Tunisia (Zarazi), Egypt (Toffahi) and Syria (Zaity and Kaissy) plus three ornamental trees (Bonsai trees with unknown origin) were analyzed. The cultivars (excepted Hojiblanca and Mazanilla de Sevilla) were already characterized in previous studies with RAPD markers and for ptDNA polymorphism (Besnard et al., 2001a, 2001b, 2002, 2007).
Geographical locations of the populations of O. europaea prospected in the present study. Native and invasive populations are respectively distinguished by circles and squares. The shaded areas (in gray) represent the native distribution of both subspecies. For each invasive population, the frequency of ptDNA haplotypes is also given. *For each plastid haplotype, a code number is used and the corresponding lineage is indicated in bracket (see Supplementary material and Besnard et al., 2007).
Genomic DNA from fresh leaves or leaves desiccated in silica gel was extracted using a cetyltrimethylammonium bromide method (Besnard et al., 2000) for the 272 sampled individuals.
Molecular markers
Plastid DNA polymorphism
Maternal origins of invasive olive populations were inferred by likening their ptDNA haplotypes with those previously described in the native range by Besnard et al. (2007). Twenty polymorphic ptDNA loci were investigated to characterize Australian and Hawaiian populations as reported by Besnard et al. (2007). PCR conditions and electrophoresis procedures were used as previously described (Weising and Gardner, 1999; Besnard et al., 2003; Baali-Cherif and Besnard, 2005). These loci were developed based on sequences that were obtained for a sample of 20 trees representative of the 6 olive subspecies (Besnard et al., 2007). They allowed the identification of 40 haplotypes and 7 ptDNA lineages distributed in specific areas. Thus, lineages E1, E2 and E3 are primarily distributed in the Mediterranean area (subsp. europaea) while lineages A, C1 and C2 are only found in trees belonging to subsp. cuspidata (Besnard et al., 2007).
After determining ptDNA haplotypes of invasive individuals, the trnT-trnL and trnS-trnG intergenic spacers as well as the matK intron were sequenced for all identified haplotypes of each provenance. Sequences were compared to those previously obtained for 20 native trees (Besnard et al., 2007). This procedure allowed us to check for potentially undetected new polymorphism in invasive populations and also to confirm the haplotype identification based on the 20 polymorphic ptDNA loci. Sequencing was performed using the Big Dye 3.1 Terminator cycle sequencing kit (Applied Biosystems, Foster City, CA, USA) according to manufacturer's instructions and an ABI Prism 3100 genetic analyzer (Applied Biosystems).
Sequencing of internal transcribed spacer-1 (ITS-1)
The nuclear ribosomal DNA variation in the olive complex was previously investigated using ITS-1 sequences by Besnard et al. (2007). Seven ITS-1 lineages were detected in the native range and each one was distributed in a specific area (for example, populations of subsp. cuspidata were characterized by lineages I, IV and VI while Mediterranean trees displayed lineages II or III; Besnard et al., 2007). In the present study, one individual per invasive population was characterized for ITS-1 sequences as described by Besnard et al. (2007). These sequences were then aligned with those obtained for native trees (Besnard et al., 2007) to determine the ITS-1 lineage(s) of the invasive populations.
Nuclear DNA microsatellite markers
To assess intrapopulation genetic diversity, all individuals were characterized by means of nuclear microsatellites (SSR) because such genetic markers are codominant and generally highly polymorphic (Belaj et al., 2003). Eight polymorphic SSR loci were used as described by Baali-Cherif and Besnard (2005): ssrOeUA-DCA1, ssrOeUA-DCA3, ssrOeUA-DCA8, ssrOeUA-DCA9, ssrOeUA-DCA14, ssrOeUA-DCA15 (Sefc et al., 2000), PA(ATT)2 (Saumitou-Laprade et al., 2000) and EMO03 (de la Rosa et al., 2002). These loci are appropriate for assessing genetic diversity in the O. europaea complex as they show good transferability in the olive complex and because of very low frequency of null alleles (Baali-Cherif and Besnard, 2005).
Data analysis
Based on nuclear SSR data, the genetic similarity between populations (excluding Amalundu for which only one individual was available) was assessed. Nei's minimum genetic distances (Nei, 1987) were computed between pairs of populations using the software Populations 1.2.28 (Langella, 1999). A phenetic tree was built using the neighbor-joining algorithm (Saitou and Nei, 1987). Bootstrap values were computed using 1000 resamplings on individuals to evaluate support of the branches. Additionally, genetic relationships among subsp. cuspidata, subsp. europaea and the four invasive populations were ascertained and visualized by a factorial correspondence analysis (FCA) on individuals using the software Genetix version 4.05.2 (Belkhir et al., 2004).
