Abstract
During the last decade, brackish northern pike populations in Denmark have been subject to stocking programmes, using nonindigenous pike from freshwater lakes, in order to compensate for drastic population declines. The present study was designed to investigate the genetic impact of stocking freshwater pike into a brackish pike population in Stege Nor, Denmark. We analysed polymorphism at eight microsatellite loci in samples representing the indigenous Stege Nor population prior to stocking (ie from 1956 to 1957), along with a sample of the contemporary Stege Nor population and samples from the three populations used for stocking. Despite large numbers of stocked fry, the results from both individual and population level admixture analyses demonstrated extremely poor performance and <1% introgression of stocked freshwater pike into the brackish pike population. Furthermore, pairwise FST estimates between samples demonstrated close genetic relationship among temporal samples from Stege Nor, indicating temporal stability over the last 45 years. We also estimated the effective population size (Ne) of pike in Stege Nor and applied a test for recent population bottlenecks. The harmonic mean of Ne was relatively high (>250), but there were indications of bottlenecks in all samples and populations. We ascribe this finding to historical rather than recent bottlenecks, possibly dating back to founder events associated with postglacial recolonisation.
Similar content being viewed by others
Introduction
During the past decades, there has been increased attention on the possible negative genetic consequences of human-mediated gene flow among genetically differentiated populations (Hindar et al, 1991; Waples, 1991; Allendorf et al, 2001; Beaumont et al, 2001; Randi and Lucchini, 2002). Thus, prevention of intentional or accidental releases of domesticated or exogenous wild individuals is now recognised as an important element in conservation programmes for plants and animals worldwide (Allendorf et al, 2001). The advent of highly variable microsatellite markers and the development of new statistical methods, combined with the ability to analyse historical samples, has improved possibilities for detecting gene flow and admixture between indigenous and transplanted or domesticated populations (eg Roy et al, 1994; Beaumont et al, 2001; Nielsen et al, 2001; Hansen, 2002; Randi and Lucchini, 2002).
Stocking has often been the first method used to alleviate problems with declining fish populations (Hindar et al, 1991). In the case of salmonid fishes, stocking based on releases of hatchery-reared fish has been conducted using strains that have been domesticated for several decades (eg Hindar et al, 1991; Hansen et al, 2001; Levin and Williams, 2002). In other species, however, transplantations of offspring of wild caught fish into different natural populations have also been conducted. Until recently, little was known about the genetic effects of transplantations of wild, exogenous individuals possibly lacking local adaptations to their new environment, compared to the effects of stocking domesticated fish into wild populations. However, northern pike provides an opportunity for addressing this issue, as in Denmark offspring of wild pike from freshwater lakes have been stocked in large numbers into brackish waters in order to increase total census population sizes in the local areas.
The northern pike (Esox lucius L.) exhibits a circumpolar geographical distribution in the Northern Hemisphere, where it is one of the most common predators in lakes, rivers and brackish waters. It is considered an economically important species for both commercial and recreational fisheries (Seeb et al, 1987). In the Baltic Sea, its distribution ranges from the inner Bothnian Bay to the southern Danish Belt Sea (Raat, 1988). During the last 40 years, the pike populations in the Baltic Sea have declined dramatically (more than 90% in many areas), possibly due to degradation and destruction of habitat (Højgaard, 1985; Westin and Limburg, 2002). To compensate for this decline, stocking of pike has been used to improve fisheries in brackish waters in Denmark. Until recently, it has not been feasible to apply genetic markers for evaluating the effects of pike stocking activity, due to extremely low levels of genetic variability at the allozyme and mitochondrial DNA level (Healy and Mulcahy, 1980; Seeb et al, 1987; Brzuzan et al, 1998; Maes et al, 2003). However, recent development of microsatellite markers for pike has provided efficient tools for studying the genetic population structure (Miller and Kapuscinski, 1996, 1997; Hansen et al, 1999; Senanan and Kapuscinski, 2000; Miller et al, 2001). The purpose of this study was to analyse the genetic impact of a decade of stocking freshwater pike into an indigenous brackish pike population of the Stege Nor fjord on the island of Møn, Denmark. Analyses were based on microsatellite DNA markers from old scale samples (1956–1957) taken prior to stocking, along with contemporary samples from both stocking material and the Stege Nor population. This rendered it possible to determine the origin of the present population, that is, to determine if the pike were descendents of indigenous pike, stocked pike or hybrids between the two groups. Furthermore, our aim was to estimate genetic differentiation among populations, and if stocking had not effected the contemporary sample, to assess whether the genetic composition of the Stege Nor population had been stable in the period between 1956–1957 and 2002. Finally, to address future conservation and management recommendations for the area, we estimated the effective population size (Ne) for the pike population in Stege Nor, using the temporal method, and tested for recent drastic reductions of effective population size, that is, bottlenecks.
