Abstract
Spiranthes romanzoffiana Cham. is restricted in Europe to the British Isles, where it is recognised as a conservation priority species due to frequent extirpation of populations along with no evidence of seed set; vegetative reproduction has been invoked as the sole means of perpetuation and dispersal. To investigate the reproductive ecology of this species, 17 populations have been sampled for chloroplast microsatellites and amplified fragment length polymorphisms (AFLPs). These markers revealed a previously unsuspected genetic–geographic split in the species, which correlates with differences in patterns of within-population variation. Northern populations were fixed for one chloroplast haplotype but showed high levels of AFLP genotypic diversity consistent with sexual reproduction (proportion of genotypes distinguishable, PD=0.98). More southerly populations showed fixed differences from the northern populations in their chloroplast haplotype and for 10 AFLP markers. They harboured only 12 unique multilocus genotypes among 113 individuals from six populations (PD=0.11). These genotypes differed mostly by single bands, and none by more than 4/138 loci, with identical multilocus genotypes occurring in widely separated populations. This uniformity in southern populations is consistent with agamospermous or autogamous reproduction, and/or an extreme population bottleneck. Finally, the observed patterns of population differentiation in S. romanzoffiana are compared with other studies of orchids, revealing a wide range of values that belie recent contrasting published generalisations that claim that orchids show either higher, or lower, levels of population differentiation than other plant families.
Similar content being viewed by others
Introduction
Molecular markers are now routinely used to contribute towards programmes concerned with the conservation of endangered species. Conservation genetics research has proved valuable in assessing the extent and consequences of genetic erosion (Young et al, 1996; Saccheri et al, 1998), in clarifying taxonomic issues to provide guidelines on which entities to preferentially conserve (Soltis and Gitzendanner, 1998), and in gaining insights into modes of reproduction and patterns of contemporary and historical dispersal (Sork et al, 1999).
This latter topic, essentially using molecular markers to investigate reproductive ecology, is relevant to many species of conservation concern, as halting and reversing the decline of a given species ultimately requires successful reproduction and dispersal. The mode of reproduction and the spatial scales of effective dispersal impact on both management strategies of individual sites and the maintenance of networks of populations at the landscape level. In plant species that can reproduce both sexually (via pollen and seed) and asexually (via vegetative spread), it can be exceedingly difficult to gauge the relative importance of each mode from field observations alone. Likewise, physical tracking of pollen and seed dispersal is also problematical beyond extremely local scales. Polymorphic genetic markers are well suited to tackling these issues, and hence are increasingly being applied in a conservation context.
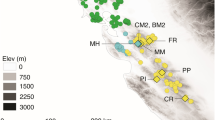
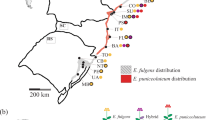
Spiranthes romanzoffiana (Irish Lady's-tresses) is a terrestrial, herbaceous, perennial orchid, widespread and relatively frequent in North America, yet confined to the western fringes of the British Isles within Europe (Preston et al, 2002; Figure 1). Its localised European distribution has led to increasing conservation interest; it is now listed in the UK Biodiversity Action Plans as a conservation priority species (UK Biodiversity Group, 1999), and recognised by the IUCN Orchid Specialist group as having ‘critically low populations’ in Europe (IUCN/Orchid Specialist Group, 1996).
Distribution of S. romanzoffiana in Europe. Closed triangles: 1987–1999 records. Open triangles: pre-1987 records (data from Preston et al, 2002). The map grid represents 100 km squares.
One of the major factors behind this conservation concern relates to the mode of reproduction of European populations of S. romanzoffiana. Pollination of this species in North America occurs via unspecialised, medium-sized long-tongued bees, which are attracted to nectar produced from two bosses near the base of the labellum. Similar potential pollinator species occur within the plant's habitat in Europe, and flower visits have been observed (J Robarts, 2002, personal communication). Despite this, no seed set has been recorded in the European range, leading to suggestions that the plant's persistence there is attributable to vegetative reproduction (Horsman, 1994). Plants produce lateral buds in the autumn, which over-winter and develop into the following year's aerial parts. The only documented mechanism for vegetative reproduction is the occasional production of twin (rarely three or four) lateral buds, which can lead to the formation of multiple aerial parts; these may eventually divide to form different ramets (Summerhayes, 1968). However, while this mechanism may lead to clonal spread over very small areas, it is difficult to imagine this mode of reproduction being effective over larger scales (eg >1 m).
