Abstract
Random amplified polymorphic DNA (RAPD) markers were used to estimate the population structure and phylogenetic relationships among the eight species of the genus Barbus that inhabit the Iberian Peninsula. Ten random oligodecamers were used to amplify DNA from 232 fish sampled from 15 populations. A total of 270 markers were detected that revealed low levels of genetic variability. The conclusions of cluster analysis indicate two main branches and three well-differentiated groups: north-eastern, Mediterranean and Atlantic. This clustering mainly reflects the evolutionary history of the genus, which is closely related to the paleogeography of the Iberian Peninsula. The contribution and application of these results to the conservation of the species, to their taxonomic status and to the process of colonization of the Iberian Peninsula by the genus Barbus are discussed.
Similar content being viewed by others
Introduction
The genus Barbus Cuvier and Cloquet 1816, one of the most diversified of the family Cyprinidae, is widely distributed across the Old World (Africa, Europe and Asia) where it is the dominant component among cyprinid fauna. In the Iberian Peninsula, the genus is one of the most important of all freshwater fauna, showing the highest specific diversity among all fish genera (Encina and Granado, 1988). At the present time, eight species of Barbus are found in Spain: Barbus meridionalis, B. haasi, B. bocagei, B. comizo, B. graellsii, B. guiraonis, B. sclateri and B. microcephalus. All of these, except the first, are endemic. Most show an allopatric distribution (Doadrio et al, 1991). The high number of endemic species is due to mountain (the Pyrenees) and maritime barriers (the Straits of Gibraltar). These provide the area with its singular biogeographical characteristics and favour the maintenance of local fish fauna within the Iberian Peninsula (Granado-Lorencio, 1992).
The phylogenetic relationships and taxonomic identity of Barbus have been the subject of debate for decades. Prior to 1990, their study in Europe was based on morphological data and geographical distribution (Tsigenopoulos et al, 1999). Osteological characters have also been employed in some studies to determine the relationships among the different species (Doadrio, 1990). Nevertheless, due to the considerable morphological diversity of Barbus (mainly related to growth, biogeographical and ecological factors (Encina and Granado, 1988)), the interpretation of these data have sometimes been very difficult.
Studies based on morphology, osteology, ecology and, to a lesser extent, on genetics, have all made contributions to the present understanding of the Barbus genus (Encina and Granado-Lorencio, 1988, 1997; Collares-Pereira and Madeira, 1990; Torralva et al, 1997; Aparicio and Sostoa, 1998; Chenuil et al, 1998, 2000; Tsigenopoulos et al, 1999, Callejas and Ochando, 2000; Tsigenopoulos and Berrebi, 2000). However, basic questions on its evolutionary differentiation remain unsolved (see Tsigenopoulos and Berrebi, 2000 and references therein). These issues can be extended to the Iberian Peninsula where, although Barbus is the most speciose genus of all freshwater fish, only a few genetic studies have tried to answer questions on their phylogenetic relationships and taxonomic status (Machordom et al, 1990, 1995; Callejas and Ochando, 1998, 2000; Zardoya and Doadrio, 1998, 1999; Machordom and Doadrio, 2001). Much more information is needed for understanding the processes of evolution that has occurred in the genus Barbus, as well as for the development of efficient strategies in future conservation programmes.
The development of the random amplified polymorphic DNA technique (RAPD) (Williams et al, 1990; Welsh and McClelland, 1990) has provided a useful tool for research into genetic variability. It consists of the PCR amplification of small, inverted repeats scattered throughout the genome, using a single, short primer of arbitrary sequence. Thus, the genome can be scanned more randomly than with conventional techniques. The ability to examine genomic variation without previous sequence information (Williams et al, 1990), the relatively low cost of the technique, and the requirement of only nanograms of template DNA, are all advantages of RAPD in population and other genetic studies.
There is now increasing evidence that the RAPD technique, which has been used in different fields, can detect nuclear variation in fish (Borowski et al, 1995; Naish et al, 1995; Sultmann et al, 1995; Bielawski and Pumo, 1997; Caccone et al, 1997; Callejas and Ochando, 1998; Mamuris et al, 1999; Allendorf and Seeb, 2000). These studies have shown that RAPD is an extremely sensitive method for detecting DNA variation and for establishing genetic relationships in closely related organisms. In the present work, RAPD markers were used to infer phylogenetic relationships among the Spanish species of genus Barbus.