We used an assignment test to compare the multilocus microsatellite genotype of each individual from the introduced range with each of the native populations. We calculated a probability of each invasive individual's genotype in each potential source population. This analysis was performed using the software Geneclass2 (Piry et al., 2004). We used the standard criterion described by Rannala and Mountain (1997), which applies Bayesian statistics to compute probabilities. In addition, we used the simulation algorithm for population assignment described by Paetkau et al. (2004). We simulated 10 000 genotypes for each population and applied an arbitrary threshold probability value of 0.05 or greater to determine origin of invasive individuals. If the individual's probability is lower than the decided threshold in a population, it considers that the individual does not originate from the population. The analysis was performed independently in subspp. cuspidata and europaea. For subsp. cuspidata, we removed the population of Amalundu (Zimbabwe), which was constituted of only one genotype. Six native populations were thus considered (South Africa, Kenya, Reunion, Yemen, Iran and China). For subsp. europaea, we considered only four groups of individuals from the native range: western Mediterranean (Portugal, Spain, Morocco and Algeria; 58 individuals), central Mediterranean (Tunisia and Libya; nine individuals), Eastern Mediterranean (Turkey and Syria; 13 individuals) and cultivars (15 individuals).
Gene diversity parameters in invasive populations were estimated and compared to their related populations (Gué de Constantine and Cape Town) in the native range (see below). Observed heterozygosity (HO) and expected heterozygosity (HS, gene diversity; Nei, 1987) were calculated from allele frequencies at each nuclear SSR locus independently using Fstat version 2.9.4 (Goudet, 2005). Deviation of observed genotypes from Hardy – Weinberg equilibrium was computed for each locus in each population using Genepop version 3.4 (Raymond and Rousset, 2003). To account for different sample sizes of populations, allelic richness (RS) was also estimated using Fstat. A Wilcoxon paired test (one-sided) was used to evaluate the significance of the differences of allelic richness and gene diversity between invasive and native populations. Pairwise FST for both nuclear SSR and ptDNA data were computed using Fstat for the four invasive populations. Significance of pairwise differentiation was assessed using standard Bonferroni corrections.
In addition, the software Bottleneck version 1.2.02 (Cornuet and Luikart, 1996; Piry et al., 1999) was used to evaluate recent effective population size reductions in the invasive populations. The two-phased model (TPM) of mutation was applied with 5% of multistep changes as recommended by Luikart and Cornuet (1998) for microsatellite markers. The same model with 30% of multistep changes and the stepwise mutation model (SMM) were also tested. This program uses the fact that recently bottlenecked populations exhibit an excess of gene diversity among polymorphic loci. The observed gene diversity (He) is compared to the gene diversity expected at mutation-drift equilibrium (Heq) given the number of alleles and the sample size (see Piry et al. (1999) and Kahru et al. (2006) for more detailed descriptions of the method). Several statistical tests have been proposed to evaluate the significance of such differences (Cornuet and Luikart, 1996), and we applied the Wilcoxon signed-rank test, as it does not require a large number of polymorphic loci and is statistically more powerful than the other tests in detecting bottlenecks (Maudet et al., 2002). Furthermore, a qualitative descriptor of allele frequency distribution ‘mode shift’ was used to help the discrimination of stable from bottlenecked populations (Luikart and Cornuet, 1998).
Results
Plastid DNA data
Four ptDNA haplotypes belonging to four distinct ptDNA lineages (E1, E2, E3 and A; Besnard et al., 2007) were revealed in the invasive populations (Supplementary material). In Brownhill Creek and Lonsdale (South Australia) populations, we detected haplotypes 1 (71%), 9 (27%) and 12 (2%), which respectively belong to the Mediterranean sublineages E1, E3 and E2 (Besnard et al., 2007). In contrast, the populations from Campbelltown and Maui mainly displayed haplotype 39 (98%), which is characteristic of the Austral African sublineage A (Besnard et al., 2007). Furthermore, one individual from Campbelltown (numbered Ca-21) displayed the Mediterranean haplotype 1. Plastid DNA sequences (EMBL accessions AM229535–AM229554) obtained for six individuals (BC-21, haplotype 9; BC-23, haplotype 1; Lo-7, haplotype 12; Ca-1, haplotype 39; Ca-21, haplotype 1 and Ma-1, haplotype 39) confirmed the haplotype identification based on the 20 ptDNA loci.
ITS-1 sequence analysis
We isolated ITS-1 sequences (EMBL accessions AM229655–AM229660) from five individuals (BC-21, Lo-7, Ma-1, Ca-1 and Ca-21). These sequences were unambiguously assigned to an ITS-1 lineage as previously described by Besnard et al. (2007). First, the sequences isolated from BC-21 and Lo-7 differed by only one substitution. These sequences were identical to those isolated from Mediterranean individuals (AJ877275 and AJ877273-AJ877274-AJ877276, respectively; lineage III). Second, the sequences obtained in Ca-1 and Ma-1 did not differ and were identical to those obtained for South African populations (EMBL accessions AJ877298–AJ877300; Lineage IV). Therefore, these nuclear data also clearly support that South Australian populations are related to Mediterranean olives while Campbelltown and Maui are related to Austral African populations. Additionally, two divergent sequences belonging to lineages III (Mediterranean) and IV (Austral Africa) were isolated from Ca-21 suggesting that this individual is a hybrid. These two sequences were identical to EMBL accessions AJ877298–AJ877300 and AJ877275, respectively.