A second area of conservation concern relates to the populations showing a high level of demographic instability; for instance, no population recorded in Scotland prior to 1981 is now known to be extant (UK Biodiversity Group, 1999). New populations have been recorded, but this frequent extirpation of populations leads to difficulties in assessing the distribution and conservation status of this species.
There is thus something of a paradox about European populations of S. romanzoffiana. On the one hand, populations appear to have a high turnover, with known populations disappearing and new populations being discovered. On the other hand, there is no satisfactory explanation for how the plant reproduces and disperses. Seed set has not been observed, and although vegetative reproduction has been invoked, the species' apparently limited capacity for clonal spread makes this an unlikely mode of the formation of new populations. We have used chloroplast microsatellites (cpSSRs; Provan et al, 2001) and amplified fragment length polymorphisms (AFLPs; Vos et al, 1995) to establish whether reproduction is primarily sexual or asexual, and to examine population differentiation in order to infer the spatial scales over which effective gene flow is occurring.
Materials and methods
Plant material and isolation of genomic DNA
To assess genetic variation in European populations of S. romanzoffiana, individuals were sampled from 17 populations: eight from Scotland and nine from Ireland (Table 1; see also Figure 2a). Sampling was designed to assess variation at the within-population, among-population and regional levels. In 10 populations, between 10 and 25 individuals were sampled and examined for both AFLP and cpSSR variation. While this sampling primarily consisted of individuals separated by at least 5 m, in eight of the populations five individuals were sampled within a 1 m2 quadrat to look for evidence of local clonal growth. From a further seven populations, between four and seven individuals were sampled for cpSSR analyses only (Table 1). For all molecular analyses, a section of leaf material approximately 30 mm long was harvested from single leaves and stored in silica gel.
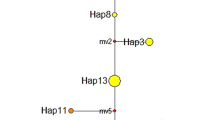
(a) Distribution map showing the location of the sampled populations. Numbers correspond to the population names in Table 1, and symbols correspond to different chloroplast haplotypes (squares A20, southern group; triangles A19, northern group). The map grid represents 100 km squares. (b) Sample electropherogram and partial sequences showing cpSSR variability. The peak traces correspond to the difference in poly-A repeat number.
DNA isolation was performed with DNeasy® Plant Mini Kits (Qiagen Ltd, UK). DNA quality and concentration was assessed by running samples alongside Hyperladder concentration marker (Bioline, UK) on a 1% agarose gel in 1 × tris borate/EDTA buffer, and visualised by staining with ethidium bromide.
Chloroplast microsatellite analysis
Five cpDNA regions (Table 2) were screened for microsatellite loci among 11 accessions representing a range of populations. Amplification of chloroplast regions was performed via polymerase chain reaction (PCR) on an MJ Research PTC-200 DNA Engine thermal cycler in 50 μl reactions containing 5 μl 10 × NH4 reaction buffer (Bioline, UK), 5 μl 2 mM deoxynucleotide triphosphate (dNTP), 2.5 μl 50 mM MgCl2, 1.5 μl of each primer (10 mM), 33.5 μl dH2O, 1.25 U Biotaq polymerase (Bioline, UK) and 1.0 μl DNA template. The thermocycle profile was: initial denaturation for 4 min at 94°C, followed by 30 cycles of 45 s at 92°C, 45 s at 53–62°C (Table 2) and 3 min at 72°C, with a final extension for 10 min at 72°C. PCR products were run on 1% agarose gels to check for amplification success and quality.
Amplified fragments were purified using QIAquick™ PCR purification kits (Qiagen Ltd, UK) following the manufacturer's protocol, and sequenced using the dideoxy chain-termination method. Cycle sequencing was performed in 20 μl reactions containing 4 μl Thermo Sequenase II (Amersham Pharmacia, UK), 0.5 μl primer (10 mM), 13.5 μl dH2O and 2 μl of purified PCR product, under the following PCR conditions: 25 cycles of 96°C for 10 s, 50°C for 5 s and 60°C for 4 min. Sequencing PCR products were purified following the manufacturer's instructions, then run on an ABI Prism™ 377 DNA Sequencer, before editing and manual alignment using Sequence Navigator™ 1.0.1 (Applied Biosystems Inc., USA).