Materials and methods
Sample collection and DNA extraction
Fifteen samples from the eight Spanish species of barbel were collected from 14 separated sites in the Iberian Peninsula. Table 1 contains the collection sites, the species collected and the number of individuals sampled (n = 232). Figure 1 is a map showing the collection sites. Specimens were dissected in situ. Liver tissue was shipped to the laboratory in liquid nitrogen and maintained at −80°C until DNA extraction.
Genomic DNA was extracted from 2 g of liver according to Benito et al (1993), with minor modifications. Samples were treated with SDS and genomic DNA isolated using phenol:chloroform extraction and isopropanol precipitation. The resulting pellets were washed in 70% ethanol, dried and resuspended in 10 mM Tris (pH 8.0), 1 mM EDTA.
RAPD-PCR analysis
Ten oligodecamers from Operon Technologies were used (C-02, C-04, C-05, C-06, C-07, C-08, C-09, C-11, C-15 and C-16). DNA amplifications were based on conditions reported by Williams et al (1990), with slight modifications.
RAPD-PCR was performed in 25 μl reaction volumes containing 25 ng barbel DNA, 5 pmoles of primer, 0.1 mM of each dNTP, 4 mM MgCl2, 2.5 μl Stoffel buffer 10× (Perkin-Elmer) and 1.2 units of Stoffel Fragment DNA polymerase (Perkin-Elmer). DNA was amplified by a PTC-100 MJ Research Thermocycler programmed to provide a first denaturation of 5 min at 94°C, followed by 45 cycles of 1 min at 94°C, 1 min at 36°C and 6 min at 72°C, and, finally, one cycle at 72°C for 6 min.
Amplification products were resolved by electrophoresis in 2% agarose gels with TAE buffer (40 mM Tris-Acetate, 1 mM EDTA pH 8.0) containing ethidium bromide. A 100 bp ladder marker was used as a molecular size standard.
The RAPD-PCR method has some limitations, such as sensitivity to reaction conditions, occasionally non-reproducible amplification products, and possible co-migration of amplified fragments. However, these problems can be solved by following a strict protocol with standardised conditions, repeating the amplification reactions twice in order to score clearly reproducible bands, and increasing the resolution of band separation (Hadrys et al, 1992). In the present work, amplification results were routinely repeatable.
Statistical analysis
Amplified bands were scored for their presence (1) or absence (0). Mean bandsharing similarity indices were calculated within (Si) and between populations (Sij) for all possible comparisons, according to Lynch (1990): Sij = 1 + S′ij −0.5 (Si + Sj) where Si is the average similarity of individuals within population i, and S′ij is the average similarity between random pairs of individuals across populations i and j. When S′ij equals the mean similarity in the two populations, Sij = 1 indicating that the populations are homogeneous.
Genetic differentiation among the samples was examined by clustering and multivariate analysis. Genetic distances were obtained from both bandsharing and frequency data. Nei’s (1972) genetic distance values were calculated from frequencies of markers using the NTSYS-PC programme (Rohlf, 1990). Lynch’s (1991) analogue of Nei’s genetic distance was calculated from mean similarity within (Si) and between population (Sij) samples. Both sets of distance values were used to construct dendrograms with the unweighted pair-group (UPGMA, Sneath and Sokal, 1973) and neighbour-joining (NJ, Saitou and Nei, 1987) methods of analysis, employing NTSYS software. Correlations between genetic distances produced using both methods were compared using a randomised test for matrix correspondence – the Mantel test (Mantel, 1967). The reliability of the trees was evaluated using 1000 bootstrap replicates with the RAPDDIST 1.0 programme from the RAPD-PCR software package (Black IV, 1997). Finally, a principal components analysis (PCA) (Sneath and Sokal, 1973) was performed using NTSYS software (Rohlf, 1990).
