Abstract
Background:
Aberrant expression of Brahma-related gene-1 (BRG1), a core component of the SWI/SNF chromatin-remodelling complex, has been implicated in cancer development; however, the biological significance of BRG1 in colorectal carcinoma (CRC) remains unknown.
Methods:
In CRC tissues, expression of BRG1 and Brahma (BRM) was investigated immunohistochemically. Colorectal carcinoma-derived DLD-1 cells were used for knockdown of BRG1 and PTEN with small interfering RNA (siRNA) and transduction of Akt. Complementary DNA (cDNA) microarray analysis was performed to explore the genes affected by BRG1.
Results:
Expression of BRG1, but not BRM, was frequently elevated in CRC specimens, and knockdown of BRG1 suppressed cell proliferation of DLD-1 cells. By cDNA microarray, we determined that PTEN expression was negatively regulated by BRG1 in DLD-1 cells, which subsequently influenced the cyclin D1 levels via the phosphoinositide 3-OH kinase (PI3K)–Akt signalling pathway. The interplay of BRG1 on cyclin D1 expression was confirmed by the introduction of Akt and knockdown of PTEN in the BRG1 siRNA-transduced DLD-1 cells. Interestingly, this positive correlation between BRG1 and cyclin D1 expression was also observed in CRC specimens.
Conclusion:
Brahma-related gene-1 has an important role in the process of CRC development by activating the PI3K–Akt signalling pathway and resultant upregulation of cyclin D1 levels.
Similar content being viewed by others
Main
Chromatin is actively remodelled during development, as indicated by the observations that the same genetic locus in different tissues varies in its sensitivity to DNase I and restriction enzymes (Weintraub and Groudine, 1976; McGhee et al, 1981). Chromatin remodelling of certain genes appears to precede the transcriptional activation of the genes, suggesting that chromatin remodelling may occur in anticipation of developmental transitions (Siebenlist et al, 1986). The SWI/SNF chromatin-remodelling complex is a multisubunit complex first identified in yeast and highly conserved among eukaryotes (Kingston and Narlikar, 1999; Wade and Wolffe, 1999). The mammalian SWI/SNF complex mediates ATP-dependent chromatin remodelling processes that are critical for transcriptional regulation by remodelling of nucleosomes, control of cellular processes and involvement in DNA repair, proliferation and differentiation (Roberts and Orkin, 2004; Dinant et al, 2008; Reisman et al, 2009). The SWI/SNF complex contains 9–12 different subunits that assemble into at least three separate complexes containing either single Brahma-related gene-1 (BRG1) or Brahma (BRM) as the ATPase subunit (Wang et al, 1996). The BRG1 and BRM possess highly conserved structures, with a sequence identity of 75% in humans, and their enzymatic properties are quite similar (Khavari et al, 1993; Chiba et al, 1994). Despite the fact that these subunits are interchangeable (Phelan et al, 1999), the mechanism by which the functions of BRG1 and BRM are distinguished in the SWI/SNF complex is currently poorly understood.
Brahma-related gene-1 has been reported to affect cell growth and to interact with the regulatory proteins involved in cellular proliferation in in vitro studies (Muchardt and Yaniv, 2001). Transduction of BRG1 into BRG1- and BRM-negative cells inhibited cell proliferation through altered expression of retinoblastoma (Rb) family members (Dunaief et al, 1994; Strober et al, 1996; Dahiya et al, 2000). In breast carcinoma cells, the induction of cell cycle arrest by reintroduction of BRG1 was accounted for by the downregulation of cyclin E and upregulation of cyclin-dependent kinase inhibitors p21 and p15 expression (Hendricks et al, 2004). In addition, BRG1 protein directly interacts with BRCA1 tumour suppressor and subsequently stimulates transcriptional activity of the p53 protein (Bochar et al, 2000). Thus, evidence has accumulated that supports the tumour-suppressive effects of BRG1 in human cancers. However, increased expression of BRG1 was oppositely oncogenic and indispensable for transformation of human cervical, rhabdoid and colon cancer cells: BRG1 permitted cancer cell proliferation in cooperation with the histone acetyl transferase protein, CREB-binding protein, to suppress p53 activity (Naidu et al, 2009). Thus, BRG1 may possibly be involved in biological processes that accelerate cell cycle progression and cell proliferation.