Initial screening revealed eight mononucleotide repeat loci >8 bp in length, located in four of the five cpDNA regions amplified (atpB-rbcL (T)10, (T)14, (T)15, (A)20; psbC-trnS (T)9; trnH-trnK (A)13; trnL-trnF (T)17, (A)19 or 20). Underlined numbers indicate repeat motifs with single base-pair interruptions. Only the poly-A repeat located in the trnL intron showed any polymorphism among the 11 accessions studied. Forward and reverse primers were designed to amplify this microsatellite region (forward primer: 5′-GGTAACTTCCAAATTCAGA-3′; reverse primer: 5′-ACAGCTTCCGTTGAGTCTC-3′) using Primer 3 (www.basic.northwestern.edu/biotools/Primer3.html). Amplification was carried out in 10 μl reactions containing 0.5 μl of each of the two primers (10 mM), 1 μl 10 × NH4 reaction buffer, 0.5 μl 50 mM MgCl2, 1 μl 2 mM dNTPs, 6 μl dH2O, 0.2 U Biotaq polymerase and 0.5 μl DNA template. PCR conditions were as follows: 7 min at 95°C, followed by 30 cycles of 15 s at 95°C, 15 s at 50°C and 15 s at 72°C, with a final extension for 15 min at 72°C. Amplification products were diluted 130-fold in dH2O and analysed on 5% Long Ranger® acrylamide gels (BMA, USA) on an ABI Prism™ 377 DNA sequencer. Fragment size was determined by comparison with Rox 500 GeneScan size standard using Genotyper 2.0® software (Applied Biosystems Inc., USA).
Amplified fragment length polymorphism
Accessions were analysed following the plant mapping protocol of the PE Applied Biosystems AFLP™ kits. In all, 13 primer combinations were assessed for clarity, reproducibility and polymorphism among 12 accessions representing a range of populations, with three primer combinations selected for further analysis (A=EcoRI-ACT/MseI-CTA, B=EcoRI-ACT/MseI-CAG, C=EcoRI-ACA/MseI-CAG). Selective amplification products were run on 2% agarose gels to check for amplification success, and then electrophoresed on 5% Long Ranger® acrylamide gels; multilocus profiles were visualised using GeneScan Analysis® 3.1.2 software (Applied Biosystems Inc., USA). Markers were scored as present/absent using Genotyper® 2.0, and a binary data matrix constructed encompassing markers in the range 50–400 bp. Markers scoring below 50 arbitrary units of fluorescence were disregarded. Reproducibility of amplification was assessed by replicate extractions and amplifications of a subset of the samples. Only repeatable and unambiguous markers were included in the analysis.
Data analysis
Standard descriptive statistics were obtained to assess patterns of within-population variation: P, the proportion of loci that are polymorphic; PD, the proportion of distinguishable genotypes; APD, the average number of pairwise differences of markers among individuals. To establish the apportionment of variation into within- and between-population components, an analysis of molecular variation (AMOVA) was undertaken using Arlequin version 2.0 (Schneider et al, 2000), and the significance of FST analogues (φST) was assessed by permutation tests.
Results
cpSSRs
Polymorphism at the trnL microsatellite locus revealed two haplotypes differing by a single base pair (A19 or A20), resulting in fragments of 139 or 140 bp, respectively (Figure 2b). No intrapopulation variation was detected, but there was clear geographically structured interpopulation variation (Figure 2a). The A19 repeat occurred in all accessions from Coll, Vatersay and Barra (hereafter referred to as the northern group), and the A20 repeat occurred in all accessions from Colonsay and Ireland (hereafter referred to as the southern group).
AFLP analysis
A total of 138 unambiguous AFLP markers were generated from the 205 individuals that gave clear amplification profiles. Of these 138 markers, 86 were polymorphic (P=0.62) either within or among populations (Table 3).
Estimation of differentiation among all populations was high (φST=0.892, P<0.01). A hierarchical AMOVA analysis was conducted based on the geographical regions corresponding to the two chloroplast haplotypes (northern and southern groups). Between-group variation accounted for 85.8% of the total variation, with 3.5% of variation among populations within groups and 10.8% among individuals within populations. In total, 10 markers were differentially fixed between the northern and southern groups. Average pairwise distances among multilocus genotypes were also high when comparing populations from the two groups (Table 4).
The northern populations (Coll, Barra and Vatersay) showed lower levels of differentiation than that among all populations (φST=0.193), but differentiation was still significant (P<0.01). Little differentiation was detected among the southern populations (Colonsay and Ireland), although one population on Colonsay showed a fixed difference for a single marker in all individuals from the other southern populations. Obtaining meaningful estimates of population differentiation within the southern group was hampered by the paucity of polymorphism within and among these populations (see below).