Genetic relationships between native and invasive populations based on nuclear SSR data
The genetic similarity between invasive and native populations was assessed based on nuclear SSR data by construction of a phenetic tree (Figure 2). Populations of the subspecies cuspidata and europaea formed two independent clusters (Figure 2). Invasive populations from Campbelltown and Maui were clearly grouped (100% of bootstrap support) and placed in the cuspidata cluster. The clade formed by populations from Cape Town (South Africa), Campbelltown and Maui is supported by a bootstrap value of 100%. The Mediterranean group (subsp. europaea) was divided into one clade grouping populations from East Mediterranean (51% of bootstrap support) and a second clade grouping populations from the west Mediterranean region (75% of bootstrap support). Libyan and Tunisian populations were placed in an intermediate position between these two clades as shown in a previous study based on RAPD markers (Besnard et al., 2001c). Both invasive populations from South Australia were clearly grouped together (98% of bootstrap support) and also placed in an intermediate position between the East and west Mediterranean populations.
Unrooted phenetic tree showing the genetic similarities between invasive and native populations using eight nuclear SSR loci. The analysis is based on minimum genetic distances (Nei, 1987) and the neighbor-joining algorithm (Nei, 1987). Bootstrap values superior to 50% for 1000 replicates are indicated on each corresponding branch. The line separates populations belonging to subspecies europaea and cuspidata. Invasive populations are indicated in bold.
The FCA performed on all cuspidata individuals (Figure 3a) confirmed that invasive populations of Campbelltown and Maui are closely related. Moreover, this analysis also highlights the close genetic affinities of these populations with South African wild trees (Cape Town). The similarity between the Cape Town population with introduced populations of Campbelltown and Maui was also observed when assignment tests were performed on individuals from the introduced range (Table 2). In this analysis, about 84% of invasive trees were significantly assigned to the population of Cape Town. No individual was assigned to another native population. Moreover, 100% of invasive trees were assigned to the population of Cape Town when an arbitrary threshold probability value of 0.01 was used (data not shown).
Factorial correspondence analyses on nuclear SSR data for individuals of subspecies cuspidata (a) and europaea (b). Percentage of explained variance is reported for the first three dimensions. These analyses were based on 147 and 142 alleles for subspecies cuspidata and europaea, respectively. For subsp. europaea, the cultivars are numbered: 1, Cornicabra; 2, Zarazi; 3, Taksrit; 4, Arbequina; 5, Hojiblanca; 6, Lechin de Sevilla; 7, Olivière; 8, Manzanilla de Sevilla; 9, Manzanilla; 10, Cailletier; 11, Kaissy; 12, Galega; 13, Frantoio; 14, Zaity, 15, Toffahi.
In the second FCA performed on all europaea individuals (Figure 3b), East (Turkey and Syria) and West (Portugal, Mallorca, Morocco and Algeria) Mediterranean trees can be separated into two groups. Libyan and Tunisian oleasters as well as cultivars are not specifically placed in one of these two groups. Syrian and Turkish oleasters are separated from invasive olives and oleasters from the west Mediterranean Basin on axis 1, while west Mediterranean trees are separated from invasive olives and oleasters from the East Mediterranean Basin on axis 2. Invasive trees from South Australia displayed an intermediate position between East and west Mediterranean oleasters. Interestingly, all cultivars originating from west Mediterranean (and particularly Arbequina, Cornicabra, Hojiblanca, Taksrit and Zarazi) cluster with invasive trees. The assignment test of South Australia invasive trees to native populations of subsp. europaea showed that invasive trees are not clearly assigned to a particular group as previously shown for subsp. cuspidata (Table 2). Forty-four (75%) individuals were assigned to at least two groups. Nevertheless, invasive trees were mostly assigned to central Mediterranean populations (81.4%) and to cultivars (72.9%).
Genetic differentiation between invasive populations
Pairwise FST values between invasive populations were firstly computed based on nuclear SSR loci (Table 3). The lowest value (0.8%, not significant) was detected between the two South Australian populations (Brownhill Creek and Lonsdale). All other comparisons are highly significant even at the 0.1% nominal level (Table 3). The population differentiation between the Maui and Campbelltown populations was of 17.5%. A higher differentiation (ranging from 29.5 to 36.1%) was found between invasive populations from South Australia and those from East Australia and Hawaii.
Pairwise FST values for ptDNA haplotypes were partially incongruent with the results obtained using nuclear SSR (Table 3). Indeed, no differentiation was observed between Maui and Campbelltown, while a high differentiation (39.4%) was observed between the two South Australian populations. As shown with nuclear SSR data, the differentiation between invasive populations from South Australia and those from East Australia and Hawaii was especially high (70.2–93%; Table 3).