Results
In the 232 specimens analysed, a total of 270 scorable bands were observed for the eight species of Barbus, ranging in size from 2000 to 300 bp. Only 17 (6.30%) were monomorphic for all species. Differences among the eight species were detected with all 10 primers. Figure 2 shows the RAPD profiles of the different species with the primer C08. As observed in Table 2, RAPD analysis also revealed intraspecific polymorphism in barbel populations, ranging from 0% in B. meridionalis, where all specimens analysed showed identical patterns, to 29.36% in B. comizo (GUA population). Frequencies of polymorphic bands were also included for species comparisons.
RAPD profiles of Spanish barbels with the primer C08. The first lane contains a 100 bp molecular weight marker. Lanes 2 and 3: Barbus microcephalus. Lanes 4–7: B. comizo. Lanes 8–12: B. sclateri. Lanes 13–18: B. bocagei. Lanes 19 and 20: B. guiraonis. Lanes 21–25: B. graellsii. Lanes 26–28: B. haasi. Lanes 29 and 30: B. meridionalis.
The matrix of the estimated values of within and between population similarity is shown in Table 3. Intrapopulation similarity indices (diagonal, Table 3) were high, ranging from 95.36% (MAN population, B. bocagei) to 100% (SER population, B. meridionalis). Interpopulation similarity values (above diagonal) were lower, ranging from 59.30% to 97.81%. The similarities between populations of the same species were clearly greater than those between populations of different species. The lowest values of between-population similarity were obtained for all comparisons of B. haasi (BAS population) and B. meridionalis (SER population) with the rest of the Spanish populations. Both species, B. haasi and B. meridionalis, inhabit the northeast of Spain. The mean similarity interpopulation value for all Spanish populations of Barbus was 75.85% ± 8.17, increasing to 80.34 ± 3.85 when B. haasi and B. meridionalis were excluded.
Genetic distance values obtained from bandsharing indices (Lynch, 1991) and marker frequencies (Nei, 1972) were also calculated. The Mantel test showed that genetic distances between populations, calculated using both methods, correlated significantly across comparisons (r = 0.9762, t = 5.4261, the one-tailed probability that observed Z was greater than random Z in 100 permutations was 0.0100). Table 3 (below diagonal) shows the values of Nei’s genetic distances. The highest intraspecific distances were found in B. bocagei (range 0.045–0.025); the lowest in B. comizo (0.006). Between any two samples, interspecific values of distance ranged from 0.115 to 0.508.
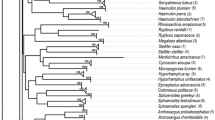
Based on Nei (1972) and Lynch (1991) genetic distances, the four dendrograms generated by the UPGMA and NJ methods showed the same topology. Therefore, Figure 3 depicts only one of them, the UPGMA dendrogram based on Nei’s genetic distances. Bootstrap values are displayed. The dendrogram exhibits two main branches: one contains the species B. haasi and B. meridionalis, both present in north-eastern Spain; the other includes the remaining Spanish species. This second branch shows that B. comizo, B. sclateri and B. bocagei, species of the Atlantic basin, cluster together, while B. graellsii and B. guiraonis, both species of the Mediterranean side of the Peninsula, group together in their own cluster. B. microcephalus (River Guadiana) is placed in the terminal node of this second main branch.
In order to visualise the gathering of populations, PCA was used to examine the variability assessed by RAPD. The first three axes covered a large portion of the total variation (89.5%), with three main groups observed (Figure 4). This PCA topology is congruent with the distribution of the groups as revealed by the dendrogram.
Plot of the first three components in a principal component analysis of RAPD data from the fifteen populations of Barbus sampled. See Table 1 for abbreviations of names of populations.
Discussion
Bands that were monomorphic in one species (and therefore its diagnostic markers) or some species, but absent in the rest, are those mainly responsible for the greater inter- rather than intraspecific polymorphism detected (Callejas and Ochando, 2001). Levels of intraspecific polymorphism ranged between 0 and 29.36% (Table 2). The higher polymorphism of some populations, such as CIJ, GUA and MAN, are due mainly to the presence of a high number of bands in low frequencies (Table 2). In general, levels of polymorphism are low compared with those seen in other fish RAPD studies (Bielawski and Pumo, 1997; Caccone et al, 1997; Gomes et al, 1998; Mamuris et al, 1999).