There is a striking difference in the levels of intrapopulation diversity in northern vs southern populations (Table 3). The vast majority (90/92) of individuals from the northern populations harboured unique multilocus genotypes (PD=0.98), the only exception being a pair of accessions from Vatersay, and shared multilocus genotypes among single accessions from Vatersay and Barra (SC-BA1). All multilocus genotypes were unique to single populations except this latter pair. All 20 samples collected within 1 m2 quadrats (five plants from each of four northern populations) had different multilocus genotypes, consistent with sexual reproduction rather than local clonal spread. In contrast, only 12 unique multilocus genotypes among 113 individuals (PD=0.11) were detected from the southern populations, most being present only at low frequencies. The majority of these multilocus genotypes differed from each other by only a single marker (APD=0.37, range 0–4). Artefactual differences attributable to subtle variations in AFLP reaction conditions cannot be excluded as the source of these differences.
Discussion
Regional geographic structure
Both AFLPs and cpSSRs provide strong evidence for a marked genetic–geographic divide in European populations of S. romanzoffiana. Differential fixation of chloroplast haplotypes and 10 AFLP markers suggests a lack of contemporary gene flow among the northern (Coll, Barra, Vatersay) and more southerly (Colonsay, Ireland) populations. This split was previously unsuspected, and does not correlate with any documented morphological or ecological differences between these population groups.
Within-region population genetic structure (northern populations)
The northern populations (Barra, Coll, Vatersay) show a moderate degree of population differentiation (φST=0.193) but high levels of intrapopulation genotypic diversity. Although seed set has not been observed in European populations of S. romanzoffiana, the genotypic diversity in these northern populations suggests that seed set and sexual recruitment has taken place. The lack of individuals with shared multilocus genotypes (even within 1 m2 quadrats) indicates that vegetative reproduction does not play a major role in the perpetuation and dispersal of individuals.
It could be argued that the high levels of genotypic diversity, coupled with an absence of observed seed set, could be attributable to trans-atlantic seed dispersal (seed set is frequently observed in North American populations). We feel that this is exceedingly unlikely given (a) the considerable number of distinct genotypes and plants involved, and (b) the marked regional differentiation, which would require differential migration of seeds with the different chloroplast haplotypes into the northern and southern regions.
Instead, the most parsimonious explanation of our data is that sexual reproduction has taken place in situ. This reproduction could be historical, or current-but-infrequent. It is difficult to distinguish between these two scenarios, but it is worth noting that an absence of observed seed set does not necessarily equate to a genuine absence of seed set. Sexual reproduction could take place via occasional overlooked fertilised flowers; a typical individual terrestrial orchid flower can produce 1000–10 000 seeds (Arditti, 1992). Another potential theory is that a low level of seed set occurs within individual capsules, such that the capsules do not swell and appear fertile, but nevertheless contain a few viable seeds (J Robarts, 2002; M Light, 2003, personal communication). Detailed observations of flowering spikes and capsules, along with pollen viability, seed viability and crossing experiments, are needed to establish the precise conditions under which reproduction occurs.
Until such studies are completed, grazing controls should be established in these northern populations of S. romanzoffiana during the flowering and fruiting period. Sheep can have devastating effects on S. romanzoffiana flower spikes, and entire populations can be stripped of all flowering spikes in a matter of hours or days when flocks are moved into the orchid-containing pastures. As these populations were believed to reproduce entirely vegetatively (Horsman, 1994), the reproductive consequences of this grazing have been considered to be small. However, as the genetic data from this study suggest that reproduction is predominantly sexual, and yet field observations suggest that seed set is rare, the flowering spikes should be protected from herbivory to maximise the chances of successful seed production and periodic sexual recruitment.
Within-region population genetic structure (southern populations)
In contrast to the data from the northern populations, the more southerly populations are typified by extremely low levels of genotypic diversity, with identical multilocus genotypes being detected in geographically disparate populations. Such genetic uniformity within and among populations could be attributed to efficient clonal growth and dispersal. In clonal plants, if propagules from a single genotype are able to disperse, large areas can be covered by ramets of a single genet (eg Hollingsworth and Bailey, 2000). However, given the lack of evidence for clonal spread in the morphologically similar northern populations, this seems an unsatisfactory explanation. This is particularly evident when one considers the substantial geographical distances involved.
Agamospermy has been reported in some Spiranthes species from North America (Catling, 1982; Schmidt and Antlfinger, 1992). Although this has never been observed in S. romanzoffiana, several species that usually reproduce via sexual out-crossing have been observed to develop agamospermous races in colonising populations near the edge of their ranges (Catling and Catling, 1991). This offers a potential explanation for the patterns of genotypic similarity among populations in Ireland and Colonsay.