Materials and methods
Sampled localities
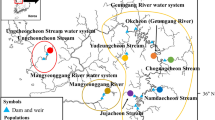
Pike were sampled in Stege Nor, a 5 km2 brackish fjord on the island of Møn, Denmark (Figure 1). This population has experienced a dramatic decline during the last 40 years. Thus, present data on commercial fish landings suggest that catches have decreased by 99% over the last four decades (L Jacobsen, Danish Institute for Fisheries Research, Department of Inland Fisheries, personal communication). Furthermore, the location of Stege Nor, between the brackish Baltic Sea (6–8‰ salinity in the central part) and the highly saline North Sea, results in periodical inflows of highly saline water. Several large inflows were registered in the winters of 1929–1930, 1950–1951, 1969–1970 and 1991–1992 (Dahl, 1961; Højgaard, 1985). Following the latest inflow in 1991–1992, the majority of pike in the area died and the fish landings dropped to <50 kg/year compared to several tonnes before the saltwater inflow. In order to augment recovery of the pike population, an intense stocking programme was initiated in 1993 using hatchery-reared pike fry from freshwater lakes. In total, nearly 400 000 fry have been stocked into the fjord in the period from 1993 to 2002. In 1993, pike fry used for stocking originated from Lake Vandet, Jutland. In later years, stocking material has been derived from three lakes on Zealand, Lake Sorø/Lake Tuel (these two lakes are connected and with no obstacles to migration of fish between the lakes) and most importantly Lake Tissø (Figure 1). Each year, spawners are caught in the wild, and the fry are reared in large tanks until stocking.
Contemporary and historical samples
Contemporary samples (fin tissue) were collected with assistance from local fishermen and fish farmers in 2001 and 2002. Following capture, all fish were length measured, then fin-clipped and the tissue placed in separate Eppendorf tubes (1.5 ml) containing 96% ethanol. Mature pike were caught at the spawning grounds in Lake Tuel/Lake Sorø in spring 2002 using nets (sample abbreviation RN02). Pike from Lake Tissø were caught in July 2002 by angling and electrofishing (sample abbreviation TS02). Samples from Lake Vandet were caught by pound net and gillnets in spring 2001 (sample abbreviation VN01). In all, 12 individuals from Stege Nor were sampled by local fishermen from July to November 2002 using a variety of methods, including gillnets and traps. The remaining 37 individuals were caught by angling and gillnets in November 2002 (sample abbreviation ST02). The size range of pike caught in Stege Nor in 2002 was 52–113 cm, representing several cohorts of pike. The indigenous pike population in Stege Nor prior to stocking was represented by historical scale samples, collected by fisheries biologists in 1956 and 1957.
Microsatellite analysis
DNA from ethanol-preserved fin clips was extracted using a proteinase K/Chelex method described by Estoup et al (1996). A phenol–chloroform extraction method, combined with the use of microconcentrators, was used for the old scale samples (Nielsen et al, 1999). Depending on the size of the scales, 1–2 scales were used for extraction. Standard polymerase chain reaction (PCR) was used to amplify eight microsatellite loci, Elu 51, Elu 64, Elu 78, Elu 87 (Miller and Kapuscinski, 1996), Elu 19, Elu 37, Elu 76 (Miller and Kapuscinski, 1997) and Elu 2 (Hansen et al, 1999). Elu 252 and Elu 276 (Miller and Kapuscinski, 1996, 1997) were initially tested in this study, but samples were later excluded due to suspected presence of null alleles and problems in obtaining reproducible results for historical samples. PCR involved 29 cycles for fresh tissue samples and 39 cycles for old scale samples. Annealing temperatures for old scale samples and fresh samples, respectively, were as follows: Elu 2 (54 and 56°C), Elu 19 (56 and 56°C), Elu 37 (52 and 58°C), Elu 51 (58 and 58°C), Elu 64 (56 and 58°C), Elu 76 (52 and 56°C), Elu 78 (56 and 58°C) and Elu 87 (56 and 58°C). For each locus, one primer was end-labelled with a fluorescent dye (CY5). Microsatellites were analysed on a Pharmacia ALFexpress automatic sequencer, and gels were scored using the software ALLELINKS version 1.00 according to the recommendations of the manufacturer (PHARMACIA). Reproducibility of results was tested by performing replicates of the molecular analyses for subsamples of the whole data set. For the fresh tissue samples and old scale samples, approx. 20 and 40% of individuals were reanalysed for each locus, respectively.