Table 3 shows the similarity values within- (on the diagonal) and between-populations (above the diagonal). Within-population indices were very high, ranging from 95.36% to 100%, and greater than those found among other species of fish using RAPD analysis (Bardakci and Skibinski, 1994; Foo et al, 1995; Bielawsky and Pumo, 1997). Likewise, the genetic similarity between populations of the same species was smaller than the intrapopulation similarities. This implies that individuals within each population are genetically more similar to each other than to individuals from any other population of the same species. In general, similarity values ranged from 94.06% between populations of B. comizo, to 97.81% between populations of B. sclateri, and were higher than interpopulation similarity values reported for other species of fish (Bielaswky and Pumo, 1997). Thus, although there is much genetic similarity between these populations, some geographical species differentiation appears to exist.
These parameters allow quantification of genetic diversity, and indicate levels of genetic variability in barbel populations to be rather low. Diversity was low within populations and was mainly found between populations of different species. Previous surveys in several species of European barbels, using allozymes, also showed their genetic variability to be low (Berrebi and Cattaneo-Berrebi, 1993; Machordom et al, 1995). Nowadays, although barbel populations are larger in numbers when compared with other cyprinids species, they are in decline due to environmental changes such as drought, freshwater contamination, river dredging, excessive water supply for human, agricultural or industrial purposes and the introduction of exotic species (García-Berthou and Moreno-Amich, 2000; Penczak and Kruk, 2000; Vila-Gispert et al, 2000; Doadrio, 2001). Thus, the low genetic variability of Barbus spp. is probably attributable, at least in part, to genetic drift owing to small population sizes arising as a consequence of habitat alteration. However, we cannot forget another important process that must be implicated in such low variability, ie founder effect in the evolutionary history of the genus due to its recent radiation in the Iberian Peninsula (Callejas and Ochando, 2000).
An understanding of the inter- and intraspecific distribution of genetic variation within Barbus is essential for the development of appropriate conservation strategies. At present, conservation programmes are being promoted to protect Barbus species (Economidis, 1998). The results of the present RAPD study reveal that nuclear DNA variation in Barbus Spanish species is low, and moreover this kind of study can provide important information for the management of future conservation strategies.
With respect to genetic distances (Table 3, below diagonal), intraspecific values varied from 0.0066 to 0.0405, and fall within the range reported by other authors for local populations of the same species (Ayala, 1975; Singh, 1989; Ward et al, 1992) Conversely, interspecific distance values found among the present species (except for all comparisons of B. haasi (BAS) and B. meridionalis (SER) with the remaining Spanish populations) are lower than values commonly reported among species within a genus. Rather, they are more in accordance with genetic distance data detected for subspecies (Ayala, 1975; Nei, 1976; Avise and Smith, 1977; Avise, 1994). Consequently, RAPD data can also provide insights into taxonomic status. The debate over whether these different groups represent species or subspecies is still open. According to the biological species concept (BSC), species are ‘groups of interbreeding natural populations that are reproductively isolated from other such groups’ (Mayr, 1970). Yet from the genetic point of view, most of the Iberian barbel groups might be better considered as different subspecies with recent radiation. This idea is supported by the finding of different hybrid barbels (Encina and Granado, 1988; Berrebi and Cattaneo-Berrebi, 1993; Almaça, 1996; Crespin and Berrebi, 1999; Callejas and Ochando, 2000).
The dendrogram and PCA elaborated from RAPD data, both highly congruent, are shown in Figures 3 and 4 respectively. The robustness of the tree was shown by the high bootstrap values. As a whole, cluster analysis of both bandsharing- and frequency-based distances indicates that B. haasi and B. meridionalis (both inhabitants of north-eastern Spain) are differentiated from the remaining Spanish Barbus species. At the same time, the Barbus species of the Atlantic basin and the south of the Iberian Peninsula (B. bocagei, B. comizo and B. sclateri) are clearly differentiated from Barbus species of the Mediterranean basin (B. graellsii and B. guiraonis). This pattern of differentiation is corroborated by the PCA results (Figure 4), and indicates that clustering of the populations reflects their geographical distribution. Four molecular studies have been performed on Iberian populations of Barbus in order to assess their relationships and to attempt to answer some taxonomic questions. Based on allozymes (Machordom et al, 1995), on cytochrome b sequences (Zardoya and Doadrio, 1998; Callejas and Ochando, 2000) and on ATPase 6 and 8 genes (Machordom and Doadrio, 2001), north Mediterranean species were differentiated from the remaining Iberian barbel species as were Mediterranean samples from their Atlantic counterparts.
