Abstract
Phytophagous insects can use different host plant species across their geographic distribution. Within a locality, however, their feeding can be restricted to one or two plant species. If host species constitute different selective regimes to herbivorous insects, genetic differentiation and host-associated local adaptation may occur. In this study, we describe the genetic structure of the aphid Brevicoryne brassicae L. associated to Brassica campestris L. and B. oleraceae var. capitata L., two plant species that occur sympatrically in four localities in the highlands of Chiapas, Mexico. The aim was to determine if the aphid populations are genetically structured in relation to the plant host species, and if such differentiation is consistent among localities. The genetic description of populations was made using 11 enzyme loci using cellulose acetate electrophoresis. Aphid genotypes were surveyed in two host plant species-associated populations within each of four localities at seven polymorphic loci (eight subpopulations in total). The genetic structure was assessed at the level of subpopulations, among localities, between hosts, and pairwise comparisons of hosts within locality, using Wright F-statistics. Genetic distance among localities and between host-associated populations within each locality was also estimated. We found that overall genetic differentiation was high (FST=0.22), and that differentiation among localities (FST=0.13) was higher than differentiation between hosts (FST=0.03). All FST estimates were statistically significant. Pairwise comparisons of FST between hosts in each locality suggest high differentiation in two of them, and low but still significant differentiation in two other localities. Given that general environmental conditions are similar within localities, selection on each host species may produce genetic divergence within and among subpopulations of B. brassicae.
Similar content being viewed by others
Introduction
Phytophagous insects are often distributed in relation to the distribution of their hosts, and may feed upon a large number of host plant species throughout their geographic range (Futuyma, 1976). However, at the level of a local population, they may use a relatively restricted number of plant species. If the host plant species constitute different selective environments to herbivorous insects, adaptive genetic differentiation among populations could arise (eg Rausher, 1984; Futuyma and Peterson, 1985; Mopper and Strauss, 1998). Local genetic differentiation of insect populations in relation to different host plants has been suggested as being an incipient stage of sympatric speciation and an important phenomenon maintaining genetic diversity in insects (Bush, 1994; Berlocher and Feder, 2002; Drès and Mallet, 2002).
Aphids constitute excellent systems for assessing host-associated genetic structuring of insect populations. This is because many aphid species can often use more than one host plant species at the local scale, and thus experience different environment and selective regimes on different host plants (eg Via, 1991; Haack et al, 2000).
The host plants are considered to be an important factor that affects the genetic structuring of aphid populations (Steiner et al, 1985; Wöhrmann et al, 1986; Eggers-Schumacher and Sander, 1988; Tomiuk, 1990). Indeed, several studies have shown evidence that the genetic structure of aphids can be related to their host plants (eg De Barro et al, 1995a, 1995b; Sunnucks et al, 1997; Via, 1999; Haack et al, 2000; Anstead et al, 2002; Lushai et al, 2002).
This study aimed to assess the genetic structure of the aphid Brevicoryne brassicae L. (Homoptera: Aphididae), both among and within localities from Chiapas, Mexico. B. brassicae uses, as host plants, Brassica campestris L. and B. oleraceae var. capitata L., which occur sympatrically in the Highlands of Chiapas, Mexico. Previous observations indicated differentiation in morphological and life history traits of this aphid in relation to these two hosts (Ruiz-Montoya and Núñez-Farfán, unpublished data). Thus, besides the genetic differentiation among populations produced by spatial isolation, genetic differentiation between subpopulations (between hosts within localities) could be promoted by selection if the hosts impose differential selective pressure. This allows the assessment of genetic differentiation as a function of a biotic factor, host plant, independently of random subdivision by other means (eg predation or abiotic factors).
Materials and methods
Biology of B. brassicae L.
B. brassicae L. displays both sexual and asexual phases on an individual plant (with no alternation between host species) (Blackman and Eastop, 2000). In the study area, B. brassicae reproduces apparently only by parthenogenesis (L Ruiz-Montoya, personal observation). B. brassicae has been collected carefully by both yellow traps and on the host plants during the last 4 years in the Highlands of Chiapas. So far, there is no evidence of sexual reproduction in this aphid's populations (eg oviparous females, eggs or males; L Ruiz-Montoya, personal observation).
In the study area, B. brassicae feeds mainly upon B. campestris L. and B. oleraceae var. capitata L. B. oleracea is cultivated from the end of autumn (November), to the end of winter (February). B. campestris is an annual weed that grows near cultivated fields of B. oleraceae. Both hosts are available to aphids mainly in the winter. Small populations of B. brassicae can be found during summer on harvested plants of B. oleraceae left in the field, or on late-emerging plants of B. campestris.
