Abstract
A general prediction of the neutral theory of evolution is that genetic diversity should correlate positively with effective population size. We show here that diversity across eight microsatellite loci was consistently and substantially lower in one common amphibian (Bufo bufo) than in another with similar life history traits (Rana temporaria) despite B. bufo having the larger breeding assemblage sizes. However, B. bufo breeding assemblages were much more highly differentiated than those of R. temporaria according to both Fst and Rst estimators. These differences occurred in shared habitats across identical geographical distances. The patterns of genetic diversity and differentiation detected in these two species were probably a consequence of high gene flow in R. temporaria but much lower gene flow among the larger but more dispersed B. bufo assemblages. These observations highlight the difficulty of defining the boundaries of wild populations, and show how two broadly similar species can exhibit very different population dynamics.
Similar content being viewed by others
Introduction
The importance of genetic diversity to the long-term viability of wild plant and animal populations is increasingly well recognised (eg Amos and Balmford, 2001; Hedrick, 2001). Neutral theory predicts that a positive relationship should exist between effective population size (Ne) and the extent of genetic variation (allelic diversity and heterozygosity) at loci not subject to strong selection (Kimura, 1983). This expectation has often been found in wild populations (Frankham, 1996) although census population sizes (Nc), or other surrogates of Ne such as biogeographical range, have usually been used in these analyses rather than Ne itself. This is because Ne is difficult to measure, and in most cases where comparisons have been possible Ne has proved much smaller than Nc (Frankham, 1995). Genetic diversity also shows interesting correlations with breeding systems, at least among plants (Hamrick and Godt, 1989). Outcrossing species exhibited, on average, significantly higher levels of genetic variation than those with predominantly selfing or mixed-mating systems. Furthermore, only 10% of diversity was partitioned among populations in wind-pollinated, outcrossing species, whereas more than 50% was partitioned in this way in selfing species. Population structure, as well as size, may therefore be important in the sustenance of neutral genetic diversity.
In natural situations, however, boundaries between populations are often indeterminate. Population structure is likely to be particularly important in this context. If what appear to be discrete populations are actually interconnected by high levels of gene flow, there may effectively be a single large population rather than multiple small ones. Understanding this distinction will be increasingly important for the conservation of isolated or fragmented populations of rare species. To assess the possible significance of this issue, we compared the population structures of two species of amphibians. This group of organisms is characterised by low individual mobility, and commonly by high genetic differentiation over relatively short geographical distances (eg Rowe et al, 2000). Furthermore, there is widespread concern about amphibian declines at many different locations in the world (Houlahan et al, 2000).
For our comparison we chose species in which many features that can affect genetic diversity are common to both. Thus the common toad Bufo bufo and the common frog Rana temporaria are widespread and abundant across much of Europe (Gasc, 1997). They share similar life histories, breeding systems and habitat requirements, and often reproduce in the same ponds (Beebee and Griffiths, 2000). Both species have generation times of 3–4 years, have similar fecundities (females typically lay between 1000–3000 eggs per year in the early spring) and experience similar annual survivorships of around 50%. However, in agricultural habitats (which account for much of the European landscape), B. bufo tends to occur in larger breeding assemblages but to use fewer of the available ponds than does R. temporaria (Cooke, 1975). Such breeding assemblages, where adults congregate and spawn in ponds, are often considered as discrete populations by conservation managers. We anticipated that this autecological distinction (few large assemblages versus many small) could have different consequences for genetic diversity. The extent to which this is true will obviously depend upon the degree of isolation between assemblages and the extent of gene flow between them (eg Scribner et al, 2001). Our hypothesis was that a breeding assemblage would define a discrete population more reliably in B. bufo than in R. temporaria.
Materials and methods
Sites and sampling
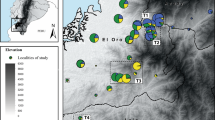
Seven assemblages of R. temporaria and B. bufo were sampled at a total of eight breeding sites. Six sites were common to both species, but because sampling at one site was interrupted by the foot and mouth disease outbreak, the seventh site was unique to one or other species (Figure 1). The sampling strategy involved two widely separated rural sites for each species in the north or east of Britain (Ainsdale for both species, Saltfleetby for B. bufo and Halesworth for R. temporaria) and five sites with assemblages of both species clustered relatively close together in Sussex, southern England. The Sussex sites included three urban locations within the city of Brighton (Crematorium, St Anne's and Withdean), and two rural locations close to Brighton (Pells and Whitelands). Samples from at least 20 separate spawn clumps (R. temporaria) or spawn strings (B. bufo) were collected at each site, with a view to maximising representation of the genetic diversity present in each assemblage. The eggs were allowed to develop in the laboratory, larvae (30–40 individuals from each site, representing all the sampled spawn) were harvested at stage 26 (Gosner, 1960) and stored in 70% ethanol prior to DNA extraction.