An alternative explanation is that reproduction in the southern populations is sexual, but that the genotypes involved are homozygous at the vast majority of the study loci. An extreme genetic bottleneck could lead to genetic uniformity among sexually reproducing individuals. Alternatively, self-pollination can lead to a rapid reduction in heterozygosity. Autogamy has been observed in other Spiranthes species (eg Catling, 1983a, 1990; Sipes and Tepedino, 1995; Sun, 1997). However, Catling's (1983b) observations on North American populations of S. romanzoffiana suggest that it is unlikely in this species. This is due to protandry, and also the column structure which prevents the pollinia from coming into contact with the stigmatic surface.
Data from variable codominant markers (eg nuclear microsatellites) are needed to distinguish between these alternative explanations. Hypervariable codominant markers could establish whether the observed genetic uniformity is associated with homozygosity (selfing), or random mating from a very narrow genetic base (bottleneck), or whether there is fixed heterozygosity (indicative of agamospermous reproduction).
Population differentiation
The overall estimate of population differentiation for European populations of S. romanzoffiana is high (φST=0.892). However, this figure includes two groups of populations fixed for different markers (apparently not exchanging genes), and potentially showing different reproductive strategies. In terms of evaluating the genetic connectivity of populations, it is perhaps more meaningful to examine patterns of differentiation among the sexual northern populations. Here, there is a smaller, but still significant, level of population differentiation (φST=0.193). It is also worth noting that if one considers pairwise differences among northern populations, an estimate of φST (0.217) is obtained between the two populations on Barra (separated by only 0.5 km), which is a similar order of magnitude to the estimate for differentiation among the northern populations per se (involving interisland distances of 70 km between Coll and Barra). This differentiation, evident even over small scales, may be related to the infrequent production of seeds, which will inevitably reduce opportunities for interpopulation gene flow.
This significant deviation from panmixia among proximal populations of S. romanzoffiana could be considered high when compared to average levels of differentiation among orchid populations. Hamrick and Godt (1996) summarised allozyme data from 16 orchid population genetic studies and obtained a mean estimate of GST for orchids of 0.087. They noted that orchids ‘had an exceptionally low mean GST… perhaps due to the species-specific pollinators characteristic of orchids, and to their tiny wind-borne seeds. Both of these traits could produce high rates of gene flow among populations’ (Hamrick and Godt, 1996, p 1297). However, it is worth stressing that allozyme work by Sun and Wong (2001, p 2186) led to exactly the opposite conclusion, that ‘gene flow appears to be much more restricted in wild orchids than in other plants’.
These opposing views triggered our own review of levels of differentiation among orchid populations (Table 5). To the best of our knowledge, this is the most comprehensive summary of orchid GST (or FST) estimates to date. Our estimates of population differentiation range from GST=0.012 to 0.924, with mean values of GST=0.187 (all studies), GST=0.219 (excluding studies examining less than five populations) and GST=0.161 (further excluding studies examining dominant marker data sets). These mean estimates are somewhat higher than the ‘exceptionally low mean GST’ of 0.087, and within the range recorded for several other plant families by Hamrick and Godt (1996). In this respect it is worth noting that while orchid species share biological attributes such as small, mostly dust-like seeds, they are also a large, heterogeneous and diverse group (Arditti, 1992; Rudall and Bateman, 2002). The Orchidaceae contains ca 19 000 species encompassing an exceptionally wide range of reproductive strategies, variation in generation times, and includes both narrow endemic and widespread species, as well as species with relatively continuously distributed ranges and species with only isolated populations. As such, they should be expected to show, and do show, considerable variation in the levels of population differentiation.
References
Aagaard JE, Harrod RJ, Shea KL (1999). Genetic variation among populations of the rare Clustered Lady-slipper orchid (Cypripedium fasciculatum) from Washington State, USA. Nat Areas J 19: 234–238.
Ackerman JD, Ward S (1999). Genetic variation in a wide-spread, epiphytic orchid: where is the evolutionary potential? Syst Bot 24: 282–291.
Alexandersson R, Ågren J (2000). Genetic structure in the nonrewarding, bumblebee-pollinated orchid Calypso bulbosa. Heredity 85: 401–409.
Arditti J (1992). Fundamentals of Orchid Biology, John Wiley and Sons: New York.
Arduino P, Cianchi R, Rossi W, Corrias B, Bullini L (1995). Genetic variation in Orchis papilionacea (Orchidaceae) from the central Mediterranean region: taxonomic inferences at the intra-specific level. Plant Syst Evol 194: 9–23.
Arduino P, Verra F, Cianchi R, Rossi W, Corrias B, Bullini L (1996). Genetic variation and natural hybridization between Orchis laxiflora and Orchis palustris (Orchidaceae). Plant Syst Evol 202: 87–109.
Arft AM, Ranker TA (1998). Allopolyploid origin and population genetics of the rare orchid Spiranthes diluvialis. Am J Bot 85: 110–122.