In human cancers, aberrant expressions of BRG1 and BRM have been documented in the development of tumours, including those of the stomach (Sentani et al, 2001; Yamamichi et al, 2007), lung (Reisman et al, 2003), prostate (Sun et al, 2007) and melanocytes (Lin et al, 2010); nevertheless, there is a major discrepancy in the biological significance of BRG1. The BRG1 gene deletion or mutation was found in SW13 adrenocortical carcinoma cells and PANC-1 pancreatic adenocarcinoma cells (Reisman et al, 2003). Also, ∼10% of primary lung cancers showed a concomitant loss of BRG1 and BRM expression, which was closely correlated with poor prognosis (Reisman et al, 2002). On the other hand, increased expressions of BRG1 and BRM were associated with development and progression of prostate cancer (Sun et al, 2007), cutaneous melanoma (Lin et al, 2010) and gastric carcinoma (Sentani et al, 2001). These findings indicate the possibility that the biological significance of these SWI/SNF chromatin remodelling complex molecules during the pathogenesis of human cancer differs according to cell and/or tissue types.
In this study, we investigated the pathological significance and underlying mechanisms of BRG1 and BRM in human colorectal carcinoma (CRC). We performed immunostaining of BRG1 and BRM in primary CRC specimens as well as their adjacent normal mucosa and adenoma. Knockdown of BRG1 by RNA interference was conducted for cell growth test and gene expression profiling experiment.
Materials and methods
Cell lines and tissue samples
Human CRC cell lines DLD-1, SW480, HCT116, LoVo and SW620 were obtained from the American Type Culture Collection (Manassas, VA, USA). Cells were cultured in RPMI-1640 medium containing 10% fetal bovine serum. Cells were treated with phosphoinositide 3-OH kinase (PI3K) inhibitor LY294002 (Sigma, St Louis, MO, USA) dissolved in DMSO at a final concentration of 20 μ M. A total of 31 cases of human CRCs and adenomas surgically or endoscopically removed at Kobe University Hospital (Kobe, Japan) were employed. Informed consent was obtained from all patients and the study was approved by the institutional review committee of the Kobe University. Histological examination was performed according to the Japanese Classification of Colorectal Carcinoma (Japanese Society for Cancer Colon and Rectum, 1998) along with the International Union Against Cancer classification (Sobin and Wittekind, 1997).
Immunohistochemistry and immunofluorescence
Immunohistochemistry was performed using the Labelled StreptAvidin-Biotin kit (Dako, Copenhagen, Denmark). Antibodies against BRG1 (Santa Cruz, Santa Cruz, CA, USA), BRM (Abcam, Cambridge, MA, USA), cyclin D1 (Cell Signaling, Beverly, MA, USA), PTEN (Cell Signaling) and phospho-Akt (Ser473) (p-Akt, Cell Signaling) were used. Sections were incubated with biotinylated goat anti-mouse/rabbit IgGs, and streptavidin conjugated to horseradish peroxidase (HRP) was used to immerse with 3,3-diaminobenzidine tetrahydrochloride. Immunoreactivities of BRG1, BRM, cyclin D1, PTEN and p-Akt were graded according to the staining intensity in individual cells: (−), <30% of tumour cells showed weak immunoreactivities; and (+), >30% of tumour cells showed intense immunoreactivities. For immunofluorescence, cells were grown on glass coverslips and then fixed with 1% formaldehyde. Antibodies against BRG1, BRM, E-cadherin (Santa Cruz), β-catenin (Cell Signaling), p-Akt, phospho-GSK-3β (Ser9) (p-GSK-3β, Cell Signaling) and cyclin D1 were used. Staining patterns were visualised with Cy2- or Cy3-conjugated antibody against rabbit/mouse IgGs (GE Healthcare Biosciences, Little Chalfont Buckinghamshire, UK). The nuclei were stained with 4,6-diamidino-2-phenylindole (DAPI).
RNA interference, gene transfection, cell growth test and flow cytometry
Cells were plated at a density of 1 × 105 cells and treated with BRG1 or negative control Stealth RNAi small interfering RNA (siRNA) duplex oligoribonucleotide at a final concentration of 20 nM (Invitrogen, Carlsbad, CA, USA) using Lipofectamine RNAi MAX (Invitrogen). We also used PTEN Stealth RNAi siRNA (Invitrogen). Wild-type Akt1 expression vector (p-Akt1; Upstate, Lake Placid, NY, USA) were transfected into DLD-1 cells using Lipofetamine 2000 (Invitrogen). For cell growth test, cells were plated at a density of 5.0 × 104. We counted the number of the viable cells with cell counting chamber. To analyse cellular DNA content, DLD-1 cells were collected and fixed in 70% methanol, treated with RNase A and stained with propidium iodide. The analysis was performed with a FACS Calibur cytometer (BD Biosciences, San Jose, CA, USA). Cell viability was evaluated from the population of cells in the subG1 DNA content.