For the purposes of our analysis, we began by equating assemblage numbers with census population sizes. Census population estimates for all the sites were based on counts of spawn clumps (R. temporaria) or adult numbers at the breeding ponds (B. bufo) averaged over 1–5 years of intermittent observations since 1980. Since female R. temporaria spawn once each year, and the sex ratio for this species is around 1:1, doubling spawn clump numbers yields a reasonable estimate of Nc, adult population size (Beebee, 1996). Bufo bufo spawn is laid as intertwined strings that cannot be counted, so estimates for this species are based on numbers of adults assembled in ponds (counted using a powerful torch at night) at the peak of the breeding season. They are therefore minimum estimates of adult census population sizes. Estimates for both species using these methods are accurate at least to within an order of magnitude (Beebee, 1996). Indeed, those for frogs, based on spawn clump counts, have relatively low variance and are generally accurate to within ±10% (Griffiths et al, 1996). Variances for toad estimates have not been formally determined, but it is clear that the true adult census size for this species must always be higher than the adult counts. Density of breeding sites for both species in the Sussex study area was determined by field survey (Beebee, 1981).
Microsatellite analysis
DNA was extracted from larvae using the Chelex 100 protocol (Walsh et al, 1991). Microsatellite loci were amplified by PCR in the presence of [α33P]dATP and locus-specific primers previously developed for these species (Brede et al, 2001; Rowe and Beebee, 2001). A total of 10 polymorphic microsatellite loci (Bbufμ14, 15, 24, 39, 46, 47, 54, 62, 63, 65) were available for B. bufo and a further 10 (Rtempμ1-10) for R. temporaria. Both sets of microsatellites were dinucleotide repeats although two in B. bufo (Bbufμ14 and Bbufμ39) and three in R. temporaria (Rtempμ1, Rtempμ2 and Rtempμ7) had short interruptions within the repeat sequences. Neither mean repeat numbers nor mean numbers of alleles per locus in the microsatellites used for our genetic analyses were significantly different between the species (two-sample t with unequal variances=−1.3, df=7.6, P=0.2303 and –1.66, df=10.4, P=0.126, respectively). PCR products were electrophoresed alongside an M13 marker on standard sequencing gels (6% w/v polyacrylamide) and alleles scored after visualisation by autoradiography (Rowe et al, 1997).
Genetic analysis
Tests for Hardy–Weinberg equilibrium and linkage disequilibrium were performed using BIOSYS-1 (Swofford and Selander, 1981) and GENEPOP 3.1 (Raymond and Rousset, 1995), respectively. Genetic diversity measurements including mean number of alleles per locus, expected (He) and observed (Ho) heterozygosities were also carried out using BIOSYS-1. Partitioning of genetic variation within and among populations was assessed by analysis of molecular variance (AMOVA) using ARLEQUIN 1.1 (Schneider et al, 1997). Differentiation amongst populations was measured using Fst (Weir and Cockerham, 1984) and Rst (Goodman, 1997). Fst values and their significance levels as well as gene flow (Nm) estimates using the private alleles method (Slatkin, 1995) were estimated using GENEPOP 3.1 (Raymond and Rousset, 1995) and FSTAT 1.2 (Goudet, 1995). Rst (variance component) estimates were made using RSTCALC 2.1 (Goodman, 1997). Isolation by distance (Slatkin, 1993) was investigated for the Sussex populations using the ISOLDE subprogram within GENEPOP 3.1 using Fst/(1−Fst), and ln geographical distance values (Rousset, 1997). Geographic site-to-site distances for this analysis were measured directly from a 1:50 000 ordnance survey map. Population bottleneck events were investigated using BOTTLENECK 1.2.02 (Cornuet and Luikart, 1996). Two-phase mutation models in which the proportion of stepwise mutation (SMM) was set at 70 or 90% were employed in the bottleneck analysis.