In general, freshwater fish have limited means of dispersal, and geological events are important factors in their evolution (Borowsky et al, 1995). The present RAPD results, concordant with those mentioned above, indicate, as might be expected, that the phylogenetic relationships among Spanish barbels (Figure 3) are closely related to the formation of fluvial basins and the palaeobiogeography of the Iberian Peninsula (López-Martínez, 1989). A special case is B. microcephalus, which is placed on the terminal node of a cluster that includes Mediterranean and Atlantic barbels (the lowest value of bootstrap). This species also has an intermediate position between both groups in the PCA. Barbus microcephalus, now inhabiting the Guadiana basin, would have a common ancestor with B. graellsii and B. guiraonis as can be inferred from the number of RAPD bands they share and from its cluster position in other studies (Machordom et al, 1995; Zardoya and Doadrio, 1998; Callejas and Ochando, 2000; Machordom and Doadrio, 2001). There is faunistic evidence that a fluvial capture process occurred between the Guadiana and Jucar Rivers (Doadrio, 1984; Perdices et al, 2000) and the ancestor of B. microcephalus would have migrated from the eastern river basins to the Guadiana basin.
On the basis of ecological traits, barbels in Europe belong to two ecophenotypes: reophilic and fluvio-lacustrine barbels. Molecular studies have been performed on barbel samples using different markers (Tsigenopoulos et al, 1999; Tsigenopoulos and Berrebi, 2000) in an attempt to clarify whether there is one lineage for each different ecophenotype or whether groups of Barbus species represent clusters of morphologically convergent taxa. The results have provided contradictory conclusions in this respect. Therefore, the clear differentiation of B. meridionalis and B. haasi (north-east of Spain) from the remaining Iberian species might be explained by a longer isolation period, or as a consequence of adaptation to different environmental conditions (given that both species are mountain barbels), or by possible interactions with other European Barbus species.
At present there are some controversies surrounding barbel organisation in the northern Mediterranean area. The classical hypothesis of freshwater fish migrations (Darlington, 1948; Banarescu, 1973), established that a lineage of Barbus came from Asia to Europe and finally colonised the Iberian Peninsula and North Africa. An alternative hypothesis (Doadrio, 1990) based on osteological characters and the fossil record, proposed two different lineages. The lineage Barbus colonised the whole of Europe, including north-east Spain, from Asia, while a second lineage, Luciobarbus, migrated from Asia through the Middle East and North Africa, occupying the Iberian Peninsula from the south. Since then, the results of this group seem to support this idea (Machordom et al, 1995; Zardoya and Doadrio, 1998, 1999; Machordom and Doadrio, 2001). In contrast, other authors, based on morphological, parasitological and genetic studies, agree with the classical hypothesis of barbel dispersion (for further details, see Berrebi, 1995; Berrebi et al, 1996).
Regardless of how the directions of colonization have taken place, differences between the species of north-eastern Spain (B. haasi and B. meridionalis) and the remaining Spanish Barbus species were observed as well as a Mediterranean and Atlantic separation of the Iberian species of Barbus.
In summary, the results demonstrate that the RAPD technique is a very advantageous and useful tool for studying the genetic structure and phylogenetics of the species and population levels of Barbus. The study of the genetic structure of populations is essential in order to devise conservation programme strategies, which can efficiently maintain genetic variability. The RAPD markers revealed generally low levels of intra- and interpopulation variation, and a north-eastern, Mediterranean and Atlantic differentiation of the Iberian species. Their dispersion and divergence during a recent period, together with an adaptation to local habitats, might indicate that most of Iberian groups are still probably under speciation process from a genetic point of view.
References
Allendorf, FW, Seeb, LW (2000). Concordance of genetic divergence among sockeye salmon populations at allozyme, nuclear DNA and mitochondrial DNA markers. Evolution, 54: 640–651.