Western blot
The cells were lysed in a buffer containing 50 mM Tris-HCl (pH 7.4), 125 mM NaCl, 0.1% Triton X-100 and 5 mM EDTA containing 1% protease inhibitor cocktail (Sigma). Proteins (20 μg) were separated by sodium dodecyl sulphate–polyacrylamide gel electrophoresis followed by electrotransfer onto Hybond C membrane (Millipore, Bedford, MA, USA). Primary antibodies against BRG1, BRM, PTEN, Akt (Cell Signaling), p-Akt, p-GSK-3β, GSK-3β (Cell Signaling), cyclin D1 and phospho-cyclin D1 at Thr286 (p-cyclin D1; Cell Signaling) were used. Anti-β-actin antibody (Sigma) was used for a loading control. After blotting with primary antibodies, HRP-conjugated anti-mouse/rabbit IgGs (1 : 1000 dilution; GE Healthcare Biosciences) were used as secondary antibodies. The signals were visualised with enhanced chemiluminescence.
cDNA microarray
Total RNAs were extracted from the BRG1 siRNA- and control siRNA-treated DLD-1 cells using the RNeasy kit (Qiagen, Hilden, Germany). Double-stranded cDNA was synthesised from 500 ng of total RNA Moloney murine leukaemia virus-reverse transcriptase (Agilent, Palo Alto, CA, USA) and poly dT primer incorporating the T7 promoter. Cy5-sample cRNA and Cy3-common reference cRNA were generated and hybridised to a Whole Human Genome oligo DNA microarray kit (Agilent Technologies), which was scanned using an Agilent DNA microarray scanner (Agilent), as described previously (Takeuchi et al, 2006). After data normalisation, significance analysis of microarray plot analysis was performed and significantly altered genes were identified in accordance with the manufacture's instructions (http://chem.agilent.com).
Quantitative real-time RT–PCR (qRT–PCR)
First-strand cDNA was synthesised using ReverTra Ace (Toyobo, Tokyo, Japan). In order to analyse the expression level of each mRNA, real-time quantitative PCR was performed using the ABI StepOne Realtime PCR system (Applied Biosystems, Foster City, CA, USA). Gene-specific primers were designed using the Primer Express software (Applied Biosystems). The primer sequences were as follows: PTEN: 5′-GACATTATGACACCGCCAAA-3′/5′-AAGTTCTAGCTGTGGTGGGTTATG-3′; and cyclin D1: 5′-GGGAGGGCAGTTTTCTAATGGA-3′/5′-CACCACAGTGGCCCACACT-3′. RT–PCR amplification was performed after 30 s of denaturation at 95 °C, and 40 cycles of PCR were performed at 95 °C for 5 s and 60 °C for 30 s. We confirmed that a band of single amplicon was detected in each real-time PCR reaction by the following electrophoresis. The CT values were determined by plotting the observed fluorescence against the cycle number. Each CT value was analysed using the comparative threshold cycle method and normalised to the CT values of cyclophilin A. The relative gene expression levels were estimated using the following formula: relative expression=2−(CT[target gene]−CT[cyclophilin A]).
Results
Increased levels of BRG1 expression in human CRC cases
To investigate the role of BRG1 and BRM during the pathogenesis of CRC, we first investigated the expressions of BRG1 and BRM in human CRC as well as normal colorectal mucosa and adenoma. In normal mucosa, weak immunoreactivities of BRG1 and BRM in the nuclei were detected, particularly in the cells located at the proliferative zone of crypts (Figure 1A). In CRC tissues, the BRG1 expression levels were dramatically increased, clearly indicating upregulation in comparison with those of adenoma (Figure 1A). However, no significant elevation of BRM levels was found in the same tissue samples (Figure 1A). We evaluated the average ratio of BRG1-positive cells in normal colorectal mucosa, adenoma and CRC as well as that of BRM-positive cells, and found that the average ratios of BRG1-positive cells in normal colorectal mucosa, adenoma and CRC were 31.9±1.6, 66.5±3.3, and 90.6±1.8%, respectively (P<0.05, Student's t-test, Figure 1B). However, there was no significant difference in the ratios of BRM-positive cells among normal colorectal mucosa, adenoma and CRC (Figure 1B).