Routine statistical tests of significance were performed using STATISTIX 7 (Analytical Software, Tallahasse, FL, USA) with either parametric (matched pair or two-sample t-tests, Pearson moment correlations) or nonparametric (Wilcoxon signed-rank or Wilcoxon rank-sum test or Spearman rank correlations) methods according to data distribution. Linear regression was also performed with STATISTIX 7 using arcsin-transformed estimates of He, Nm (which was normally distributed), and with log10-transformed estimates of population size. All transformations were checked for normal distribution of residuals by the Shapiro–Wilks test. Randomization tests were performed with the program RT 2.1 (Manly, 1997).
Results
Genetic diversity
Two loci for each species (Bbufμ24 and Bbufμ65, Rtempμ5 and Rtempμ6) deviated significantly from Hardy–Weinberg equilibrium in several populations and were excluded from subsequent analyses. These loci all had substantial heterozygote deficits, possibly as a result of widespread null alleles. At the species level (ie averaged across all populations), B. bufo had an estimated mean He of 0.579 with an average of 5.07 alleles per locus. R. temporaria had an estimated mean He of 0.669 with an average of 7.87 alleles per locus. Figure 2a and b show that individual R. temporaria populations consistently had greater genetic diversity than those of B. bufo. Rana averaged 17% higher He than Bufo and 58% higher allelic diversity across the seven populations of each species. Indeed, the frequency distributions of allelic diversities were mutually exclusive between the two anurans. Both mean heterozygosity (Mann–Whitney U=5, 44, P=0.015) and mean number of alleles per locus (Mann–Whitney U=0, 49, P=0.0022) were significantly higher in R. temporaria than in B. bufo. By contrast, mean census population sizes (Nc) averaged more than 10-fold higher for Bufo (at >103) than for Rana (at >102) as shown in Figure 2c.
Genetic diversity and census population size ranges of seven B. bufo and R. temporaria populations. (a) Frequency distributions of mean expected heterozygosities; (b) frequency distributions of mean allelic diversities; (c) frequency distributions of census population sizes. Solid bars, R. temporaria; open bars, B. bufo.
For B. bufo, there was almost a significant positive correlation between mean number of alleles and mean Nc (rs=0.7748, P=0.0532), but not for mean He and mean Nc rs=0.2091. With R. temporaria, correlations between Nc and genetic diversity were negative. For Nc and mean number of alleles, rs=−0.5406, while for Nc and mean He rs=−0.9550, P=0.0064. Although sample size (n=7) and therefore power was relatively low in this analysis, there was a clear indication that small frog populations were the most genetically diverse.
One population of B. bufo (Withdean) and one of R. temporaria (Whitelands) were significantly (P<0.05) out of Hardy–Weinberg equilibrium at three and four of the remaining eight loci, respectively. Of the remaining 48 loci × population assessments, only one in B. bufo and three in R. temporaria deviated significantly from Hardy–Weinberg equilibrium after Bonferroni correction for multiple tests using P=0.001. Of 168 pairwise combinations of loci, only one in B. bufo but 15 in R. temporaria (10 in a single population) showed evidence of linkage disequilibrium after Bonferroni correction using P=0.0003.
Genetic differentiation
AMOVA of all seven populations of each species indicated differences between frogs and toads in the partitioning of genetic variation. For B. bufo, 86.6% of variation across the eight microsatellite loci was within populations while 13.4% of variation was distributed among populations. By contrast, 100% of genetic variation in R. temporaria was estimated to be within populations.
We then compared levels of genetic differentiation among four Sussex populations, excluding the fifth Sussex populations of each species that deviated significantly from Hardy–Weinberg equilibrium at multiple loci. The more distant populations (Ainsdale, Saltfleetby and Halesworth) were excluded from this part of the analysis because gene flow estimates over large distances have little meaning for amphibians, with their low powers of dispersal. Reducing the number of populations lowered the statistical power of the analysis but was the most biologically realistic test. Finally, we also excluded one locus for each species (Bbufμ47 and Rtempμ3) because they yielded Fst estimates discordant from the other seven and thus might have been affected by selection.