Evidence of hybridization
To study the relationships between all invasive individuals, an FCA was applied on nuclear SSR data (Figure 4). As expected, populations from South Australia (group 1) were clearly distinguished from Campbelltown and Maui populations (group 2). This distinction was mainly supported by the first axis of the FCA. In addition, Campbelltown and Maui populations were also distinguished based on the second axis of the FCA. Interestingly, individual Ca-21 was clearly in an intermediate position between groups 1 and 2. In addition, this individual has a hybrid pattern at all loci which allow clear distinction of both groups (ssrOeUA-DCA3 (alleles 233–253), ssrOeUA-DCA8 (alleles 127–139), ssrOeUA-DCA14 (alleles 150–190) and EMO03 (alleles 213–223)), indicating that it is probably a hybrid of first generation.
FCA on nuclear SSR data for all individuals of invasive O. europaea populations. Percentage of explained variance is reported for the first three dimensions. South Australian (group 1, subsp. europaea) and Campbelltown–Maui clusters (group 2, subsp. cuspidata) are indicated. Individual Ca-21 is in an intermediate position between subspecies europaea and cuspidata.
Assessment of the loss of genetic diversity in invasive populations
The genetic diversity of invasive populations was investigated and directly compared to one related population sampled in the native range. Thus, Lonsdale and Brownhill Creek populations were compared to a native population from the Mediterranean region (Gué de Constantine; 34 individuals), whereas Campbelltown and Maui populations were compared to a South African population (Cape Town; 18 individuals). For the Campbelltown population, we excluded individual Ca-21 because it is assumed to be an inter-subspecific hybrid. In all populations, the observed heterozygosity was not significantly different from the expected heterozygosity at each nuclear SSR locus, suggesting no excess of homozygosity (Table 4). The allelic richness (RS) and gene diversity (HS) of South Australian populations were not significantly different from Gué de Constantine's. In contrast, the populations from Campbelltown and Maui display a lower allelic richness and gene diversity than the Cape Town population (Table 4; Wilcoxon paired tests (6 d.f.): RS (Campbelltown) <RS (Cape Town), V=28, P<0.01; RS (Maui) <RS (Cape Town), V=28, P<0.01; HS (Maui) <HS (Cape Town), V=28, P<0.01). Moreover, the Maui population displayed a particularly low allelic richness (RS=16.8) and gene diversity (HS=0.41). Among the 17 alleles detected in this population, 16 were also detected at Campbelltown (of which 13 with a frequency superior to 15%).
Results of bottleneck tests are presented in Table 5. Once again, we excluded individual Ca-21. As expected, no evidence of bottleneck was observed for the two native populations from Gué de Constantine and Cape Town. From the four invasive populations studied, only two (Maui and Brownhill Creek) showed a significant evidence of a recent bottleneck (Table 5). However, the allele frequency distribution of the Brownhill Creek population (South Australia) was approximately L-shaped, which is a property of populations at mutation-drift equilibrium. For the six analyzed populations, when using the TPM of mutation with 30% of multistep changes or the SMM, congruent qualitative results were obtained (data not shown).
Discussion
Origins of invasive olive populations
In the present study, four plastid haplotypes characteristic of both subspecies europaea (1, 9 and 12) and cuspidata (39; Besnard et al., 2007) were identified in invasive populations from Australia and Hawaii. Confirming morphological observations, populations from South Australia display haplotypes of subsp. europaea, while the populations from Maui and Campbelltown mostly shared one haplotype of subsp. cuspidata (Austral Africa). Similarly, ITS-1 sequences and nuclear SSR markers (Figures 2 and 3) support the hypothesis that South Australian populations belong to the Mediterranean taxon (subsp. europaea) whereas Maui and Campbelltown populations are closely related to South African populations (subsp. cuspidata). These data clearly confirm the introduction of two divergent olive subspecies in Australia from which invasive populations originated (Spennemann and Allen, 2000). The strong genetic differentiation observed between East and South Australian populations (Figure 2; Table 3) can therefore be mainly explained by the historical divergence between cuspidata and europaea populations (Rubio de Casas et al., 2006; Besnard et al., 2007).
The invasive populations from Campbelltown and Maui (subsp. cuspidata) are closely related to the Cape Town population (Figures 2 and 3; Table 2). Moreover, they display particularly low nuclear and ptDNA diversity (Figure 1; Table 4), suggesting a single introduction of trees from southern Africa. Conversely to East Australian and Hawaiian populations, the origin of South Australian populations (subsp. europaea) remains elusive as they probably originated from cultivated forms (Spennemann and Allen, 2000). The detection of three divergent Mediterranean ptDNA haplotypes clearly indicates that europaea populations originated from multiple individuals. Based on nuclear SSR data, East and west Mediterranean oleaster populations appear to be differentiated, as shown in previous studies (Besnard et al., 2001c, 2002; Lumaret et al., 2004). The phenetic analysis and the FCA (Figures 2 and 3b) placed both South Australian populations at an intermediate position between the two main groups of native populations with some cultivars and oleasters from Libya and Tunisia. The similarity between cultivars, central Mediterranean and South Australian populations was also observed using assignment tests (Table 2). Based on our data, we suppose that cultivars introduced in Australia have mostly originated from the west or central Mediterranean Basin, because olive cultivars from these areas are known to have resulted mainly from hybridizations between Eastern and western Mediterranean trees (Besnard et al., 2001b, 2002).