The SWI/SNF chromatin-remodelling BRG1 and BRM expression in human CRC tissues. (A) Immunohistochemical results of BRG1 and BRM expression in the representative normal mucosa, adenoma and adenocarcinoma of the colorectum. (B) Average percentages of BRG1- and BRM-positive cells. *P<0.05, Student's t-test.
BRG1 knockdown reduces cell proliferation and induces morphological changes
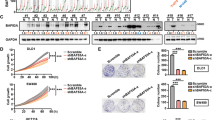
To examine the biological function of BRG1 in CRC cells, we collected five CRC cell lines and examined the protein levels of BRG1 and BRM. DLD-1, SW480 and HCT116 cells showed high levels of BRG1 and BRM expression, whereas LoVo and SW620 cells expressed neither BRG1 nor BRM (Figure 2A). Nuclear localisation of both BRG1 and BRM in DLD-1 cells are shown in Figure 2B. Then, we performed an RNA interference of BRG1 to assess the impact of silencing BRG1 on cell proliferation and morphology. The BRG1 siRNA specifically decreased BRG1 expression; however, there was no significant effect on BRM levels (Figure 2C). Transduction of BRG1 siRNA significantly reduced cell growth (P<0.05; Student’ t-test; Figure 2D) and increased the population of cells in the G1/G0 phase (Figure 2E). As transduction of BRG1 siRNA did not increase the population of cells in the subG1 DNA content, we considered that knockdown of BRG1 did not influence cell viability (Figure 2E). Also, BRG1 knockdown caused the morphologic changes: although cells treated with the negative control siRNA transfectant formed stable cell-to-cell junctions, and E-cadherin and β-catenin were linearly localised at the cell–cell borders, the distribution of these molecules was disrupted by the BRG1 siRNA transfectant (Figure 2F).
Effects of knockdown of BRG1 in human CRC cell lines. (A) Expressions of BRG1 and BRM in CRC cell lines. β-Actin was used as a loading control. (B) Nuclear localisation of BRG1 and BRM protein in DLD-1 cells. (C) Silencing of BRG1 by transduction of BRG1 siRNA into DLD-1 cells. The negative control siRNA was also transduced. (D) Results of cell growth test. *P<0.05, Student's t-test. (E) Results of cell cycle analysis. The percentages of cells in the subG1, G0/G1 and S/M/G2 DNA content are shown. (F) Morphological changes and altered expressions of E-cadherin and β-catenin in the BRG1 siRNA-transfected DLD-1 cells.
Knockdown of BRG1 upregulates the PTEN mRNA levels
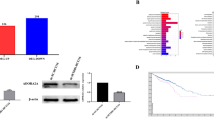
To explore the genes affected by BRG1 that are attributed to elevated cell growth, we performed a cDNA microarray gene expression profile using cDNAs from DLD-1 cells in the presence or absence of the BRG1 knockdown. We confirmed an approximately eight-fold reduction of BRG1 transcripts by transduction of BRG1 siRNA in the cDNA microarray experiment (data not shown). Then, we extracted a total of 12 cell cycle-related genes whose expressions were up- or down-regulated ⩾3-fold by transduction of BRG1 siRNA (Table 1). According to the results, we hypothesised that PTEN might have an important role as a downstream target of BRG1 in human CRC, because PTEN was a key tumour suppressor by suppressing the PI3K–Akt signalling pathway in a variety of human cancers (Li et al, 1997; Steck et al, 1997; Semba et al, 2009). Upregulation of PTEN expression was confirmed in DLD-1 (Figure 3) and SW480 cells (Supplementary Data S1). We investigated the altered expressions of the PI3K–Akt signalling-related genes. Although there was no significant difference in the PIK3CA, PIK3CB, PIK3CD and PIK3CG genes, decreased levels of gene expression of PDK1 and PI3KD1 transcripts were detected by transduction of BRG1 siRNA (Supplementary Data S2). Furthermore, no change of the Rb and p53 mRNA and protein levels were found in the BRG1 siRNA-treated DLD-1 cells (data not shown).