Matrices of mean Fst, Rst and geographic distances between the remaining four Sussex populations for each species are shown in Table 1. Inclusion of the eighth locus made negligible differences (average <2.5%) to R. temporaria pairwise Fst estimates, and reduced those of B. bufo uniformly by around 10%. Both sets of Fst estimates yielded similar results in subsequent analyses, and results are presented from those based on the seven concordant loci. Pairwise Fst estimates were on average significantly higher for B. bufo than for R. temporaria (randomisation test, 5000 permutations, P=0.003), as were pairwise mean Rst estimates (P=0.003). For the three populations common to both species (Crematorium, Pells and St Anne's), pairwise Fst estimates averaged >5-fold higher for B. bufo (mean 0.265) than for R. temporaria (mean 0.051), and pairwise Rst values were almost 60-fold higher for B. bufo (mean 0.221) than for R. temporaria (mean 0.004). Bufo populations were therefore much more strongly differentiated than those of Rana across identical geographical distances.
Mantel tests (1000 iterations) indicated no significant correlations between Fst and Rst for either species (B. bufo rs=−0.1429; R. temporaria rs=0.3143). Fst estimates were generally higher than Rst estimates throughout. Differences between Fst and Rst were significant for R. temporaria in Manley randomisation tests (5000 permutations, P=0.002) with mean pairwise Fst=0.050 and mean pairwise Rst=0.015. However, there were no significant differences between the two estimators among B. bufo populations (P=0.107). Permutation tests (200 cycles) showed that all pairwise Fst and Rst estimates for B. bufo (with means of 0.222 and 0.168, respectively) were significantly different from zero (5% level) after adjustment for multiple comparisons. By contrast, only one Fst and two Rst pairwise comparisons were significantly different from zero in R. temporaria.
Isolation by distance using Fst statistics with Mantel tests (Figure 3, 1000 permutations each) demonstrated no significant distance effects for B. bufo populations (Spearman Rank correlation, rs=−0.0309, P=0.537), but a significant correlation in R. temporaria despite the small number (n=4) of populations available for this test (Spearman Rank correlation, rs=0.8117, P=0.045).
Bottleneck tests
One possible consequence of the high level of genetic differentiation evident in B. bufo is a greater risk of bottlenecking, due to greater population isolation than that experienced by R. temporaria. Heterozygosity excess, a signal of recent population bottlenecks (Cornuet and Luikart, 1996), was investigated assuming that a 70–90% stepwise mutation model (Schlotterer, 2000) covers the most probable range for amphibian microsatellites (Table 2). Six out of seven B. bufo populations exhibited heterozygote excess at >50% of loci using a 70% SMM, and five out of seven showed heterozygote excess using a 90% SMM. By contrast, only a maximum of three out of seven R. temporaria populations exhibited heterozygote excess at >50% of loci under the same models. Including all polymorphic loci or excluding loci deviating from Hardy–Weinberg equilibrium gave identical results. However, no R. temporaria population and just one B. bufo population showed a significant bottleneck effect (and then only with a 70% stepwise mutation model) after Bonferroni correction of probability levels for each species (P=0.007). No population of either species showed evidence of heterozygote deficiency.
Multiple regression analysis
We investigated linear regression models to explain the levels of genetic diversity in frogs and toads. For these analyses, we included the five Sussex populations of each species, with data normalised as described in Materials and methods. For each population, its average pairwise Nm (derived using the private alleles method) with the other four populations was calculated and used as a measure of population structure. This measure of differentiation, unlike Fst, is statistically independent of He (Hedrick, 1999). Mean Nm between population pairs for R. temporaria was 1.55, some three-fold higher than the average of 0.54 for B. bufo. Interpopulation differences in Nm were highly significant between the two species (matched pair t=6.84, df=4, P=0.002). If gene flow is important in maintaining high levels of genetic diversity in these populations, heterozygosity should correlate with mean Nm.
For B. bufo:

This regression accounted for 69.1% of the variance in arcsin He and was just statistically significant (F=9.94, df=1, P=0.05). Including population size as a second independent variable failed to improve on this relationship.
For R. temporaria, there was no significant relationship between arcsin He and Nm. However,

This model explained 95.2% of the variance in arcsin He and was significant (F=40.62, df=2, P=0.024). In this case, a simpler model with population size alone was individually significant and accounted for 93.5% of variance in arcsin He.