Almaça, C (1996). Peixes dos rios de Portugal. Inapa: Lisboa.
Aparicio, E, Sostoa, A (1998). Reproduction and growth of Barbus haasi in a small stream in the N.E. of the Iberian Peninsula. Arch Hydrobiol, 142: 95–110.
Avise, JC (1994). Molecular Markers, Natural History and Evolution. Chapman & Hall: New York.
Avise, JC, Smith, MH (1977). Gene frequency comparisons between sunfish (Centrarchidae) populations and various stages of evolutionary divergence. Syst Zool, 26: 319–335.
Ayala, FJ (1975). Genetic differentiation during the speciation process. Evol Biol, 8: 1–78.
Banarescu, P (1973). Origin and affinities of the freshwater fish fauna of Europe. Ichthyologia, 5: 1–8.
Bardakci, F, Skibinski, DOF (1994). Application of the RAPD technique in tilapia fish: species and subspecies identification. Heredity, 73: 117–123.
Benito, C, Figueiras, AM, Zaragoza, C, Gallego, FJ, de La Peña a, (1993). Rapid identification of Triticiae genotypes from single seeds using the PCR. Plant Mol Biol, 21: 181–183.
Berrebi, P (1995). Speciation of the genus Barbus in the North Mediterranean Basin: recent advances from biochemical genetics. Biol Conserv, 72: 237–249.
Berrebi, P, Cattaneo-Berrebi, G (1993). Natural hybridization of two species of tetraploid barbels: Barbus meridionalis and Barbus barbus (Osteichthyes, Cyprinidae) in southern France. Biol J Linn Soc, 48: 319–333.
Berrebi, P, Kottelat, M, Skelton, P, Rab, P (1996). Systematics of Barbus: State of the art and heuristic comments. Folia Zool, 45: 5–12.
Bielawski, JP, Pumo, DE (1997). Randomly amplified polymorphic DNA (RAPD) analysis of Atlantic Coast striped bass. Heredity, 78: 32–40.
Black, WC IV (1997). FORTRAN programs for the analysis of RAPD-PCR markers in populations. Colorado State University: Fort Collins, CO-80523.
Borowsky, L, McClelland, M, Cheng, R, Welsh, J (1995). Arbitrarily primed DNA fingerprinting for phylogenetic reconstruction in vertebrates: the Xiphophorus model. Mol Biol Evol, 12: 1022–1032.
Caccone, A, Allegrucci, G, Fortunato, C, Sbordoni, V (1997). Genetic differentiation within the European sea bass (D. labrax) as revealed by RAPD-PCR assays. J Hered, 88: 316–324.
Callejas, C, Ochando, MD (1998). Identification of Spanish barbel species using the RAPD technique. J Fish Biol, 53: 208–215.
Callejas, C, Ochando, MD (2000). Recent radiation of Iberian barbel fish (Teleostei, Cyprinidae) inferred from cytochrome b genes. J Hered, 91: 231–238.
Callejas, C, Ochando, MD (2001). Molecular identification (RAPD) of the eight species of the genus Barbus (Cyprinidae) in the Iberian Peninsula. J Fish Biol, 59: 1589–1599.
Chenuil, A, Cattaneo-Berrebi, G, Binda, O, Poncin, P (1998). Reproductive success of males in polyandrous crosses of Barbusmeridionalis (Cyprinidae) revealed by microsatellite genotyping. Folia Zool, 47: 53–60.
Chenuil, A, Crespin, L, Pouyaud, L, Berrebi, P (2000). Movements of adult fish in a hybrid zone revealed by microsatellite genetic analysis and recapture data. Freshwater Biol, 43: 121–131.
Collares-Pereira, MJ, Madeira, JM (1990). Cytotaxonomic studies in Iberian cyprinids. III. Karyology of Portuguese populations of Barbus Cuvier 1817, with some reconsiderations on thekaryological evolution of Cyprinidae. Caryologia, 43: 17–26.
Crespin, L, Berrebi, P (1999). Asymmetrical introgression in a freshwater fish hybrid zone as revealed by a morphological index of hybridization. Biol J Linn Soc, 67: 57–72.
Darlington, PJ (1948). The geographical distribution of cool-blooded vertebrates. Q Rev Biol, 23: 105–123.