Knockdown of BRG1 upregulates PTEN levels in DLD-1 cells. (A) Results of real-time quantitative RT–PCR analysis. The cyclophilin mRNA levels were examined as a quality and quantity of control of mRNA. *P<0.05, Student's t-test. (B) Increased levels of PTEN protein in the BRG1 siRNA transfectant. β-Actin was used as a loading control.
Knockdown of BRG1 downregulates the cyclin D1 expression levels via inhibition of the PI3K–Akt signalling pathway
The major role of PTEN is to suppress tumourigenesis as a negative regulator of the PI3K–AKT signalling pathway (Franke et al, 1997). We examined altered expression levels of the key proteins involved in the PI3K–Akt signalling pathway when the cells were transfected with BRG1 siRNA. Silencing of BRG1 remarkably reduced not only the phosphorylated forms of Akt and GSK-3β levels, but also the total amount of cyclin D1 protein levels in DLD-1 cells, whereas transduction of BRG1 siRNA did not significantly decrease p-cyclin D1 levels (Figure 4A and B). Treatment by the LY294002 PI3K inhibitor showed the same effects on the reduced levels of p-Akt, p-GSK-3β and cyclin D1 expression (Figure 4A). The similar effect of silencing of BRG1 on the negative regulation of the PI3K–Akt signalling pathway was confirmed in SW480 cells (Supplementary Data S1). GSK-3β-induced nuclear accumulation of β-catenin is another main pathway targeting cyclin D1 by upregulation of cyclin D1 mRNA transcripts (Shtutman et al, 1999; Tetsu and McCormick, 1999). Therefore, we examined the amount of β-catenin in subcellular fractions and the cyclin D1 mRNA levels in the BRG1 siRNA- and control siRNA-transfected cells; however, we did not detect any significant changes in either experiment (Supplementary Data S3). Furthermore, co-transfection of BRG1 siRNA and p-Akt1 restored cell growth accompanied by upregulation of p-Akt, p-GSK-3β and cyclin D1 levels (Figure 4C and D), whereas transduction of PTEN siRNA decreased tumour-suppressing effect of BRG1 silencing (Figure 4E and F).
The impact of BRG1 on the cyclin D1 levels via the PI3K–Akt signalling pathway in DLD-1 cells. (A) Results of western blot analysis. Cells were also treated with PI3K inhibitor LY294002. β-Actin was used as a loading control. (B) The status of p-Akt, p-GSK-3β and cyclin D1 expressions in the BRG1 siRNA transfectant. (C) Transduction of recombinant Akt expression vector (p-Akt1) inhibited tumour-suppressive effects of BRG1 siRNA. β-Actin was used as a loading control. (D) Results of the growth test. *P<0.05, Student's t-test. (E) Co-transduction of BRG1 siRNA and PTEN siRNA. β-Actin was used as a loading control. (F) Results of the growth test. *P<0.05, Student's t-test.
High levels of BRG1 are associated with the cyclin D1 status in human CRC tissues
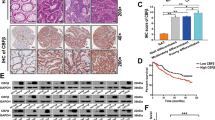
Finally, we investigated whether the expression pattern of BRG1 was consistent with that of cyclin D1 expression in human CRC tissues (Figure 5A). The results are summarised in Figure 5B. Of 31 CRCs, 23 (74%) cases showed positive immunoreactivity against BRG1. Interestingly, positive immunoreactivity against cyclin D1 was frequently detected in 21 (91%) of 23 BRG1(+) CRCs and only in 1 (13%) of 8 BRG(−) CRCs. As for the status of PTEN and p-Akt levels, there was no significant difference between BRG1(+) and BRG1(−) CRCs. Overall, 11 (35%) of BRG1(+)/PETN(−)/p-Akt(+)/cyclin D1(+) CRC cases were detected, which supported the results obtained in in vitro experiments.
The status of BRG1 expression correlates with high cyclin D1 levels in CRC cases. (A) Representative illustrations of immunoreactivities against cyclin D1, PTEN and p-Akt antibodies. (B) Summary of immunohistochemical analyses. Immunoreactivities against BRG1, cyclin D1, PTEN and p-Akt were evaluated as described in the text.