Discussion
Despite substantially greater (5–10-fold) census population sizes at breeding sites, B. bufo exhibited significantly lower levels of genetic diversity than R. temporaria at multiple localities widely distributed across Britain. There are several possible explanations for this difference between the species. Systematic differences between the suites of microsatellite loci can probably be excluded as a cause (see Materials and methods). Indeed, more of the frog than the toad loci had small interruptions in the repeats that tend to reduce mutation rates (Richards and Sutherland, 1994). There is no reason to believe, therefore, that the frog microsatellites were inherently more mutable than the toad ones. However, there could be consistent differences in Ne:Nc ratios between frog and toad populations. B. bufo has a very low Ne:Nc ratio (⩽0.01) based on genetic estimates that took account of variances in sex ratio and reproductive success, but not long-term fluctuations in population size over multiple generations (Scribner et al, 1997). Although no comparable studies have been reported for frogs, R. temporaria has a similar reproductive biology and seems unlikely to differ greatly from B. bufo in this respect. Long-term variations in population size are the most important determinants of Ne, but again there is no reason to suppose that these differ systematically between the two species. Both R. temporaria and B. bufo can experience substantial fluctuations in Nc over long time periods (Halley et al, 1996; Meyer et al, 1998).
A more likely explanation of our results is that the definitions of populations as breeding assemblages, and thus the Nc estimates, were inappropriate surrogates of Ne in one or both species. This situation could arise if, for example, animals breeding at several ponds were really part of a single weakly differentiated population. B. bufo populations were highly differentiated with little gene flow over relatively small geographical distances. By contrast, there must be substantial gene flow among the smaller R. temporaria populations over identical distances. Breeding pond density in Sussex is some 4–5-fold higher for R. temporaria than for B. bufo (Beebee, 1981), no doubt favouring intersite migration by Rana. The situation for Rana differs from metapopulation models in which local extinctions and high population turnover are expected to result in reduced levels of genetic diversity (Harrison and Hastings, 1996). R. temporaria exists as a complex of interconnected, stable subpopulations in which extinctions seem very rare and definitions of ‘population’ boundaries are scarcely possible. Indeed, when all five Sussex ‘populations’ were analysed as a single entity, all eight loci were in Hardy–Weinberg equilibrium. B. bufo was different, with populations that behaved more like discrete units. These were larger in average size than individual R. temporaria ‘populations’, but their isolation may result in a lower overall effective population size at large geographical scales. Analysis of the five Sussex B. bufo ‘populations’ as a single entity yielded three of the eight loci significantly out of Hardy–Weinberg equilibrium.
In regression models, population structure explained a substantial amount of the variance in He in B. bufo. This was not true of R. temporaria. The negative correlation between Nc and mean heterozygosity in R. temporaria populations was both striking and unexpected. We do not know the cause of this correlation, but high levels of intersite migration in R. temporaria might increase local genetic diversity in inverse relation to local population size. A single immigrant will presumably make a greater difference to a small population than to a large one. Allozyme analyses of 12 R. temporaria and 12 B. bufo populations (Hitchings and Beebee, 1997, 1998), including some of those used in the present study, indicated a positive correlation between population size and the mean heterozygosity in B. bufo (rs=0.644, P=0.026), but no significant relationship in R. temporaria (rs=0.361, P=0.264). However, variation among allozyme loci was much lower than that detected using microsatellites in the current study. The mean allozyme heterozygosities in B. bufo (27 loci, mean population size=1300) and R. temporaria (19 loci, mean population size 385) were 0.023 and 0.065, respectively. Both types of loci therefore indicated higher levels of genetic variation in R. temporaria than in B. bufo.
The different population structures of R. temporaria and B. bufo probably arise from relatively minor autecological features. Both species are generally philopatric with respect to breeding sites (eg Elmberg, 1990; Reading et al, 1991), but nevertheless colonise new ponds effectively when these are created near existing breeding sites (Baker and Halliday, 1999). However, because toads are more selective than frogs with respect to breeding site choice, the distances between suitable ponds are usually greater for the former species. It is probably this site selectivity that reduces gene flow for Bufo relative to Rana in most landscapes. This difference is, however, likely to be habitat context dependent. Among populations of the same species on Baltic islands, genetic differentiation at allozyme loci was stronger for R. temporaria than for B. bufo, although relatively weak (Fst<0.07) in both cases (Seppa and Laurila, 1999).