Interestingly, the South Australian populations are strongly differentiated based on ptDNA markers (FST=39.4%; Table 3) while no differentiation (FST=0.8%; Table 3) is observed on nuclear markers. The high differentiation revealed on the plastid genome (which is maternally inherited) is surprising since these populations have been sampled in the same area. Nevertheless, this pattern indicates that gene flow between invasive populations is more intense by pollen than by seeds, as already shown in native populations (Besnard et al., 2002). Furthermore, this high differentiation on maternal markers results either from a historical effect (founder effect) and/or from a selective effect. Indeed, about 50% of trees from Brownhill Creek display haplotype 9 (referred as CCK in Besnard et al., 2000), which was shown to be strictly associated to a cytoplasmic male sterility (CMS) in cultivars (Besnard et al., 2000). This CMS may play a significant role in the dynamics of invasion since male sterile individuals are known to be very vigorous in olives (for example, cultivars Zarazi, Chemlal or Olivière; Besnard et al., 2000) and should therefore be favored in the initial establishment of populations compared to hermaphrodites as shown in Thymus vulgaris (Manicacci et al., 1996).
Loss of genetic diversity during the invasion process
Genetic diversity of subsp. cuspidata was significantly reduced in invasive populations compared to the South African population. This clear difference in genetic diversity between invasive and native populations fits the prediction that introduction of few founder individuals and their subsequent expansion in the new area will lead to a reduced genetic diversity (Husband and Barrett, 1991). Furthermore, East Australia could have been an intermediate source for the introduction to Hawaii as indicated by historical records (for example, Wagner et al., 1999; Spennemann and Allen, 2000; Starr et al., 2003). In our study, this hypothesis is supported by the fact that the Maui population is the most genetically impoverished and displays a subsample of alleles found in the Campbelltown population. In addition, the strong and recent bottleneck evidenced in the Maui population is probably due to consecutive founder events in the history of introduction to the Hawaiian Archipelago (from Austral Africa to Australia and then to Hawaii). Conversely, the East Australian population displays no significant evidence of a recent bottleneck, probably due to its more ancient origin. Furthermore, based on nuclear SSRs, we revealed a surprisingly high genetic differentiation between these populations (FST=17.5%). The number of generations since introduction in Australia (200 years ago) may not exceed 25. This means that genetic differentiation between distant populations originating from the same pool may rapidly appear due to genetic drift and isolation.
Southern Australian populations display no significant difference in genetic diversity when compared to a Mediterranean population. This supports the hypothesis that multiple individuals of subsp. europaea (cultivars) have been introduced in Australia, as the supply of divergent genotypes from disparate source populations alleviates the loss of genetic diversity occurring during introduction (Amsellem et al., 2000; Bossdorf et al., 2005; Genton et al., 2005). One of these two populations (Brownhill Creek) also exhibits evidence of a recent bottleneck. However, because of high frequency of the cytoplasm associated to CMS (52%) and the proximity of this population with present orchards, it is possible that recurrent gene flow from cultivated clones may have affected the results of the bottleneck test used (based on He>Heq).
Inter-subspecies hybridization in the invasive range
A single hybrid (Ca-21) was detected throughout the whole study area. It was clearly identified based on both ptDNA and nuclear DNA data. Further studies with a more thorough sampling of invasive populations would be necessary to detect other hybrids as well as estimating the relative fitness of hybrids versus pure lines to determine the importance of hybridization in the invasion process of O. europaea in Oceania and Pacific Islands. It could also be of interest to compare the frequency of hybrids in old and young trees to determine if inter-subspecies hybridizations would increase in time and space.
Concluding remarks
The present study on two subspecies of O. europaea has identified two clearly distinct cases of olive invasion. One taxon (from the wild) suffers from reduced genetic diversity due to bottlenecks and the other (from cultivars) enjoys higher genetic diversity due to the introduction of multiple individuals and yet both are successful invaders. This study confirmed that successful invasion can occur even after an important bottleneck (for example, Hawaii). Furthermore, subspecies cuspidata and europaea invaded different areas, with the former mainly found in warm temperate to tropical areas and the latter in areas displaying a typical Mediterranean climate. However, as both subspecies increase their ranges, chances for hybridization and introgression will increase, putatively leading to higher within-population genetic diversity and increasing potential for evolutionary changes (Kolbe et al., 2004; Facon et al., 2006). Nevertheless, the genomic, ecological and phenological compatibilities between europaea and cuspidata populations are still largely unknown and should be explored, to understand the impact of hybridization in the evolution of this invasive species.