Discussion
In this study, we examined the biological significance of BRG1, the SWI/SNF chromatin-remodelling factor, during the development of CRC. Our results showed that BRG1 expression was frequently elevated in CRC tissues and that BRG1 knockdown in DLD-1 cells reduced cell proliferation by suppressing the activity of the PI3K–Akt signalling pathway by induction of PTEN expression and resultant downregulation of cyclin D1 expression. Interestingly, this correlation between BRG1 and cyclin D1 expression was also observed in human CRC tissues. To our knowledge, this is the first report indicating that aberrant BRG1 expression may promote tumour development and growth through the PI3K–Akt pathway in CRC. In these clinical specimens of human CRC, we found significantly increased expression of BRG1 but not BRM compared with the normal mucosa. In addition, the average ratio of BRG1(+) cells in adenoma and CRC was significantly higher than that in normal mucosa. This positive correlation between BRG1 expression and tissue malignancy was in agreement with the results of gastric cancer (Sentani et al, 2001), prostate cancer (Sun et al, 2007) and cutaneous melanoma (Lin et al, 2010) but not with those of lung cancer (Reisman et al, 2003).
In our BRG1 knockdown experiments, we found remarkable reduction of cell proliferation and cessation of cell cycle. As a molecular mechanism of BRG1 affecting cell proliferation, we provided evidence that BRG1 possibly suppressed PTEN expression at the mRNA and protein levels and then downregulated the PI3K–Akt signalling pathway. As reported previously, loss of PTEN (Frattini et al, 2005; Sawai et al, 2008) and phosphorylation of Akt (Itoh et al, 2002) are essential for the tumour progression of CRC cells. Although PTEN transcription is regulated by several molecules, including Egr-1 (Virolle et al, 2001), c-Jun (Hettinger et al, 2007), Id-1 (Lee et al, 2009) and TGF-β (Chow et al, 2010), little is known about the gene regulating PTEN transcription during CRC development. Our results may offer a key to understanding the regulatory basis for the PTEN mRNA in CRC. Although BRG1 contributed to tumour suppression by interaction with Rb (Dunaief et al, 1994; Strober et al, 1996; Dahiya et al, 2000) and p53 (Bochar et al, 2000), other BRG1-related mechanisms may exist for the promotion of tumour growth in CRC. Here, we have proposed the possibility that the transcription of PTEN is regulated by BRG1; further study is required to determine what mechanism BRG1 uses for transcriptional regulation of PTEN, whether by the disruption of histone-DNA contacts by ATP-dependent chromatin remodellers, or by histone tail modifications including methylation and acetylation or by direct regulation.
Cyclin D1 is an important target of the PI3K–Akt signalling pathway, and overexpression of cyclin D1 may be a significant predictor of CRC progression (Ogino et al, 2009). Nuclear β-catenin interacts with DNA-binding proteins of the TCF/LEF family and acts as a transcriptional activator of many target genes including cyclin D1 (Shtutman et al, 1999; Tetsu and McCormick, 1999). Despite previous reports showing that β-catenin expression in the nuclei can be a prognostic marker in CRC patients (Horst et al, 2008), and that BRG1 directly interacts with β-catenin to promote target gene activation (Barker et al, 2001), our results show that BRG1 expression is significantly associated with cyclin D1 expression but not nuclear β-catenin expression. This finding derives from our experiments using DLD-1 cells: BRG1 knockdown downregulated expression of cyclin D1 without subcellular change of β-catenin (Supplementary Data S3), and p-cyclin D1 expression level was relatively higher than total cyclin D1 levels (Figure 4A). Therefore, it seems reasonable that the aberrant expression of BRG1 can regulate the degradation of cyclin D1 via direct phosphorylation by GSK-3β (Diehl et al, 1998), but not via nuclear β-catenin accumulation.
The data from our cDNA microarray study using BRG1 siRNA- and negative control siRNA-treated cells suggests the further possibility that BRG1 is involved in regulating other tumour-related genes as well as the PTEN gene. In genes upregulated by BRG1 siRNA transduction, the regulator of G-protein signalling 2 (RGS2) gene is suggested to have an important role as tumour suppressor in several human cancers (Cao et al, 2006; Smalley et al, 2007). The zipper sterile-α-motif kinase (ZAK) gene might possess tumour-suppressive activity via the ERK and JNK pathways in CRC development as well as in lung cancer (Yang et al, 2010). In addition, recent studies have shown that Annexin1 (ANXA1), which was listed as a gene downregulated by BRG1 knockdown, regulates TGF-β signalling and promotes metastasis formation (de Graauw et al, 2010); loss of ANXA1 was associated with tumour progression in human breast cancer (Shen et al, 2006). In this study, we provide novel evidence that BRG1 suppresses tumour growth via PTEN transcription; however, it is possible that PTEN cooperates with the other genes mentioned above. Additional study of the genes potentially regulated by BRG1 could uncover a multifaceted role of BRG1 in CRC development.