These observations highlight a need to assess population structure and its environmental context when interpreting differences in genetic diversity between species. They are also relevant to conservation biology and the genetic management of animal and plant populations. B. bufo has experienced substantial declines in large areas of England in recent years, at a time when R. temporaria has apparently been stable (Carrier and Beebee, 2003). It may be that differences in population structure in Britain have made B. bufo more vulnerable than R. temporaria to environmental change. It will be increasingly important to maintain gene flow, by the use of habitat corridors or more generally by less intensive landscape use, if species such as B. bufo are to persist as viable metapopulations. We cannot conclude from a comparative study of just two species as to how widespread differences of this kind are likely to be, but B. bufo and R. temporaria serve as a useful example of how two animals with broadly similar biology can show very different genetic structuring in a common landscape.
References
Amos W, Balmford A (2001). When does conservation genetics matter? Heredity 87: 257–265.
Baker JMR, Halliday TR (1999). Amphibian colonization of new ponds in an agricultural landscape. Herpetol J 9: 55–63.
Beebee TJC (1981). Habitats of the British amphibians (4): agricultural lowlands and a general discussion of requirements. Biol Conserv 21: 127–139.
Beebee TJC (1996). Ecology and Conservation of Amphibians, Chapman & Hall: London.
Beebee TJC, Griffiths RA (2000). Amphibians and Reptiles, Harpercollins: London.
Brede EG, Rowe G, Trojanowski J, Beebee TJC (2001). Polymerase chain reaction primers for microsatellite loci in the common toad Bufo bufo. Mol Ecol Notes 1: 308–310.
Carrier J-A, Beebee TJC (2003). Recent, substantial and unexplained declines of the common toad Bufo bufo in lowland England. Biol Conserv 111: 395–399.
Cooke AS (1975). Spawn site selection and colony size of the frog (Rana temporaria) and the toad (Bufo bufo). J Zool London 175: 29–38.
Cornuet JM, Luikart G (1996). Description and evaluation of two tests for detecting recent Bottlenecks. Genetics 88: 267–272.
Elmberg J (1990). Long-term survival, length of the breeding season and operational sex ratio in a boreal population of the common frog Rana temporaria L. Canad J Zool 68: 121–127.
Frankham R (1995). Effective population size/adult population size ratios in wildlife: a review. Genet Res 66: 95–107.
Frankham R (1996). Relationship of genetic variation to population size in wildlife. Conserv Biol 10: 1500–1508.
Gasc J-P (1997). Atlas of Amphibians and Reptiles in Europe, Museum National d'Histoire: Paris.
Goodman SJ (1997). RstCALC: a collection of computer programs for calculating estimates of genetic differentiation from microsatellite data and determining their significance. Molecular Ecology 6: 881–885.
Gosner K (1960). A simplified table for staging anuran embryos and larvae with notes on identification. Herpetologica 16: 183–190.
Goudet J (1995). FSTAT (V.1.2): a computer program to estimate F-statistics. J Hered 86: 485–486.
Griffiths RA, Raper SJ, Brady LD (1996). Evaluation of a standard method for surveying common frogs (Rana temporaria) and newts (Triturus cristatus, T. helveticus and T. vulgaris). Joint Nature Conservancy Council Report 21, Joint Nature Conservancy Council Report 21. Peterborough, UK.
Halley JM, Oldham RS, Arntzen JW (1996). Predicting the persistence of amphibian populations with the help of a spatial model. J Applied Ecol 33: 455–470.
Hamrick JL, Godt MJW (1989). Allozyme diversity in plant species. In: Brown AH, Clegg DMT, Kahler AL, Weir BS (eds) Plant Population Genetics, Breeding, and Genetic Resources, Sinauer: New York pp 43–63.
Harrison S, Hastings A (1996). Genetic and evolutionary consequences of metapopulation structure. Trends Ecol Evol 11: 180–183.
Hedrick PW (1999). Perspective: highly variable loci and their interpretation in evolution and conservation. Evolution 53: 313–318.
Hedrick PW (2001). Conservation genetics: where are we now? Trends Ecol Evol 16: 629–636.