Bateman RM, Pridgeon AM, Chase MW (1997). Phylogenetics of subtribe Orchidinae (Orchidoideae, Orchidaceae) based on nuclear ITS sequences. 2. Infrageneric relationships and reclassification to achieve monophyly of Orchis sensu stricto. Lindleyana 12: 113–141.
Borba EL, Felix JM, Solferini VN, Semir J (2001). Fly-pollinated Pleurothallis (Orchidaceae) species have high genetic variability: evidence from isozyme markers. Am J Bot 88: 419–428.
Brzosko E, Ratkiewicz M, Wroblewska A (2002). Allozyme differentiation and genetic structure of the Lady's slipper (Cypripedium calceolus) island populations in north-east Poland. Bot J Linn Soc Lond 138: 433–440.
Bullini L, Cianchi R, Arduino P, De Bonis L, Mosco MC, Verardi A et al (2001). Molecular evidence for allopolyploid speciation and a single origin of the western Meditteranean orchid Dactylorhiza insularis (Orchidaceae). Biol J Linn Soc Lond 72: 193–201.
Case MA (1993). High levels of allozyme variation within Cypripedium calceolus (Orchidaceae) and low levels of divergence among its varieties. Syst Bot 18: 663–677.
Case MA (1994). Extensive variation in the levels of genetic diversity and degree of relatedness among five species of Cypripedium (Orchidaceae). Am J Bot 81: 175–184.
Case MA, Mlodozeniec HT, Wallace LE, Weldy TW (1998). Conservation genetics and taxonomic status of the rare Kentucky Lady's Slipper: Cypripedium kentuckiense (Orchidaceae). Am J Bot 85: 1779–1786.
Catling PM (1982). Breeding systems of north-eastern North American Spiranthes (Orchidaceae). Can J Bot 60: 3017–3039.
Catling PM (1983a). Autogamy in eastern Canadian Orchidaceae: a review of current knowledge and some new observations. Nat Can 110: 37–53.
Catling PM (1983b). Pollination of north-eastern North American Spiranthes (Orchidaceae). Can J Bot 61: 1080–1093.
Catling PM (1990). Auto-pollination in the Orchidaceae. In: Arditti J (ed) Orchid Biology: Reviews and Perspectives, Timber Press: Portland Vol 5, pp 121–158.
Catling PM, Catling VR (1991). A synopsis of breeding systems and pollination in North American orchids. Lindleyana 6: 187–210.
Chiang T-Y, Schaal BA, Peng C-I (1998). Universal primers for amplification and sequencing a non-coding spacer between the atpB and rbcL genes of chloroplast DNA. Bot Bull Acad Sin 39: 245–250.
Chung MY, Chung MG (2000). Allozyme diversity in populations of Cymbidium goeringii (Orchidaceae). Plant Biol 2: 77–82.
Corrias B, Rossi W, Arduino P, Cianchi R, Bullini L (1991). Orchis longicornu Poiret in Sardinia: genetic, morphological and chorological data. Webbia 45: 71–101.
Culley TM, Wallace LE, Gengler-Nowak KM, Crawford DJ (2002). A comparison of two methods of calculating G ST, a genetic measure of population differentiation. Am J Bot 89: 460–465.
Demesure B, Sodzi N, Petit RJ (1995). A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol Ecol 4: 129–131.
Ehlers BK, Pedersen H (2000). Genetic variation in three species of Epipactis (Orchidaceae): geographic scale and evolutionary inferences. Biol J Linn Soc Lond 69: 411–430.
Gustafsson S (2000). Patterns of genetic variation in Gymnadenia conopsea, the fragrant orchid. Mol Ecol 9: 1863–1872.
Hamrick JL, Godt MJW (1996). Effects of life history traits on genetic diversity in plant species. Phil Trans R Soc Lond B 351: 1291–1298.
Hedrén M, Klein E, Teppner H (2000). Evolution of polyploids in the European orchid genus Nigritella: evidence from allozyme data. Phyton-Ann Rei Bot 40: 239–275.
Hollingsworth ML, Bailey JP (2000). Evidence for massive clonal growth in the invasive weed Fallopia japonica (Japanese Knotweed). Bot J Linn Soc Lond 133: 463–472.
Hollingsworth PM, Dickson JH (1997). Genetic variation in rural and urban populations of Epipactis helleborine in Britain. Bot J Linn Soc Lond 123: 321–331.
Horsman F (1994). Spiranthes romanzoffiana. In: Stewart A, Pearman DA, Preston CD (eds) Scarce Plants in Britain, Joint Nature Conservancy Council: Peterborough p 398.