Data analysis
Microsatellite polymorphism was quantified by allelic richness, observed (Ho) and expected (He) heterozygosity for the different loci in the populations, using FSTAT version 2.9.3.2 (Goudet, 1995). Tests for deviation from Hardy–Weinberg equilibrium for each locus and tests for gametic phase equilibrium (linkage) for pairs of loci were conducted using a Markov chain Monte Carlo (MCMC) method (Guo and Thompson, 1992) implemented in GENEPOP version 3.1c (Raymond and Rousset, 1995). Levels of significance for multiple tests were determined using sequential Bonferroni adjustments for simultaneous tests (Rice, 1989). Genetic differentiation among all populations and between pairs of populations was estimated by θ (Weir and Cockerham, 1984), an unbiased estimator of FST, and 95% confidence intervals (CIs) were estimated based on bootstrapping 10 000 times over loci. These analyses were conducted using FSTAT version 2.9.3.2 (Goudet, 1995). The method and software LEADMIX (Wang, 2003) was used for estimating admixture proportions of the Stege Nor gene pool, that is, the proportion of the contemporary population derived from stocked and indigenous pike, respectively. This was carried out by using the historical Stege Nor samples and the populations used for stocking as baseline samples. Individual admixture proportions (q), that is, the part of an individual's genome that is derived from either indigenous or stocked fish, were estimated using STRUCTURE (Pritchard et al, 2000), a Bayesian clustering approach, based on MCMC. The number of populations represented by the sampled individuals was estimated by calculating the probability of the data assuming from one to six populations, as described by Pritchard et al (2000). These analyses were based on 200 000 burn-in steps followed by 1 000 000 MCMC replicates. Individual admixture proportions and their 90% posterior probability intervals were estimated based on the following prior assumptions: (a) Loci within baseline samples were at Hardy–Weinberg equilibrium and linkage equilibrium. (b) Baseline samples ST57, VN01, RN02 and TS02 consisted of nonadmixed individuals. (c) Individuals from the ST02 sample were either pure individuals or possible admixtures between pure populations. In all simulations, the length of the burn-in period was set to 20 000 iterations, followed by another 50 000 iterations collecting data. To assess the power of STRUCTURE for identifying admixed individuals, three groups of individuals were simulated, each consisting of 50 individuals. One group consisted of simulated individuals based on the allele frequencies of the ST57 population, the second group consisted of simulated individuals based on the allele frequencies of the three populations used for stocking and the last group consisted of admixed individuals (F1 hybrids) between ST57 and TS02. Simulation of individuals was conducted using the program HYBRIDLAB version 0.9 (Nielsen et al, 2001).