Change history
29 March 2012
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Barker N, Hurlstone A, Musisi H, Miles A, Bienz M, Clevers H (2001) The chromatin remodelling factor Brg-1 interacts with β-catenin to promote target gene activation. EMBO J 20: 4935–4943
Bochar DA, Wang L, Beniya H, Kinev A, Xue Y, Lane WS, Wang W, Kashanchi F, Shiekhattar R (2000) BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell 102: 257–265
Cao X, Qin J, Xie Y, Khan O, Dowd F, Scofield M, Lin MF, Tu Y (2006) Regulator of G-protein signaling 2 (RGS2) inhibits androgen-independent activation of androgen receptor in prostate cancer cells. Oncogene 25: 3719–3734
Chiba H, Muramatsu M, Nomoto A, Kato H (1994) Two human homologues of Saccharomyces cerevisiae SWI2/SNF2 and Drosophila brahma are transcriptional coactivators cooperating with the estrogen receptor and the retinoic acid receptor. Nucleic Acids Res 22: 1815–1820
Chow JY, Ban M, Wu HL, Nguyen F, Huang M, Chung H, Dong H, Carethers JM (2010) TGF-β downregulates PTEN via activation of NF-κB in pancreatic cancer cells. Am J Gastrointest Liver Physiol 298: G275–G282
Dahiya A, Gavin MR, Luo RX, Dean DC (2000) Role of the LXCXE binding site in Rb function. Mol Cell Biol 20: 6799–6805
de Graauw M, van Miltenburg MH, Schmidt MK, Pont C, Lalai R, Kartopawiro J, Pardali E, Le Dévédec SE, Smit VT, van der Wal A, Van’t Veer LJ, Cleton-Jansen AM, ten Dijke P, van de Water B (2010) Annexin A1 regulates TGF-β signaling and promotes metastasis formation of basal-like breast cancer cells. Proc Natl Acad Sci USA 107: 6340–6345
Diehl JA, Cheng M, Roussel MF, Sherr CJ (1998) Glycogen synthase kinase-3β regulates cyclin D1 proteolysis and subcellular localization. Genes Dev 12: 3499–3511
Dinant C, Houtsmuller AB, Vermeulen W (2008) Chromatin structure and DNA damage repair. Epigenetics Chromatin 1: 9
Dunaief JL, Strober BE, Guha S, Khavari PA, Alin K, Luban J, Begemann M, Crabtree GR, Goff SP (1994) The retinoblastoma protein and BRG1 form a complex and cooperate to induce cell cycle arrest. Cell 79: 119–130
Franke TF, Kaplan DR, Cantley LC, Toker A (1997) Direct regulation of the Akt proto-oncogene product by phosphatidylinositol-3,4-bisphosphate. Science 275: 665–668
Frattini M, Signoroni S, Pilottie S (2005) Phosphatase protein homologue to tensin expression and phosphatidylinositol-3 phosphate kinase mutations in colorectal cancer. Cancer Res 65: 11227
Hendricks KB, Shanahan F, Lees E (2004) Role for BRG1 in cell cycle control and tumor suppression. Mol Cell Biol 24: 362–376
Hettinger K, Vikhanskaya F, Poh MK, Lee MK, deBelle I, Zhang JT, Reddy SA, Sabapathy K (2007) c-Jun promotes cellular survival by suppression of PTEN. Cell Death Differ 14: 218–229
Horst D, Reu S, Kriegl L, Engel J, Kirchner T, Jung A (2008) The intratumoral distribution of nuclear β-catenin is a prognostic marker in colon cancer. Cancer 115: 2063–2070
Itoh N, Semba S, Ito M, Takeda H, Kawata S, Yamakawa M (2002) Phosphorylation of Akt/PKB is required for suppression of cancer cell apoptosis and tumor progression in human colorectal carcinoma. Cancer 94: 3127–3134
Japanese Society for Cancer Colon and Rectum (1998) General Rules for Clinical Pathological Studies on Cancer of the Colon, Rectum and Anus, 6th edn. Kanehara: Tokyo
Khavari PA, Peterson CL, Tamkun JW, Mendel DB, Crabtree GR (1993) BRG1 contains a conserved domain of the SWI2/SNF2 family necessary for normal mitotic growth and transcription. Nature 366: 170–174
Kingston RE, Narlikar GJ (1999) ATP-dependent remodeling and acetylation as regulators of chromatin fluidity. Genes Dev 13: 2339–2352