Sampled localities
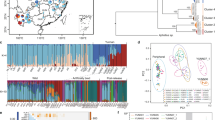
Four localities in the Highlands of Chiapas (Mexico), where the two host species occur sympatrically, were chosen for genetic analysis. Localities differ slightly in general environmental conditions (Figure 1), and are, on average, 12 km apart from one another. The altitude within the region varies from 1500 to 2600 m a.s.l. The natural vegetation is a Pinus–Quercus forest in different succesional stages (González-Espinosa et al, 1997). The mean annual temperature and rainfall are 16°C and 1500 mm, respectively. Rains occur from June to the end of September.
Sampling
Aphids were collected from plants of each host plant species selected at random in the localities of Balún, Chamula, Mitzitón and Teopisca (Figure 1). Within each locality, individuals of the two host plant species were separated from a few meters up to 200 m. In each locality, the nearest population of B. campestris to the cultivated field of B. oleracea was selected for aphid sampling. B. campestris grows in abandoned fields or in natural plant communities. Local farmers eliminate regularly the weeds from their cultivated fields, including B. campestris. Aphids were placed in vials, labelled and maintained in liquid nitrogen. In the laboratory, aphids were stored in a deep-freeze (−70°C) until electrophoretic analysis. At least 30 apterous adult aphids from different individual plants of each host species, in each locality, were screened to detect variation at the enzyme loci chosen. The total sample was: four localities (populations) and two subpopulations per locality (one for each host plant).
Electrophoresis
This was performed using horizontal cellulose acetate electrophoresis (Hebert and Beaton, 1993). Aphids were macerated in 15 μl of grinding bufffer (10 mg NADP and 100 μl β-mercaptoethanol in 100 ml water) (see Fisk et al, 1992). Five buffer systems were tested: CAAPM and TGE (Hebert and Beaton, 1993), BORATE (Richardson et al, 1986), and PHOSPHATE-1 and PHOSPHATE-2 (Fisk et al, 1992). In total, 18 enzymes/isozymes were assayed following the staining protocols of Hebert and Beaton (1993). Only the CAAPM and PHOSPHATE-1 buffer systems gave good resolution for 10 enzymes (Table 1). The electrophoresis was carried out using cellulose acetate plates of 76 mm × 76 mm (Helena Laboratories, Beaumont, TX, USA), run at ambient temperature, 95 V and 10 mA for 80 min.
Analysis
For each locality and host-associated population of B. brassicae, genotypic frequencies were scored and used to calculate the observed mean heterozygosity (Ho), and allelic frequencies. Allelic frequencies were used to estimate the mean number of alleles per locus (A), the average proportion of polymorphic loci using the 99% criterion (P), and expected mean heterozygosity (He) (Hartl and Clark, 1997). Heterogeneity of allelic frequencies among populations was evaluated by testing χ2 (Workman and Niswander, 1970). Unbiased estimates of Nei's (1978) genetic distance (D) were obtained using the program TFPGA (Miller, 1997). Average genetic distances were estimated for all populations and host-associated populations.
In order to assess population genetic structure, estimates of f, F, and θ of Weir and Cockerham (1984), the equivalent of Wright's (1965) F-statistics, FIS, FIT, and FST, respectively, were calculated. F-statistics were estimated in four ways: (A) among all (eight) subpopulations of B. brassicae; (B) among the four localities (each including the two host-associated subpopulations); (C) between hosts (pooling localities for each host); (D) pairwise comparisons of FST between host species in each locality were obtained. The estimation of genetic differentiation was performed following the procedure of Weir and Cockerham (1984), using the FSTAT program (version 2.9.3; Goudet, 2001). Jack-knife and Bootstrap methods were used to obtain standard errors and confidence intervals of F-statistics. For each locus, FST departure from zero was tested by χ2=2NFST (k−1), where k is the number of alleles, N the total number of analysed organisms, and (k−1)(s−1) degrees freedom, where s is the number of subpopulations (Workman and Niswander, 1970). The statistical test for FIS and FIT was obtained by χ2=(F)2N(k−1), with k(k−1) degrees of freedom (Nei, 1977).
Indirect estimates of gene flow Nm, the mean number of individuals or migrants exchanged among groups per generation, was calculated as Nm≈1/4(1/FST−1) since FST≈(1/(1+4Nm)) (Wright, 1951).