IUCN/SSC Orchid Specialist Group (1996). Orchids – Status Survey and Conservation Action Plan, IUCN: Gland, Switzerland and Cambridge, UK.
Machon N, Bardin P, Mazer SJ, Moret J, Godelle B, Austerlitz F (2003). Relationship between genetic structure and seed and pollen dispersal in the endangered orchid Spiranthes spiralis. New Phytol 157: 677–687.
Murren CJ (2003). Spatial and demographic population genetic structure in Catasetum viridiflavum across a human-disturbed habitat. J Evol Biol 16: 333–342.
Nielsen LR, Siegismund HR (1999). Interspecific differentiation and hybridization in Vanilla species (Orchidaceae). Heredity 83: 560–567.
Peakall R, Beattie AJ (1991). The genetic consequences of worker ant pollination in a self-compatible, clonal orchid. Evolution 45: 1837–1848.
Peakall R, Beattie AJ (1996). Ecological and genetic consequences of pollination by sexual deception in the orchid Caladenia tentactulata. Evolution 50: 2207–2220.
Peakall R, James SH (1995). Outcrossing in an ant pollinated clonal orchid. Heredity 62: 161–167.
Preston CD, Pearman DA, Dines TD (2002). New Atlas of the British and Irish Flora, Oxford University Press: Oxford.
Provan J, Powell W, Hollingsworth PM (2001). Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16: 142–147.
Reinhammar L-G, Hedrén M (1997). Allozyme differentiation between lowland and alpine populations of Pseudorchis albida s. lat. (Orchidaceae) in Sweden. Nord J Bot 18: 7–14.
Rossi W, Corrias B, Arduino P, Cianchi R, Bullini L (1992). Gene variation and gene flow in Orchis morio (Orchidaceae) from Italy. Plant Syst Evol 179: 43–58.
Rudall PJ, Bateman RM (2002). Roles of synorganisation, zygomorphy and heterotopy in floral evolution: the gynostemium and labellum of orchids and other lilioid monocots. Biol Rev 77: 403–441.
Saccheri I, Kuussaari M, Kankare M, Vikman P, Fortelius W, Hanski I (1998). Inbreeding and extinction in a butterfly metapopulation. Nature 392: 491–494.
Scacchi R, De Angelis G (1989). Isozyme polymorphism in Gymnadenia conopsea and its inferences for systematics within this species. Biochem Syst Ecol 17: 25–33.
Scacchi R, De Angelis G, Corbo RM (1991). Effect of the breeding system on the genetic structure in three Cephalanthera spp. (Orchidaceae). Plant Syst Evol 176: 53–61.
Scacchi R, De Angelis G, Lanzara P (1990). Allozyme variation among and within eleven Orchis species (fam. Orchidaceae), with special reference to hybridizing aptitude. Genetica 81: 143–150.
Scacchi R, Lanzara P, De Angelis G (1987). Study of electrophoretic variability in Epipactis helleborine (L.) Crantz, E. palustris (L.) Crantz and E. microphylla (Ehrh.) Swartz (Orchidaceae). Genetica 72: 217–224.
Schmidt JM, Antlfinger AE (1992). The level of agamospermy in a Nebraska population of Spiranthes cernua (Orchidaceae). Am J Bot 79: 501–507.
Schneider S, Roessli D, Excoffier L (2000). Arlequin 2.000: a software for population genetic data analysis, Genetics and Biometry Laboratory, University of Geneva, Switzerland.
Sharma IK, Clements MA, Jones DL (2000). Observations of high genetic variability in the endangered Australian terrestrial orchid Pterostylis gibbosa R. Br. (Orchidaceae). Biochem Syst Ecol 28: 651–663.
Sharma IK, Jones DL (1996). An electrophoretic study of variability in Diuris sulphurea R. Br. (Orchidaceae) in the Canberra region. J Orch Soc Ind 10: 19–24.
Sharma IK, Jones DL, French CJ (2002). Unusually high genetic variability revealed through allozymic polymorphism of an endemic and endangered Australian orchid, Pterostylis aff. picta (Orchidaceae). Biochem Syst Ecol 31: 513–526.
Sharma IK, Jones DL, Young AG, French CJ (2001). Genetic diversity and phylogenetic relatedness among six endemic Pterostylis species (Orchidaceae; series Grandiflorae) of western Australia, as revealed by allozyme polymorphisms. Biochem Syst Ecol 29: 697–710.
Sipes SD, Tepedino VJ (1995). Reproductive biology of the rare orchid Spiranthes diluvialis: breeding system, pollination, and implications for conservation. Conserv Biol 9: 929–938.