References
Abbott RJ, James JK, Milne RI, Gillies ACM (2003). Plant introductions, hybridization and gene flow. Philos Trans Roy Soc Lond Ser B 358: 1123–1132.
Amsellem L, Noyer JL, Le Bourgeois T, Hossaert-Mckey M (2000). Comparison of genetic diversity of the invasive weed Rubus alceifolius Poir. (Rosaceae) in its native range and in areas of introduction, using amplified fragment length polymorphism (AFLP) markers. Mol Ecol 9: 443–455.
Baali-Cherif D, Besnard G (2005). High genetic diversity and clonal growth in relict populations of Olea europaea subsp. laperrinei (Oleaceae) from Hoggar, Algeria. Ann Bot 96: 823–830.
Baker HG (1974). The evolution of weeds. Annu Rev Ecol Syst 5: 1–24.
Bass DA, Crossman ND, Lawrie SL, Lethbridge MR (2006). The importance of population growth, seed dispersal and habitat suitability in determining plant invasiveness. Euphytica 148: 97–109.
Belaj A, Satovic Z, Cipriani G, Baldoni L, Testolin R, Rallo L et al. (2003). Comparative study of the discriminating capacity of RAPD, AFLP and SSR markers and of their effectiveness in establishing genetic relationships in olive. Theor Appl Genet 107: 736–744.
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (2004). GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations. Laboratoire Génome, Populations, Interactions CNRS UMR 5000, Université de Montpellier II, Montpellier (France). http://www.univ-montp2.fr/~genetix/genetix/genetix.htm.
Besnard G, Baradat P, Bervillé A (2001a). Genetic relationships in the olive (Olea europaea L.) reflect multilocal selection of cultivars. Theor Appl Genet 102: 251–258.
Besnard G, Baradat P, Breton C, Khadari B, Bervillé A (2001b). Olive domestication from structure of wild and cultivated populations using nuclear RAPDs and mitochondrial RFLPs. Genet Sel Evol 33 (Suppl 1): S251–S268.
Besnard G, Baradat P, Chevalier D, Tagmount A, Bervillé A (2001c). Genetic differentiation in the olive complex (Olea europaea) revealed by RAPDs and RFLPs in the rRNA genes. Genet Res Crop Evol 48: 165–182.
Besnard G, Khadari B, Baradat P, Bervillé A (2002). Olea europaea (Oleaceae) phylogeography based on chloroplast DNA polymorphism. Theor Appl Genet 104: 1353–1361.
Besnard G, Khadari B, Villemur P, Bervillé A (2000). Cytoplasmic male sterility in the olive (Olea europaea L.). Theor Appl Genet 100: 1018–1024.
Besnard G, Rubio de Casas R, Vargas P (2003). A set of primers for length and nucleotide-substitution polymorphism in chloroplastic DNA of Olea europaea L. (Oleaceae). Mol Ecol Notes 3: 651–653.
Besnard G, Rubio de Casas R, Vargas P (2007). Plastid and nuclear DNA polymorphism reveals historical processes of isolation and reticulation in the olive tree complex (Olea europaea L.). J Biogeogr 34: 736–752.
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005). Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144: 1–11.
Cooke J, Willis T, Groves R (2005). Impacts of woody weeds on Cumberland Plain Woodland biodiversity. Impacts of woody weeds on Cumberland Plain woodland biodiversity. In: The Ecology and Management of Cumberland Plain Habitats. University of W Sydney: Campbelltown.
Cornuet JM, Luikart G (1996). Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144: 2001–2014.
Csurhes S, Edwards R (1998). Potential Environmental Weeds in Australia: Candidate Species for Preventative Control. Queensland Department of Natural Resources: Canberra.
D’Antonio CM, Vitousek PM (1992). Biological invasions by exotic grasses, the grass/fire cycle, and global change. Annu Rev Ecol Syst 23: 63–87.
de la Rosa R, James CM, Tobutt KR (2002). Isolation and characterization of polymorphic microsatellites in olive (Olea europaea L.) and their transferability to other genera in the Oleaceae. Mol Ecol Notes 2: 265–267.
Ellstrand NC, Schierenbeck KA (2006). Hybridization as a stimulus for the evolution of invasiveness in plants? Euphytica 148: 35–46.
Elton CS (1958). The Ecology of Invasions by Animal and Plants. Methuen: London.
Facon B, Genton BJ, Shykoff J, Jarne P, Estoup A, David P (2006). A general eco-evolutionary framework for understanding bioinvasions. Trends Ecol Evol 21: 130–135.
Genton BJ, Shykoff JA, Giraud T (2005). High genetic diversity in French invasive populations of common ragweed, Ambrosia artemisiifolia, as a result of multiple sources of introduction. Mol Ecol 14: 4275–4285.