Results
Seven out of 11 isozyme loci tested were polymorphic (Table 1), with an average of two alleles per locus (Table 1). High levels of polymorphism (45.5–63.6%) were found in all populations (Table 2). Heterogeneity in allele frequency between hosts within locality was found for five isozymes in Mitzitón, three in Balún, four in Chamula, and two in Teopisca (Figure 2). Likewise, differences in allele frequency in the same host were detected among localities (Figure 2).
High genetic diversity, expressed as expected heterozygosity (He), was found in all populations (range 0.262–0.42). At the locality level, the difference between the observed and expected heterozygosity (fixation index) for all loci indicates a deficiency of heterozygous individuals in three localities (ie inbreeding). However, at the level of host-associated populations within a locality, the fixation index indicated inbreeding in Mitzitón (in B. campestris) and Teopisca (in B. oleraceae). Contrary, there was an excess of heterozygous individuals in Mitzitón (B. oleraceae), Chamula (B. campestris), and Balún (B. campestris) (Table 2). These deviations from random mating expectations affect the magnitude of genetic differentiation (F-statistics) between hosts and among localities (see below).
Genotypic frequencies at G6PDH, EST-2, and 6PGDH loci deviated significantly from expected frequencies in all populations associated with B. oleraceae. In B. campestris, frequencies at the MDH, EST-2 and 6PGDH loci deviated from random mating expectations in three localities (Figure 3). A similar pattern was observed among populations.
All values of FIT and FIS obtained for each locus and average were significantly different from zero among subpopulations, localities, and between hosts. As expected for clonal populations, there is a deficiency of heterozygotes at the local (FIS) and whole-population (FIT) levels (Table 3 a). Similarly, there is significant inbreeding among localities and between hosts (Table 3b, c).
The FST value estimated for the whole population was high and larger than the FST estimated among localities (Table 3a, b). Thus, the FST for the whole population includes a fraction explained by differentiation between hosts within locality. The value of FST between hosts (localities pooled within each host) indicates small but significant differentiation (Table 3c). Pairwise comparisons of FST within each locality detected high differentiation between hosts in two localities (Mitzitón and Balún, Table 4). The larger differentiation was observed in the locality of Balún (Table 4).
Genetic distance between the localities of Teopisca and Chamula was small (Figure 4). Furthermore, in each of these localities, the host-associated subpopulations showed small genetic distances between them (Figure 3). Contrary to this, Balún and Mitzitón showed high genetic distances in relation to Teopisca and Chamula. Also, subpopulations of B. brassicae in Balún and Mitzitón showed high genetic distances between hosts (Figure 3), agreeing with the pairwise estimates of FST (cf Table 4). A small amount of gene flow (≪1; indirect estimate, Nm) among all subpopulations was detected (Table 3a). Gene flow among localities and between hosts within localities resulted in higher estimates than those for all subpopulations (>1; Table 3b, c).
Discussion
A major goal of population genetics studies is to determine and quantify population substructure along with the level of gene flow among subdivided populations. In this sense, we identified genetic differentiation among all populations of B. brassicae, between localities and also, locally, we found further population subdivision associated with the host plant species used by this aphid. The host-associated genetic structure of insect populations is important in relation to host-race formation, a process that precedes speciation. The existence of host-mediated genetic differentiation is relevant for the analysis of the ecological causes that promote differentiation (Nason et al, 2002).
The genetic structure of aphid species is maintained by selection adjusting populations to the local environmental conditions (Tomiuk and Wöhrmann, 1984; Eggers-Schumacher and Sander, 1988; Tomiuk et al, 1991), by reduced gene flow (Loxdale and Brookes, 1990; Loxdale et al, 1998; Johnson et al, 2002; Massonnet et al, 2002), itself a function of species-specific flight behaviour dependent on their tendency and their ability to fly (see Loxdale et al, 1993, for a review), and by genetic drift (De Barro et al, 1995a). Perhaps, as in the case of the aphid Methopolophium dirhodum, differentiation between hosts in the localities of Mitzitón and Balún could be better explained if founder effects or genetic drift have occurred (see De Barro et al, 1995a).