Effective population size (Ne) of the Stege Nor population was estimated using two different likelihood-based temporal methods implemented in the softwares TM3 (Berthier et al, 2002) and MLNE2 (Wang and Whitlock, 2003), respectively. TM3 estimates the effective population size based on two temporal samples from the same population. In this study, we based our estimates on 20 000 MCMC replicates, and results are presented as 5, 50 (median) and 95 percentiles of the posterior distribution of Ne based on an average of three simulations (as recommended by Berthier et al, 2002). MLNE2 also estimates Ne based on temporal samples using a maximum-likelihood approach. The method allows for estimating both Ne and migration rate, if samples representing possible sources of migration are included in the analysis. In the present case, however, we estimated Ne assuming an isolated population, thus representing a direct comparison to the results obtained using TM3. The mean age of spawners was used as a proxy for generation length (Hill, 1979; Miller and Kapuscinski, 1997). The estimated mean age at reproduction of males and females was 3 and 4 years, respectively, resulting in an average mean age of spawners estimated to 3.5 years. Thus, the 45 year time span between the two temporal samples corresponds to 12.9 generations. The detection of reductions in population size in the two Stege Nor samples and in the samples from the three populations used for stocking was conducted using the procedure (M_P_VAL) of Garza and Williamson (2001), which is based on the ratio M of the number of alleles compared to the range in allele size. The programme detects a bottleneck as an average of M across loci, based on the assumption that a reduction in population size is expected to reduce the number of alleles faster than the range of allele sizes (Garza and Williamson, 2001). Based on recommendations by the authors (Garza and Williamson, 2001), values of ps (proportion of one-step mutations) and Δg (average size of non one-step mutations) were set to 90 and 3.5%, respectively. The parameter θ was estimated as θ=4Nμ of the population (N and μ are the effective population size and the mutation rate, respectively). Several values of θ ranging from 0.02 to 2 were tested assuming different mutation rates and/or effective population sizes. Simulations were conducted using 10 000 replicates and the average value of M over all replicates is reported.
Results
Nearly all individuals from both contemporary and historical samples were successfully scored at the eight microsatellite loci. Only two individuals failed to amplify at one locus. Despite successful amplification and scoring, it cannot be ruled out entirely that degradation and small quantities of DNA have caused some allelic dropouts in the historical samples (Miller et al, 2002). However, tests for Hardy–Weinberg equilibrium did not suggest that dropouts affected our results. Tests for Hardy–Weinberg equilibrium and gametic phase equilibrium (linkage) showed no significant deviations after adjustment for multiple comparisons, using a sequential Bonferroni correction (Rice 1989). Summary statistics of variation at the eight microsatellite loci are provided in Table 1.
Genetic differentiation between populations
Pairwise tests for genetic differentiation between populations resulted in θ-values ranging from 0.014 to 0.30 (Table 2), with an overall θ for all samples of 0.17 (95% CI 0.11–0.23). There was no significant differentiation between the two temporal samples from Stege Nor. In contrast, significant genetic differentiation was observed between all three populations used for stocking, as well as between each of the stocking populations and the two Stege Nor samples.
Population and individual admixture proportion analysis
Estimated population admixture proportions of the contemporary Stege Nor population, using the method by Wang (2003), resulted in 99.7% (95% CI 94.3–100) of the present population being of indigenous origin. Only 0.3% was ascribed to exogenous origin.
The number of populations estimated using STRUCTURE showed that the probability of the data representing four populations was virtually 1 (data not shown). Consequently, the estimates of individual admixture proportions (q) for the present pike population in Stege Nor were based on a model assuming four populations. The power of STRUCTURE for identifying admixed individuals (hybrids) was illustrated by simulating 150 individuals (Figure 2a). Individual admixture proportions (q) for simulated individuals and their 90% posterior probability intervals were ranked and plotted (Figure 2a). Values close to zero denote pike of exogenous origin, whereas q-values close to one represent individuals that are most likely originating from the indigenous Stege Nor population. Individuals with intermediate q-values are either hybrids between stocked and indigenous pike, or individuals with uninformative genotypes. ‘Uninformative genotypes’ denote individuals with alleles at all studied loci that are relatively common in all populations, a problem which makes it difficult to distinguish between admixed and nonadmixed individuals. The figure shows that both stocked and indigenous pike were correctly ‘assigned’ to their population of origin. For simulated hybrids, wide 90% probability intervals were observed, indicating low power for unambiguous identification of admixed individuals (Figure 2a). To further test the precision of STRUCTURE, two individuals representing pike stocked in 2002 originating from Lake Tissø were included in the analysis as ‘positive’ controls (Figure 2b). They were clearly assigned to the Lake Tissø population with very narrow 90% probability intervals, providing a clear illustration of the power to identify stocked individuals in the present population. Individual admixture proportions (q) and their 90% posterior probability intervals were plotted for the 49 contemporary pike from Stege Nor (Figure 2b). In all, 45 of the 49 individuals sampled during 2002 in Stege Nor were most likely descendents of the indigenous population, whereas the remaining four individuals were either hybrids or individuals with uninformative genotypes (Figure 2b). The very wide 90% probability intervals for these four individuals make it difficult to unequivocally determine their population of origin based on individual admixture proportions.