Doadrio, I (1984). Relaciones filogenéticas y biogeográficas de los barbos (Barbus, Cyprinidae) de la Península Ibérica, y aportes corológicos y biogeográficos a su ictiofauna continental. PhD Thesis, University Complutense.
Doadrio, I (1990). Phylogenetic relationships and classification of western Palaearctic species of the genus Barbus (Osteichthyes, Cyprinidae). Aquat Living Resour, 3: 265–282.
Doadrio, I (ed) (2001). Atlas y libro rojo de los peces continentales de España. Dirección General de Conservación de la naturaleza. Museo Nacional de Ciencias Naturales: Madrid.
Doadrio, I, Elvira, B, Bernat, Y (1991). Peces continentales españoles Inventario y clasificación de zonas fluviales. ICONA-CSIC: Madrid.
Economidis, PS (1998). Evolution of barbel species: what is new in 1997? Open problems and perspectives. Folia Zool, 47: 3–6.
Encina, L, Granado, C (1988). Multivariate analysis of some morphometric characters in the genus Barbus (Pisces, Cyprinidae). Folia Zool, 37: 273–288.
Encina, L, Granado-Lorencio, C (1997). Food habits and food resource partitioning in three coexisting Barbus species. Folia Zool, 46: 325–336.
Foo, CL, Dinesh, KR, Lim, TM, Chan, WK, Eng Phan, VP (1995). Inheritance of RAPD markers in the guppy fish Poecilia reticulata. Zooll Sci, 12: 535–541.
Garcia-Berthou, E, Moreno-Amich, R (2000). Introduction of exotic fish into a Mediterranean lake over a 90-year period. Arch Hydrobiol,, 149: 271–284.
Gomes, C, Dales, RB, Oxendorf, HA (1998). The amplification of RAPD markers in stock discrimination of the four-wing flying fish, Hirundichthys affinis in the central western Atlantic. Mol Ecol, 7: 1029–1039.
Granado-Lorencio, C (1992). Fish species ecology in Spanish freshwater ecosystems. Limnetica, 8: 255–261.
Hadrys, H, Balick, M, Schierwater, B (1992). Applications of random amplified polymorphic DNA (RAPD) in molecularecology. Mol Ecol, 1: 55–63.
López-Martínez, J (1989). Paleontologia. In: Aguirre E (ed) Colección Nuevas Tendencias. CSIC: Madridpp271–296.
Lynch, M (1990). The similarity index and DNA fingerprinting. Mol Biol Evol, 7: 478–484.
Lynch, M (1991). Analysis of population genetic structure by DNA fingerprinting. In: Burke T et al (eds) DNA Fingerprinting: Approaches and Applications. Birkhäuser Verlag: Basel pp113–126.
Machordom, A, Berrebi, P, Doadrio, I (1990). Spanish barbel hybridization detected using enzymatic markers: Barbus meridionalis Risso × Barbus haasi Mertens (Osteichthyes, Cyprinidae). Aquat Living Resour, 3: 295–303.
Machordom, A, Doadrio, I (2001). Evidence of a cenozoic betic-kabilian connection based on freshwater fish phylogeography (Luciobarbus, Cyprinidae). Mol Phylogenet Evol, 18: 252–263.
Machordom, A, Doadrio, I, Berrebi, P (1995). Phylogeny and evolution of the genus Barbus in the Iberian Peninsula as revealed by allozyme electrophoresis. J Fish Biol, 47: 211–235.
Mamuris, Z, Stamatis, C, Triantaphyllidis, C (1999). Intraspecific genetic variation of striped red mullet (Mullus surmuletus L.) in the Mediterranean Sea assessed by allozyme and random amplified polymorphic DNA (RAPD) analysis. Heredity, 83: 30–38.
Mantel, NA (1967). The detection of disease clustering and a generalized regression approach. Cancer Res, 27: 209–220.
Mayr, E (1970). Populations, Species, and Evolution. Belknap Press: Harvard University, Cambridge.
Naish, KA, Warren, M, Bardacki, F, Skibinski, DOF, Carvalho, GR, Mair, GC (1995). Multilocus DNA fingerprinting and RAPD reveal similar genetic relationships between strains of Oreochromis niloticus (Pisces, Cichlidae). Mol, Ecol, 4: 271–274.