Further subdivision in the genetic structure of populations may occur if the local environment is heterogeneous. The presence of different host plants in a given habitat constitutes a heterogeneous environment to insect populations. Heterogeneity in gene frequencies and high differentiation between host-associated populations of B. brassicae in, at least, two localities (Table 4) suggests that either drift or selection may contribute to further genetic structuring of B. brassicae populations in the Highlands of Chiapas. The magnitude of genetic differentiation (FST) between hosts found in this study falls within the range of estimations obtained for other host-associated populations of aphids (range from 0.012 to 0.22; Via, 1999; Delmotte et al, 2002). In addition, Cole and Lynn (1996) found differences in allele frequency of B. brassicae at short distances (<1 km) using the 6PGDH locus, and such differentiation might be related to different host plant species. The geographic distances separating pairs of host-associated populations in the present study were even shorter than 1 km, and heterogeneity in allele frequencies was common (Figure 2).
Genetic differentiation between subpopulations of B. brassicae may be associated with the different environmental conditions represented by the two host plants (Via, 1999). These differences between hosts include nutritional, physiological, or micro-environmental conditions (Loxdale et al, 1985; Steiner et al, 1985; Berlocher et al, 1993; De Barro et al, 1995a, 1995b; Eggers-Schumacher and Sander, 1988; Via, 1999). B. campestris and B. oleraceae are known to differ in nutritional quality (nitrogen content, humidity, minerals, crude protein; L Ruiz-Montoya, unpublished data), and secondary chemistry (Kjaer, 1976). Nutritional quality affects reproduction in aphids (Dixon, 1998). However, to what extent nutritional quality selects genotypes of B. brassicae that use one or the other host plant remains unknown.
Genetic differentiation and restricted gene flow between host-associated populations may promote local adaptation in phytophagous insects (Berlocher and Feder, 2002). Ample evidence of host plant differentiation has recently been found in other aphids (eg De Barro et al, 1995a, 1995b; Sunnucks et al, 1997; Via, 1999; Lushai et al, 2002), supporting the view that this phenomenon is much more widespread than previously realized. Natural selection can contribute to host-related genetic structuring in B. brassicae if intense selection occurs during establishment in different hosts. A reciprocal transference experiment between hosts indicated that B. brassicae survived better on B. campestris than in B. oleracea (Ruiz-Montoya and Núñez-Farfán, unpublished data). Thus, further analysis of local adaptation and selection will be important in understanding the patterns of morphological, life history, and genetic differentiation found in this aphids species.
Several studies have revealed greater amounts of genetic variability among aphid populations (eg RAPD-PCR, DNA-microsatellite; Vanlerbergh-Masutti and Chavigny, 1998; Wilson et al, 2002; Llewellyn et al, 2003), and hence offer the potential to further assess host-associated genetic differentiation (Lushai et al, 2002).
References
Anstead JA, Burd JD, Shufran KA (2002). Mitochondrial DNA sequence divergence among Schizaphis graminum (Hemiptera: Aphididae) clones from cultivated and non-cultivated hosts: haplotype and host associations. Bull Entomol Res 92: 17–24.
Berlocher SH, Feder JI (2002). Sympatric speciation in phytphagous insects: moving beyond controversy? Annu Rev Entomol 47: 773–815.
Berlocher SH, McPheron BA, Feder JL, Bush GL (1993). Genetic differentiation at allozyme loci in the Rhagoletis pomonella (Diptera: Tephritidae) species complex. Ann Entomol Soc Am 86: 716–727.
Blackman RL, Eastop VF (2000). Aphids on the World's Crops. An Identification and Information Guide. 2nd edn. Wiley: London.
Bush GL (1994). Sympatric speciation in animals: new wine in old bottles. Trends Ecol Evol 9: 285–288.
Cole RA, Lynn J (1996). Spatial and temporal population structure of Brevicoryne brassicae. Entomol Exp Appl 78: 121–127.
De Barro PJ, Sherratt TN, Carvalho GR, Nicol D, Iyengar A, Maclean N (1995a). Geographic and microgeographic genetic differentiation in two aphid species over southern England using the multiloucs (GATA)4 probe. Mol Ecol 4: 375–382.
De Barro PJ, Sherrat TN, David O, Maclean N (1995b). An investigation of the differential performance of clones of the aphid S. avenae on two hosts. Oecologia 104: 379–385.
Delmotte F, Leterme N, Gauthier J-P, Rispe C, Simon J-C (2002). Genetic structure of sexual and asexual populations of the aphid Rhopalosiphum padi based on allozyme and microsatellite markers. Mol Ecol 11: 711–723.
Dixon AFG (1998). Aphid Ecology. Chapman & Hall: New York.
Drès M, Mallet J (2002). Host races in plant-feeding insects and their importance in sympatric speciation. Phil Trans R Soc Lond B 357: 471–492.