Goudet J (2005). FSTAT (version 2.9.4): a program to estimate and test population genetics parameters. http://www2.unil.ch/popgen/softwares/fstat.htm.
Green PS (1994). Flora of Australia. Vol. 49: Oceanic islands 1. Australian Government Publishing Service: Canberra.
Green PS (2002). A revision of Olea L. (Oleaceae). Kew Bull 57: 91–140.
Guerin J, Mekuria G, Burr M, Collins G, Sedgley M (2003). Selection of improved olive cultivars. Acta Hort 622: 231–234.
Heenan PB, de Lange PJ, Glenny DS, Breitwieser I, Brownsey PJ, Ogle CC (1999). Checklist of dicotyledons, gymnosperms, and pteridophytes naturalised in New Zealand: additional records 1997–1998. NZ J Bot 37: 629–642.
Hobbs RJ, Mooney HA (1986). Community changes following shrub invasion of grassland. Oecologia 4: 508–513.
Husband BC, Barrett SCH (1991). Colonisation history and population genetic structure of Eichornia paniculata in Jamaica. Heredity 66: 287–296.
Kahru A, Vogl C, Moran GF, Bell JC, Savolainen O (2006). Analysis of microsatellite variation in Pinus radiata reveals effects of genetic drift but no recent bottlenecks. J Evol Biol 19: 167–175.
Keane M, Crawley J (2002). Exotic plant invasions and the enemy release hypothesis. Trends Ecol Evol 17: 164–170.
Kolbe JJ, Glor RE, Schettino LR, Lara AC, Larson A, Losos JB (2004). Genetic variation increases during biological invasion by a Cuban lizard. Nature 431: 177–181.
Langella O (1999). POPULATIONS 1.2.28 software. http://www.pge.cnrs-gif.fr/bioinfo/populations.
Lee CE (2002). Evolutionary genetics of invasive species. Trends Ecol Evol 17: 386–391.
Levine JM, D’Antonio CM (2003). Forecasting biological invasions with increasing international trade. Conserv Biol 17: 322–326.
Lodge DM (1993). Biological invasions: lessons for ecology. Trends Ecol Evol 8: 133–137.
Luikart G, Cornuet JM (1998). Empirical evaluation of a test for identifying recently bottlenecked populations from allele frequency data. Conserv Biol 12: 228–237.
Lumaret R, Ouazzani N, Michaud H, Vivier G, Deguilloux MF, Di Giusto F (2004). Allozyme variation of oleaster populations (wild olive tree) (Olea europaea L.) in the Mediterranean Basin. Heredity 92: 343–351.
Manicacci D, Couvet D, Belhassen E, Gouyon PH, Atlan A (1996). Founder effects and sex ratio in the gynodioecious Thymus vulgaris L. Mol Ecol 5: 63–72.
Maudet C, Miller C, Bassano B, Breitenmosser-Würsten C, Gauthier D, Obexer-Ruff G et al. (2002). Microsatellite DNA and recent statistical methods in wildlife conservation management: applications in Alpine ibex (Capra ibex (ibex)). Mol Ecol 11: 421–436.
Mekuria GT, Collins G, Sedgley M (2002). Genetic diversity within an isolated olive (Olea europaea L.) population in relation to feral spread. Sci Hort 94: 91–105.
Milne RI, Abbott RJ (2004). Geographic origin and taxonomic status of the invasive Privet, Ligustrum robustum (Oleaceae), in the Mascarene Islands, determined by chloroplast DNA and RAPDs. Heredity 92: 78–87.
Mooney HA, Cleland EE (2001). The evolutionary impact of invasive species. Proc Natl Acad Sci USA 98: 5446–5451.
Nei M (1987). Molecular Evolutionary Genetics. Columbia University Press: New York.
Paetkau D, Slade R, Burden M, Estoup A (2004). Direct, real-time estimation of migration rate using assignment methods: a simulation-based exploration of accuracy and power. Mol Ecol 13: 55–65.
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004). GENECLASS2: A software for genetic assignment and first-generation migrant detection. J Hered 95: 536–539.
Piry S, Luikart G, Cornuet JM (1999). BOTTLENECK: A computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90: 502–503.
Rannala B, Mountain JL (1997). Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci USA 94: 9197–9201.
Raymond M, Rousset F (2003). GENEPOP 3.4., an updated version of GENEPOP V.1.2 (1995): population genetics software for exact tests and ecumenicism. J Hered 86: 248–249.
Rey PJ, Alcántara JM (2000). Recruitment dynamics of a fleshy-fruited plant (Olea europaea): connecting patterns of seed dispersal to seedling establishment. J Ecol 88: 622–633.
Rieseberg LH, Sinervo B, Linder CR, Ungerer MC, Aria DM (1996). Role of gene interactions in hybrid speciation: evidence from ancient and experimental hybrids. Science 272: 741–744.
Rubio de Casas R, Besnard G, Schöenswetter P, Balaguer L, Vargas P (2006). Extensive gene flow blurs phylogeographic but not phylogenetic signal in Olea europaea L. Theor Appl Genet 113: 575–583.