Hitchings SP, Beebee TJC (1997). Genetic substructuring as a result of barriers to gene flow in urban common frog (Rana temporaria) populations: implications for biodiversity conservation. Heredity 79: 117–127.
Hitchings SP, Beebee TJC (1998). Loss of genetic diversity and fitness in common toad (Bufo bufo) populations isolated by inimical habitat. J Evol Biol 11: 269–283.
Houlahan JE, Findlay CS, Schmidt BR, Meyer AH, Kuzmin SL (2000). Quantitative evidence for global amphibian population declines. Nature 404: 752–755.
Kimura M (1983). The Neutral Theory of Molecular Evolution, Cambridge University Press: Cambridge.
Manly BFJ (1997). RT, A program for Randomization Testing V2.1, Western EcoSystems Technology Inc: Cheyenne, USA.
Meyer AH, Schmidt BR, Grossenbacher K (1998). Analysis of three amphibian populations with quarter-century long time series. Proc R Soc London B 265: 522–528.
Raymond M, Rousset F (1995). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86: 248–249.
Reading CJ, Loman J, Madsen T (1991). Breeding pond fidelity in the common toad, Bufo bufo. J Zool London 225: 201–211.
Richards RI, Sutherland GR (1994). Simple repeat DNA is not replicated simply. Nat Genet 6: 114–116.
Rousset F (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145: 1219–1228.
Rowe G, Beebee TJC (2001). Polymerase chain reaction primers for microsatellite loci in the common frog Rana temporaria. Mol Ecol Notes 1: 6–7.
Rowe G, Beebee TJC, Burke T (1997). PCR primers for polymorphic microsatellite loci in the anuran amphibian Bufo calamita. Mol Ecol 6: 401–402.
Rowe G, Beebee TJC, Burke T (2000). A microsatellite analysis of natterjack toad (Bufo calamita) metapopulations. Oikos 88: 641–651.
Schlotterer C (2000). Evolutionary dynamics of microsatellite DNA. Chromosoma 109: 365–371.
Schneider S, Kueffer J-M, Roessli D, Excoffier L (1997). ARLEQUIN ver 11: A software for population genetic analysis, Genetics and Biometry Laboratory, University of Geneva.
Scribner KT, Arntzen JW, Burke T (1997). Effective numbers of breeding adults in Bufo bufo estimated from age-specific variation at minisatellite loci. Mol Ecol 6: 701–712.
Scribner KT, Arntzen JW, Cruddace N, Oldham RS, Burke T (2001). Environmental correlates of toad abundance and population genetic diversity. Biol Conserv 98: 201–210.
Seppa P, Laurila A (1999). Genetic structure of island populations of the anurans Rana temporaria and Bufo bufo. Heredity 82: 309–317.
Slatkin M (1993). Isolation by distance in equilibrium and non-equilibrium populations. Evolution 47: 264–279.
Slatkin M (1995). Rare alleles as indicators of gene flow. Evolution 49: 53–65.
Swofford DL, Selander RB (1981). BIOSYS-1: a Fortran program for the comprehensive analysis of electrophoretic data in population genetics and systematics. J Hered 82: 309–317.
Walsh PS, Metzger DA, Higuchi R (1991). Chelex R100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10: 506–513.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Acknowledgements
E Brede was supported by a University of Sussex research studentship. We thank the various landowners for permissions to visit the study sites, the wardens of Ainsdale nature reserve for their help in collecting samples, and two anonymous referees for helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brede, E., Beebee, T. Contrasting population structures in two sympatric anurans: implications for species conservation. Heredity 92, 110–117 (2004). https://doi.org/10.1038/sj.hdy.6800391
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800391
Keywords
This article is cited by
-
Urban environment determines population genetics in the green toad, Bufotes viridis
European Journal of Wildlife Research (2023)
-
The common ground in landscape effects on gene flow in two newt species in an agroecosystem
Conservation Genetics (2023)
-
Differential dispersal shapes population structure and patterns of genetic differentiation in two sympatric pond breeding salamanders
Conservation Genetics (2015)
-
Genetic diversity of four populations of Qualea grandiflora Mart. in fragments of the Brazilian Cerrado
Genetica (2014)
-
Limited effects of suburbanization on the genetic structure of an abundant vernal pool-breeding amphibian
Conservation Genetics (2013)