Eggers-Schumacher HA, Sander E (1988). Spatial and seasonal genetic (Allozyme) variation within field populations of Phorodon humuli Schrank (Homoptera: Aphididae) on winter and summer hosts in southern Germany (FRG). Entomologist 107: 110–121.
Fisk JH, Carver M, Eastop VF (1992). An electrophoretic study of Eriosoma lanigerum (Hausmann) on apple and Eriosoma sp. from galls on Elm in Canberra, Australia (Hemiptera: Aphididae). J Aust Ent Soc 31: 231–232.
Futuyma DJ (1976). Food plant specialization and environmental predictability in lepidoptera. Am Nat 110: 285–292.
Futuyma DJ, Peterson SC (1985). Genetic variation in the use of resources by insects. Annu Rev Entomol 30: 217–238.
González-Espinosa M, Ochoa-Gaona S, Ramírez-Marcial N, Quintana-Ascencio PF (1997). Contexto vegetacional y florístico de la agricultura en Los Altos de Chiapas. In: Parra VR, Díaz-Hernández B (eds) Agricultura y crisis rural. Tomo I. Recursos Naturales, El Colegio de la Frontera Sur: Chiapas, México. pp 179–211.
Goudet J (2001). FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available from http://www.unil.ch/izea/softwares/fstat.html.
Haack L, Simon J-C, Gauthier J-P, Plantegenest M, Dedryver C-A (2000). Evidence for predominant clones in a cyclically parthenogenetic organism provided by combined demographic and genetic analyses. Mol Ecol 9: 2055–2066.
Hartl DL, Clark AG (1997). Principles of Population Genetics, 3rd edn. Sinauer Associates, Inc.: Canada.
Hebert PDN, Beaton MJ (1993). Methodologies for allozyme analysis using cellulose acetate electrophoresis. Technical Manual of Cellulose Acetate Electroporesis. Helena Laboratories, TX, USA.
Johnson PCD, Whitfield JA, Foster WA, Amos W (2002). Clonal mixing in the soldier-producing aphid Pemphigus spyrothecae (Homoptera: Aphididae). Mol Ecol 11: 1525–1531.
Kjaer A (1976) Glucosinolates in the cruciferae. In: Vaughan JG, Macleod AJ, Jones BMG (eds) The Biology and Chemistry of the Cruciferae, Academic Press: New York. pp 145–167.
Llewellyn KS, Loxdale HD, Harrington R, Brokes CP, Clark SJ, Sunnucks P (2003). Migration and genetic structure of the grain aphids (Sitobion avenae) in Britain related to climate and clonal fluctuation as revealed using microsatellites. Mol Ecol 12: 21–34.
Loxdale HD, Brookes CP (1990). Temporal genetic stability within and restricted migration (gene flow) between local populations of the blackberry-grain aphid Sitobion fragariae in south-east England. J Anim Ecol 59: 497–514.
Loxdale HD, Brookes CP, Wynne IR, Clark SJ (1998). Genetic variability within and between English populations of the damson-hop aphid, Phorodon humuli (Hemiptera: Aphididae), with special reference to esterases associated with insecticide resistance. Bull Entomol Res 88: 513–526.
Loxdale HD, Hardie J, Halber S, Fottit R, Kidd NAC, Carter CI (1993). The relative importance of short-and long-range movement of flying aphids. Biol Rev 68: 291–311.
Loxdale HD, Tarr IJ, Weber CP, Brookes CP, Digby PGN, Castañera P (1985). Electrophoretic study of enzymes from cereal aphid populations. III. Spatial and temporal genetic variation of populations of Sitiobion avenae (F.) (Hemiptera: Aphididae). Bull Ent Res 75: 121–141.
Lushai G, Markovitch O, Loxdale HD (2002). Host-based genotype variation in insects revisited. Bull Entomol Res 92: 159–164.
Massonnet B, Simon J-C, Weisser WW (2002). Metapopulation dynamics and small spatial scale genetic differentiation in the specialised herbivoe Macrosiphoniella tanacetaria (Homoptera: Aphididae). Mol Ecol 11: 2511–2521.
Miller MP (1997). Tools for Population Genetic Analyses (TFPGA). Ver. 1.3. A Window program for the analysis of allozymes and molecular population genetic data. Northern Arizona University, USA.
Mopper S, Strauss SY (1998). Genetic Structure and Local Adaptation in Natural Insect Populations. Chapman & Hall: New York, USA. p xiii.