Soltis PS, Gitzendanner MA (1998). Molecular systematics and the conservation of rare species. Conserv Biol 13: 471–483.
Sork VL, Nason J, Campbell DR, Fernandez JF (1999). Landscape approaches to historical and contemporary gene flow in plants. Trends Ecol Evol 14: 219–223.
Squirrell J, Hollingsworth PM, Bateman RM, Dickson JH, Light MHS, MacConaill M et al (2001). Partitioning and diversity of nuclear and organelle markers in native and introduced populations of Epipactis helleborine (Orchidaceae). Am J Bot 88: 1409–1418.
Summerhayes VS (1968). Wild Orchids of Britain, 2nd edn. Collins: London.
Sun M (1996). Effects of population size, mating system, and evolutionary origin on genetic diversity in Spiranthes sinensis and S. hongkongensis. Conserv Biol 10: 785–795.
Sun M (1997). Genetic diversity in three colonising orchids with contrasting mating systems. Am J Bot 84: 224–232.
Sun M, Wong KC (2001). Genetic structure of three orchid species with contrasting mating systems using RAPD and allozyme markers. Am J Bot 88: 2180–2188.
Taberlet P, Gielly L, Pautou G, Bouvet J (1991). Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17: 1105–1109.
Thornhill AD (1999). Species-level and population-level patterns of genetic variation in the riparian orchid, Epipactis gigantea Dougl. ex Hook. (Orchidaceae) http://conbio.rice.edu/at/research/thesis_introduction.html.
Tremblay RL, Ackerman JD (2001). Gene flow and effective population size in Lepanthes (Orchidaceae): a case for genetic drift. Biol J Linn Soc Lond 72: 47–62.
UK Biodiversity Group (1999). Irish Lady's-tresses (Spiranthes romanzoffiana). In: Tranche 2 Action Plans, Volume 3, Plants and Fungi, English Nature: Peterborough pp 339–341.
Vos P, Hogers R, Bleeker M, Reijans M, Van De Lee T, Hornes M et al (1995). AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23: 4407–4414.
Wallace LE (2002). Examining the effects of fragmentation on genetic variation in Platanthera leucophaea (Orchidaceae): inferences from allozymes and random amplified polymorphic DNA markers. Plant Species Biol 17: 37–49.
Weicker JJ, Brumfield RT, Winker K (2001). Estimating the unbiased estimator θ for population genetic survey data. Evolution 55: 2601–2605.
Wong KC, Sun M (1999). Reproductive biology and conservation genetics of Goodyera procera (Orchidaceae). Am J Bot 86: 1406–1413.
Young A, Boyle T, Brown A (1996). The population genetic consequences of habitat fragmentation in plants. Trends Ecol Evol 11: 413–418.
Acknowledgements
We thank TGF Curtis, E Grant, RL Gulliver, M Gulliver, B Hamill, M Keirnen, J Robarts and M Wright for the collection of plant material. Technical assistance and advice was provided by A Ponge and J Squirrell. Useful discussion and comments were made by RL Gulliver, F Horsman, M Light, J Robarts and CJ Sheviak. This research was supported by NERC (Grant NER/S/M/2000/03993), the Newby Trust Ltd, the Oleg Polunin Memorial Fund, the Sidney Perry Foundation, Mr and Mrs B Higginson, and a bursary from the Royal Botanic Garden Edinburgh and University of Edinburgh. The Royal Botanic Garden Edinburgh is supported by the Scottish Executive Environment and Rural Affairs Department.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Forrest, A., Hollingsworth, M., Hollingsworth, P. et al. Population genetic structure in European populations of Spiranthes romanzoffiana set in the context of other genetic studies on orchids. Heredity 92, 218–227 (2004). https://doi.org/10.1038/sj.hdy.6800399
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800399
Keywords
This article is cited by
-
Spatial genetic structure in the Madeiran endemic Dactylorhiza foliosa (Orchidaceae)
Plant Systematics and Evolution (2022)
-
Systematics and conservation of British and Irish orchids: a "state of the union" assessment to accompany Atlas 2020
Kew Bulletin (2022)
-
Orchid conservation and research: An analysis of gaps and priorities for globally Red Listed species
Ambio (2020)
-
Pollen-mediated gene flow ensures connectivity among spatially discrete sub-populations of Phalaenopsis pulcherrima, a tropical food-deceptive orchid
BMC Plant Biology (2019)
-
Patterns of Genetic Diversity in Rare and Common Orchids Focusing on the Korean Peninsula: Implications for Conservation
The Botanical Review (2018)