Plots of individual admixture proportions (q) including 90% probability interval. Individuals are ranked from lowest to highest q-value. q-Values close to zero denote stocked pike, whereas pike with q-values close to 1 denote indigenous pike. Individuals with intermediate values are possible ‘hybrids’ between stocked and indigenous pike. (a) Illustration of the power of STRUCTURE to identify indigenous, stocked and admixed individuals. In all, 150 individuals were simulated, consisting of 50 simulated stocked pike from Lake Tissø (TS), followed by 50 individuals that are simulated F1 ‘hybrids’ between stocked and indigenous pike, and finally 50 simulated indigenous pike. (b) Individual admixture proportions of the 49 pike captured in Stege Nor in 2002 and two positive controls from the stocking event in 2002 (denoted by a circle).
Effective population size (Ne)
Estimates of effective population size were calculated based on the two temporal samples from the Stege Nor population. Assuming a Ne-MAX of 1000 and 12.9 generations between samples resulted in an estimated Ne of 523 (90% probability interval 256–935) using TM3. The corresponding result using MLNE2 assuming a Ne-MAX of 1000 and 12.9 generations between samples was 942 (95% CI 344–992). Assuming an Ne for the Stege Nor population corresponding to the lower percentile and median (approx. 250–500) estimated using TM3 and assuming a mutation rate of 10−5–10−3, the method by Garza and Williamson (2001) suggested that a significant reduction in population size had occurred in both the present population, ST02 and in the historical sample ST57 (see Table 3). Moreover, significant reductions were also suggested to have occurred in the three populations used for stocking (Table 3).
Discussion
Genetic differentiation among pike populations
We found no significant differentiation between the two temporal samples from the pike population in Stege Nor (Table 2). This indicates that the genetic composition of the population has not changed significantly during the past 45 years, despite the occurrence of at least two major inflows of highly saline water into Stege Nor during the period, causing a massive reduction in the pike population. The observation of temporal stability in the pike population in Stege Nor corresponds with previous studies on temporal stability of both trout and salmon populations over time scales as long as 80–100 years (Nielsen et al, 1997; Tessier and Bernatchez, 1999; Hansen et al, 2002; Hansen, 2002). In addition, we found a high degree of genetic differentiation among all pike populations studied, which corresponds with previous studies on pike populations in Denmark and around the Finnish part of the Baltic Sea (Hansen et al, 1999; Senanan and Kapuscinski, 2000; Jakobsen et al, 2005). These observations reflect the evolutionary history of the area and the relatively small effective population sizes within individual lakes (Miller and Kapuscinski, 1997). As a result, the small lake populations in Denmark may have drifted considerably in the time period after postglacial recolonization and subsequent isolation approx. 8000–10 000 years ago (Hansen et al, 1999).
Genetic effects of stocking
The present Stege Nor population consists almost exclusively of indigenous individuals. The results of population level admixture analysis using LEADMIX (Wang, 2003) showed that 99.7% of the contemporary gene pool was most likely of indigenous origin. Moreover, the individual admixture analyses conducted using STRUCTURE (Pritchard et al, 2000) showed that it is unlikely that any of the pike caught in Stege Nor 2002 were stocked individuals. However, this analysis cannot entirely rule out the possibility that a few individuals are possible descendants of stocked exogenous pike, hybridized with indigenous pike. Based on our results, we conclude that stocked pike have performed very poorly in Stege Nor and few, if any, stocked pike are present in the population. It could be argued that the low stocking success was the result of poor performance of stocked pike in just one or a few years. However, the result of the population level admixture analysis represents both past and current gene flow from stocked pike to the indigenous population. Furthermore, different sized pike representing several cohorts were included in the contemporary Stege Nor sample. Therefore, we rule out that the very low genetic contribution by stocked pike to the contemporary Stege Nor sample is due to a failure of stocking in just one or a few years. More than one million pike fry have been stocked in brackish waters in Denmark during the last decade, and the present results raise the possibility that all stocking attempts have failed in a similar way as observed for the stocking activity in Stege Nor.