Nason JD, Heard SB, Williams FR (2002). Host-associated genetic differentiation in the goldenrod elliptical-gall moth, Gnorimoschema gallaesolidaginis (Lepidoptera: Gelechiidae). Evolution 56: 1475–1488.
Nei M (1977). F-statistics and analysis of gene diversity in subdivided populations. Am J Hum Genet 41: 225–233.
Nei M (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89: 583–590.
Rausher MD (1984). Trade-offs in performance on different hosts: evidence from within and between-site variation in the beetle Deloyala guttata. Evolution 38: 582–595.
Richardson BJ, Baverstock PR, Adams N (1986). Allozyme Electrophoresis. A Handbook for Animal Systematics and Populations Studies. Academic Press: London.
Steiner WWM, Voegtlin DJ, Irwin ME (1985). Genetic differentiation and its bearing on migration in north American populations of the corn leaf aphid, Rhopalosiphum maidis (Fitch) (Homoptera: Aphididae). Ann Entomol Soc Am 78: 518–525.
Sunnucks P, De Barro PJ, Lushai G, Maclean N, Hales D (1997). Genetic structure of an aphid studied using microsatellites: cyclic parthenogenesis, differentiated lineages and host specialization. Mol Ecol 6: 1059–1073.
Tomiuk J (1990). Genetic stability in aphid clones and its implication for host plant interactions. In: Campbell RK, Eikenbary RD (eds) Aphid–Plant Genotype Interaction, Elsevier: Amsterdam. pp 51–66.
Tomiuk J, Hales DF, Wörhmann K, Morris D (1991). Genotypic variation and structure in Australian populations of the aphid Schoutedenia lutea. Hereditas 115: 17–23.
Tomiuk J, Wöhrmann K (1984). Genotypic variability in natural populations of Macrosiphum rosae (L.) in Europe. Biol Zbl 103: 113–122.
Vanlerbergh-Masutti F, Chavigny P (1998). Host-based genetic differentiation in the aphids Aphis gossypii Glover, evidenced from RAPD fingerprints. Mol Ecol 7: 905–914.
Via S (1991). The genetic structure of host plant adaptation in spatial patchwork: demographic variability among reciprocally transplanted aphid clones. Evolution 45: 827–852.
Via S (1999). Reproductive isolation between sympatric races of pea aphids. I. Gene flow restriction and habitat choice. Evolution 53: 1446–1457.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wilson ACC, Sunnucks P, Blackman RL, Hales DF (2002). Microsatellite variation in cyclically parthenogenetic populations of Myzus persicae in south-eastern Australia. Heredity 88: 258–266.
Wöhrmann K, Tomiuk J, Weber G (1986). The search for hidden enzymatic variation in the aphid Macrosiphum rosae (L.). Theor App Genet 73: 77–81.
Workman PL, Niswander JD (1970). Populations studies on southwerstern Indian tribes II. Local genetic differentiation in the Papago. Amer J Hum Gen 22: 24–49.
Wright S (1951). The genetical structure of populations. Ann Eugenics 15: 323–354.
Wright S (1965). The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19: 395–420.
Acknowledgements
We thank Manuel Girón and Vianey Méndez for their field assistance, and César A. Domínguez, Jorge Sarquís, and Pablo Liedo for valuable advice to the project and suggestions to the manuscript. We acknowledge a thorough revision and suggestions provided by an anonymous reviewer. Financial support was provided by the Consejo Nacional de Ciencia y Tecnología (CONACyT), México (Projects No. 31543-B and 990039-DO granted to LR-M). This is a partial fulfilment of the doctoral dissertation of LR-M.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ruiz-Montoya, L., Núñez-Farfán, J. & Vargas, J. Host-associated genetic structure of Mexican populations of the cabbage aphid Brevicoryne brassicae L. (Homoptera: Aphididae). Heredity 91, 415–421 (2003). https://doi.org/10.1038/sj.hdy.6800338
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800338
Keywords
This article is cited by
-
Phenotypic and Genetic Variations in Obligate Parthenogenetic Populations of Eriosoma lanigerum Hausmann (Hemiptera: Aphididae)
Neotropical Entomology (2015)
-
Differentiation and adaptation in Brassica nigra populations: interactions with related herbivores
Oecologia (2011)
-
Phloem Alkaloid Tolerance Allows Feeding on Resistant Lupinus angustifolius by the Aphid Myzus persicae
Journal of Chemical Ecology (2006)
-
Changes in morphological traits of the cabbage aphid (Brevicoryne brassicae) associated with the use of different host plants
Ecological Research (2005)