Saitou N, Nei M (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4: 406–425.
Sakai AK, Allendorf FW, Holt JS, Lodge DM, Molofsky J, With KA et al. (2001). The population biology of invasive species. Annu Rev Ecol Syst 32: 305–332.
Sala OE, Chapin III FS, Armesto JJ, Berlow E, Bloomfield J, Dirzo R et al. (2000). Global biodiversity scenarios for the year 2100. Science 287: 1770–1774.
Saumitou-Laprade P, Vassiliadis C, Epplen JT, Hardt C (2000). Isolation of microsatellite loci for paternity testing in Phillyrea angustifolia L. (Oleaceae). Mol Ecol 9: 112–114.
Sefc KM, Lopes MS, Mendonca D, Rodrigues dos Santos M, Laimer da Câmara Machado M, da Câmara Machado A (2000). Identification of microsatellite loci in olive (Olea europaea) and their characterization in Italian and Iberian olive trees. Mol Ecol 9: 1171–1173.
Spennemann DHR, Allen LR (2000). Feral olives (Olea europaea) as future woody weeds in Australia: a review. Aust J Exp Agric 40: 889–901.
Starr F, Starr K, Loope L (2003). Plants of Hawaii. http://www.hear.org/starr/hiplants/reports/html/index.html.
Sweeney S, Davies G (1998). The Olive industry. In: Hide K (ed). The New Rural Industries: A Handbook for Farmers and Investors. RIRDC: Barton. pp. 405–411.
Terral JF, Alonso N, Capdevila RBI, Chatti N, Fabre L, Fiorentino G et al. (2004). Historical biogeography of olive domestication (Olea europaea L.) as revealed by geometrical morphometry applied to biological and archaeological material. J Biogeogr 31: 63–77.
Vitousek PM (1990). Biological invasions and ecosystem process: towards an integration of population biology and ecosystem studies. Oikos 57: 7–13.
Vitousek PM, D’Antonio CM, Loope LL, Rejmanek M, Westbrooks R (1997a). Introduced species: a significant component of human-caused global change. NZ J Ecol 21: 1–16.
Vitousek PM, D’Antonio CM, Loope LL, Westbrooks R (1996). Biological invasions as global environmental change. Am Sci 84: 468–478.
Vitousek PM, Mooney HA, Lubchenco J, Melillo JM (1997b). Human domination of earth's ecosystems. Science 277: 494–499.
Wagner WL, Herbst DR, Sohmer SH (1999). Manual of the Flowering Plants of Hawai’i Revised edition. University of Hawai’i Press: Honolulu.
Walker BH, Steffen W (1997). An overview of the implications of global change for natural and managed terrestrial ecosystems. Conserv Ecol 1: 2.
Weising K, Gardner RC (1999). A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledonous angiosperms. Genome 42: 9–19.
West CJ (2002). Eradication of allien plants on Raoul Island, Kermadec Islands, New Zealand. In: Veitch CR and Clout MN (eds). Turning the Tide: The Eradication of Invasive Species. IUCN SSC Invasive Species Specialist Group: Cambridge. pp 365–373.
Wilcove DS, Rothstein D, Dubow J, Phillips A, Loso E (1998). Quantifying threats to imperiled species in the United States. Bioscience 48: 607–615.
Williamson M (1996). Biological Invasions. Chapman & Hall: London.
Zohary D, Hopf M (2000). Domestication of Plants in the Old World. Clarendon Press: Oxford, UK.
Acknowledgements
We thank J Goudet and N Juillet for help on statistical analysis; we also thank L Amsellem, G Martin, N Galland, D Pio and PA Christin for helpful comments on the manuscript. We especially thank D Baali-Cherif, F Starr and K Starr, Y Spanoudis, R Rubio de Casas, Y Wang and A Jambois for olive sampling.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Supplementary information
Rights and permissions
About this article
Cite this article
Besnard, G., Henry, P., Wille, L. et al. On the origin of the invasive olives (Olea europaea L., Oleaceae). Heredity 99, 608–619 (2007). https://doi.org/10.1038/sj.hdy.6801037
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801037
Keywords
This article is cited by
-
Land use change rather than surrounding vegetation affects fungal endophyte assemblages in the African wild olive
Landscape Ecology (2023)
-
High genetic variability of Nosema ceranae populations in Apis mellifera from East Asia compared to central Asia and the Americas
Biological Invasions (2022)
-
Genome skimming-based simple sequence repeat (SSR) marker discovery and characterization in Grevillea robusta
Physiology and Molecular Biology of Plants (2021)
-
Identification of an olive (Olea europaea L.) core collection with a new set of SSR markers
Genetic Resources and Crop Evolution (2021)
-
Genomic evidence for recurrent genetic admixture during the domestication of Mediterranean olive trees (Olea europaea L.)
BMC Biology (2020)