Previous studies have shown poor survival rates of stocked individuals when stocking was conducted using fish that had been maintained in a hatchery environment for several generations and consequently may have been subject to domestication selection (eg Hindar et al, 1991; Nielsen et al, 1997; Hansen et al, 2001; Hansen, 2002). In the present study, where the stocked fish were of wild origin, it is difficult to point out the precise factors underlying their poor performance. Reduced performance of stocked pike can reflect lack of local adaptations, either physiologically or behaviourally. We assume that the main problem for stocked pike in Stege Nor is the change in environmental conditions between their natural habitat in freshwater and the brackish water that they were stocked into, causing a high rate of osmotic stress. However, nongenetic factors associated with stocking activity, such as timing of release, size of the released fish and proportion of released fish in the total population of new recruits, may also be important for the apparent lack of success of released hatchery-reared pike fry. Unpublished results from the Danish Institute for Fisheries Research on survival of pike after stocking in Stege Nor indicates survival rates of approx. 80% after 10 days of exposure in enclosures (L Jacobsen, personal communication). To our knowledge, there are no published studies on the long-term effects of this type of change of environment experienced by the stocked pike, but similar effects have been studied in salmonid fishes. Thus, Järvi (1990) and Järvi and Uglem (1993) showed that transfer of Atlantic salmon (Salmo salar) from freshwater to sea water was associated with high mortality. This mortality has been ascribed to stress from a combination of predator activity and problems with osmotic regulation (eg Järvi, 1990; Järvi and Uglem, 1993). Furthermore, Westin and Limburg (2002) demonstrated that eggs from female pike collected in freshwater rivers did not hatch at salinities exceeding 6‰, whereas this did not affect resident brackish pike. With an average annual salinity of 9–10‰ in Stege Nor, this may also affect the eggs of freshwater pike that have been stocked into brackish waters, if they survive to age of reproduction and attempt to reproduce in the fjord. Consequently, osmotic stress due to the higher salinity in Stege Nor may affect both survival and possible reproductive success of stocked pike, thus making environmental conditions in the area unsuitable for freshwater pike.
Population declines
Despite results presented by M_P_VAL showing significant bottlenecks, we observed a relatively high effective population size of approx. 250–940 in Stege Nor. Several factors (high Ne and temporal stability) indicate that Ne was high during most generations, while the method developed by Garza and Williamson (2001) indicates population declines in both temporal samples from Stege Nor. These conflicting results may indicate a large historical effective population size that has decreased drastically to a current low effective population size. However, significant bottlenecks were also implied for the three populations used for stocking. This is a surprising outcome and cannot be explained by the parameters assumed in the tests of the significance of M; the numerical values of M are in all cases low and considerably below 0.8, the smallest value observed by Garza and Williamson (2001) among data sets from large, stable populations. Founder events or ancient bottlenecks could, however, provide an explanation for the low M-values. Simulations by Garza and Williamson (2001) show that if a population experiences a drastic decline and remains small, then it may take several hundreds of generations for M to recover. Two other recent studies of European pike populations, based on microsatellite analysis (Jakobsen et al, 2005) and sequencing of the mitochondrial DNA d-loop (Maes et al, 2003), suggest that northern pike have experienced one or more bottlenecks either before establishing its current distribution in Europe or as a result of founder events in connection with postglacial recolonization. It would therefore be of considerable interest to analyse microsatellite DNA variation in more populations from Europe to see if low M-values are a general finding, thereby providing further support for historical bottlenecks within the species.
Conclusion
In summary, our results show that one decade of stocking with exogenous pike has not significantly affected the indigenous gene pool of the Stege Nor pike population. The stocking programme has had surprisingly little effect, despite the large number of stocked fry. In addition, the study reveals a considerable temporal stability of genetic composition and a relatively high effective population size of the Stege Nor population during the last 45 years, despite incidences of high mortality caused by several saltwater inflows. Given the poor performance of exogenous stocked pike, we recommend that future stocking should be conducted by supportive breeding, using offspring of locally caught pike and taking into account that the effective number of breeders should be sufficiently high to avoid a lowering of the total effective population size (Ryman and Laikre, 1991).
References
Allendorf FW, Leary RF, Spruell P, Wenburg JK (2001). The problems with hybrids: setting conservation guidelines. Trends Ecol Evol 16: 613–622.