Nei, M (1972). Genetic distance between populations. Am Natural, 106: 283–292.
Nei, M (1976). In: Karlin S, Nevo E (eds) Population Genetics and Ecology. Academic: New York. p 723
Perdices, A, Doadrio, I, Côte, IM, Machordom, A, Economidis, P, Reynolds, JD (2000). Genetic divergence and origin of Mediterranean populations of the river blenny Salaria fluviatilis (Teleostei: Blenniidae). Copeia, 3: 723–731.
Penczak, T, Kruk, A (2000). Threatened obligatory riverine fishes in human-modified Polish rivers. Ecol Freshwater Fish, 9: 109–117.
Rohlf, J (1990). Numerical Taxonomy and Multivariate Analysis System NTSYS-pc. Department of Ecology and Evolution: New York.
Saitou, N, Nei, M (1987). The neighbor-joining method, a new method for reconstructing phylogenetic trees. Mol Biol Evol, 4: 406–425.
Singh, RS (1989). Genetic studies of species differences and their relevance to the problem of species formation in Drosophila. In: Loxdale HD, Hollander J (eds) Electrophoretic Studies on Agricultural Pests, Clarendon Press: Oxford pp 17–49.
Sneath, PHA, Sokal, RR (1973). Numerical Taxonomy. Freeman: San Francisco.
Sultmann, H, Mayer, WE, Figueroa, F, Tichy, H, Klein, J (1995). Phylogenetic analysis of cichlid fishes using nuclear DNA markers. Mol Biol Evol, 12: 1033–1047.
Torralva, MM, Puig, MA, Fernández-Delgado, C (1997). Effect of river regulation on the life-history patterns of Barbus sclateri in the Segura river basin (southeast Spain). J Fish Biol, 51: 300–311.
Tsigenopoulos, CS, Berrebi, P (2000). Molecular phylogeny of North Mediterranean freshwater barbs (Genus Barbus: Cyprinidae) inferred from cytochrome b sequences: biogeographic and systematic implications. Mol Phylogenet Evol, 14: 165–179.
Tsigenopoulos, CS, Karakousis, Y, Berrebi, P (1999). The North Mediterranean Barbus lineage: phylogenetic hypotheses and taxonomic implications based on allozyme data. J Fish Biol, 54: 267–286.
Vila-Gispert, A, Zamora, L, Moreno-Amich, R (2000). Use of the condition of Mediterranean barbel (Barbus meridionalis) to assess habitat quality in stream ecosystems. Arch Hydrobiol, 148: 135–145.
Ward, RD, Skibinski, DOF, Woodwark, M (1992). Protein heterozygosity, protein structure, and taxonomic differentiation. Evol Biol, 26: 73–159.
Welsh, J, McClelland, M (1990). Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res, 18: 7213–7218.
Williams, JKG, Kubelik, AR, Livak, KJ, Rafalsky, JA, Tynger, SV (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res, 18: 6531–6535.
Zardoya, R, Doadrio, I (1998). Phylogenetic relationships of Iberian cyprinids: systematic and biogeographical implications. P Roy Soc Lond, 265: 1365–1372.
Zardoya, R, Doadrio, I (1999). Molecular evidence on the evolutionary and biogeographical patterns of European cyprinids. J Mol Evol, 49: 227–237.
Acknowledgements
The authors are very grateful to A Callejas, L Campos, Dr D García de Jalón, AL López, A Mora and Dr C Pla for their kind help in providing barbel material. We would like to thank to Dr B Elvira, Dr D García de Jalón and Dr C Pla for their useful help in the morphological identification of samples. We also would like to thank to Dr JP Calvo for his valuable paleogeographical comments and assistance. During the present work, C Callejas was awarded with a Spanish doctoral grant by the Comunidad Autónoma de Madrid (CAM).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Callejas, C., Ochando, M. Phylogenetic relationships among Spanish Barbus species (Pisces, Cyprinidae) shown by RAPD markers. Heredity 89, 36–43 (2002). https://doi.org/10.1038/sj.hdy.6800091
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800091