Beaumont MA, Barratt EM, Gottelli D, Kitchener AC, Daniels MJ, Pritchard JK (2001). Genetic diversity and introgression in the Scottish wildcat. Mol Ecol 10: 319–336.
Berthier P, Beaumont MA, Cornuet J-M, Luikart G (2002). Likelihood-based estimation of the effective population size using temporal changes in allele frequencies: a genealogical approach. Genetics 160: 741–751.
Brzuzan P, Lucxynski M, Kuzniar PA (1998). Mitochondrial DNA variation in two samples of northern pike, Esox lucius L. Aquacult Res 29: 521–526.
Dahl J (1961). Age and growth in Danish and Swedish brackish pike (In Danish: Alder og vækst hos danske og svenske brakvandsgedder). Ferskvandsfiskeribladet 2: 34–38.
Estoup A, Largiader CR, Perrot E, Chourrout D (1996). Rapid one-tupe DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol Mar Biol Biotechnol 5: 295–298.
Garza JC, Williamson EG (2001). Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10: 305–318.
Goudet J (1995). Fstat version 1.2: a computer program to calculate F-statistics. J Heredity 86: 485–486.
Guo SW, Thompson EA (1992). Performing the exact test of Hardy–Weinberg proportion for multiple alleles. Biometrics 48: 359.
Hansen MM (2002). Estimating the long-term effects of stocking domesticated trout into wild brown trout (Salmo trutta) populations: an approach using microsatellite DNA analysis of historical and contemporary samples. Mol Ecol 11: 1003–1015.
Hansen MM, Ruzzante DE, Nielsen EE, Bekkevold D, Mensberg K-LD (2002). Long-term effective population sizes, temporal stability of genetic composition and potential for local adaptation in anadromous brown trout (Salmo trutta) populations. Mol Ecol 11: 2523–2535.
Hansen MM, Ruzzante DE, Nielsen EE, Mensberg K-LD (2001). Brown trout (Salmo trutta) stocking impact assessment using microsatellite DNA markers. Ecol Appl 11: 148–160.
Hansen MM, Taggart JB, Meldrup D (1999). Development of new VNTR markers for pike and assessment of variability at di- and tetranucleotide repeat microsatellite loci. J Fish Biol 55: 183–188.
Healy JA, Mulcahy MF (1980). A biochemical genetic analysis of populations of northern pike, Esox lucius L., from Europe and North America. J Fish Biol 17: 317–324.
Hill WG (1979). A note on effective population size with overlapping generations. Genetics 92: 317–322.
Hindar K, Ryman N, Utter F (1991). Genetic effects of cultured fish on natural fish populations. Canad J Fish Aquat Sci 48: 945–957.
Højgaard B (1985). Possibilities for Using Northern Pike (Esox lucius L.) for Aquaculture and Sea Ranching in the Bøgestrømmen Area. (In Danish: Muligheder for at anvende gedden (Esox lucius L.) som akvakulturfisk og til searanching i Bøgestrømsområdet). MSc Thesis, University of Aarhus.
Jakobsen BH, Hansen MM, Loeschcke V (2005). Microsatellite DNA analysis of northern pike (Esox lucius L.) populations: insights into the genetic structure and demographical history of a genetically depauperate species. Biol J Linn Soc 84: 91–101.
Järvi T (1990). Cumulative acute physiological stress in Atlantic salmon smolts: the effect of osmotic imbalance and the presence of predators. Aquaculture 89: 337–350.
Järvi T, Uglem I (1993). Predator training improves the anti-predator behaviour of hatchery reared Atlantic salmon (Salmo salar) smolt. Nord J Freshwater Resour 68: 63–71.
Levin PS, Williams JG (2002). Interspecific effects of artificially propagated fish: an additional conservation risk for salmon. Conser Biol 16: 1581–1587.
Maes GE, Van Houdt JKJ, De Charleroy D, Volckaert FAM (2003). Indications for a recent Holarctic expansion of pike based on a preliminary study of mtDNA variation. J Fish Biol 63: 254–259.
Miller CR, Joyce P, Waits LP (2002). Assessing allelic dropout and genotype reliability using maximum likelihood. Genetics 160: 357–366.
Miller LM, Kallemeyn L, Senanan W (2001). Spawning-site and natal-site fidelity by northern pike in a large lake: mark-recapture and genetic evidence. Trans Am Fish Soc 130: 307–316.
Miller LM, Kapuscinski AR (1996). Microsatellite DNA markers reveal new levels of genetic variation in northern pike. Tran Am Fish Soc 125: 971–977.
Miller LM, Kapuscinski AR (1997). Historical analysis of genetic variation reveals low effective populations size in a northern pike (Esox lucius) population. Genetics 147: 1249–1258.
Nielsen EE, Hansen MM, Bach LA (2001). Looking for a needle in a haystack: discovery of indigenous Atlantic salmon (Salmo salar L.) in stocked populations. Conserv Genet 2: 219–232 (www.evalife.dk/applications/hybridlab).
Nielsen EE, Hansen MM, Loeschcke V (1997). Analysis of microsatellite DNA from old scale samples of Atlantic salmon: a comparison of genetic composition over sixty years. Mol Ecol 6: 487–492.
Nielsen EE, Hansen MM, Loeschcke V (1999). Analysis of DNA from old scale samples: technical aspects, application and perspectives for conservation. Hereditas 130: 265–276.
Pritchard JK, Stephens M, Donnelly P (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Raat AJP (1988). Synopsis of biological data on the northern pike Esox lucius Linnaeus, 1758. Fao Fisheries Synopsis 30, Fao Rome.
Randi E, Lucchini V (2002). Detecting rare introgression of domestic dog genes into wild wolf (Canis lupus) populations by Bayesian admixture analyses of microsatellite variation. Conserv Genet 3: 31–45.
Raymond M, Rousset F (1995). GENEPOP (Version 1.2): a population genetics software for exact tests and ecumenicism. J Heredity 86: 248–249.
Rice WR (1989). Analysing tables of statistical tests. Evolution 43: 223–225.
Roy MS, Geffen E, Smith D, Ostrander EA, Wayne RK (1994). Patterns of differentiation and hybridization in North American wolf like canids, revealed by analysis of microsatellite loci. Mol Biol Evol 11: 553–570.
Ryman N, Laikre L (1991). Effects of supportive breeding on the genetically effective population size. Conserv Biol 5: 325–329.
Seeb JE, Seeb LW, Oates DW, Utter FM (1987). Genetic variation and postglacial dispersal of populations of northern pike (Esox lucius) in North America. Canad J Fish Aquat Sci 44: 556–561.
Senanan W, Kapuscinski AR (2000). Genetic relationships among populations of northern pike (Esox lucius). Canad J Fish Aquat Sci 57: 391–404.
Tessier N, Bernatchez L (1999). Stability of population structure and genetic diversity across generations assessed by microsatellites among sympatric populations of landlocked Atlantic salmon (Salmo salar L.). Mol Ecol 8: 169–179.
Wang J (2003). Maximum-likelihood estimation of admixture proportions from genetic data. Genetics 164: 747–765.
Wang J, Whitlock MC (2003). Estimating effective population size and migration rates from genetic samples over space and time. Genetics 163: 429–446.
Waples RS (1991). Genetic interactions between hatchery and wild salmonids: lessons from the Pacific Northwest. Canad J Fish Aquat Sci 48: 124–133.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Westin L, Limburg KE (2002). Newly discovered reproductive isolation reveals sympatric populations of Esox lucius in the Baltic. J Fish Biol 61: 1647–1652.
Acknowledgements
We are grateful to two anonymous reviewers for useful comments on a previous version of the manuscript, Karen-Lise D Mensberg and Dorte Meldrup for technical assistance, Henrik Baktoft for graphical assistance, Lene Jacobsen and Christian Skov for many useful discussions on pike biology and several anonymous fishermen and fishery biologists for collecting contemporary samples and historical scale samples in 2002 and 1956–1957. Peter Foged Larsen acknowledges financial support from the WWF WorldWide Fund for Nature/Novo Nordisk Biodiversity Funds and the Saltbækvig Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Larsen, P., Hansen, M., Nielsen, E. et al. Stocking impact and temporal stability of genetic composition in a brackish northern pike population (Esox lucius L.), assessed using microsatellite DNA analysis of historical and contemporary samples. Heredity 95, 136–143 (2005). https://doi.org/10.1038/sj.hdy.6800671
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800671