Lee JY, Kang MB, Jang SH, Qian T, Kim HJ, Kim CH, Kim Y, Kong G (2009) Id-1 activates Akt-mediated Wnt signaling and p27kip1 phosphorylation through PTEN inhibition. Oncogene 28: 824–831
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, Puc J, Miliaresis C, Rodgers L, McCombie R, Bigner SH, Giovanella BC, Ittmann M, Tycko B, Hibshoosh H, Wigler MH, Parsons R (1997) PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275: 1943–1947
Lin H, Wong RP, Martinka M, Li G (2010) BRG1 expression is increased in human cutaneous melanoma. Br J Dermatol 163: 502–510
McGhee JD, Wood WI, Dolan M, Engel JD, Felsenfeld G (1981) A 200 base pair region at the 5′ end of the chicken adult β-globin gene is accessible to nuclease digestion. Cell 27: 45–55
Muchardt C, Yaniv M (2001) When the SWI/SNF complex remodels…the cell cycle. Oncogene 20: 3067–3075
Naidu SR, Love IM, Imbalzano AN, Grossman SR, Androphy EJ (2009) The SWI/SNF chromatin remodeling subunit BRG1 is a critical regulator of p53 necessary for proliferation of malignant cells. Oncogene 28: 2492–2501
Ogino S, Nosho K, Irahara N, Kure S, Shima K, Baba Y, Toyoda S, Chen L, Giovannucci EL, Meyerhardt JA, Fuchs CS (2009) A cohort study of cyclin D1 expression and prognosis in 602 colon cancer cases. Clin Cancer Res 15: 4431–4438
Phelan ML, Sif S, Narlikar GJ, Kingston RE (1999) Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol Cell 3: 247–253
Reisman D, Glaros S, Thompson EA (2009) The SWI/SNF complex and cancer. Oncogene 28: 1653–1668
Reisman DN, Sciarrotta J, Wang W, Funkhouser WK, Weissman BE (2003) Loss of BRG1/BRM in human lung cancer cell lines and primary lung cancers: correlation with poor prognosis. Cancer Res 63: 560–566
Reisman DN, Strobeck MW, Betz BL, Sciariotta J, Funkhouser Jr W, Murchardt C, Yaniv M, Sherman LS, Knudsen ES, Weissman BE (2002) Concomitant down-regulation of BRM and BRG1 in human tumor cell lines: differential effects on RB-mediated growth arrest vs CD44 expression. Oncogene 21: 1196–1207
Roberts CW, Orkin SH (2004) The SWI/SNF complex – chromatin and cancer. Nat Rev Cancer 4: 133–142
Sawai H, Yasuda A, Ochi N, Ma J, Matsuo Y, Wakasugi T, Takahashi H, Funahashi H, Sato M, Takeyama H (2008) Loss of PTEN expression is associated with colorectal cancer liver metastasis and poor patient survival. BMC Gastroenterol 8: 56
Semba S, Satake S, Matsushita M, Yokozaki H (2009) Phosphatase activity of nuclear PTEN is required for CDX2-mediated intestinal differentiation of gastric carcinoma. Cancer Lett 274: 143–150
Sentani K, Oue N, Kondo H, Kuraoka K, Motoshita J, Ito R, Yokozaki H, Yasui W (2001) Increased expression but not genetic alteration of BRG1, a component of the SWI/SNF complex, is associated with the advanced stage of human gastric carcinomas. Pathobiology 69: 315–320
Shen D, Nooraie F, Elshimali Y, Lonsberry V, He J, Bose S, Chia D, Seligson D, Chang HR, Goodglick L (2006) Decreased expression of annexin A1 is correlated with breast cancer development and progression as determined by a tissue microarray analysis. Hum Pathol 37: 1583–1591
Shtutman M, Zhurinsky J, Simcha I, Albanese C, D’Amico M, Pestell R, Ben-Ze’ev A (1999) The cyclin D1 gene is a target of the beta-catenin/LEF-1 pathway. Proc Natl Acad Sci USA 96: 5522–5527
Siebenlist U, Durand DB, Bressler P, Holbrook NJ, Norris CA, Kamoun M, Kant JA, Crabtree GR (1986) Promoter region of interleukin-2 gene undergoes chromatin structure changes and confers inducibility on chloramphenicol acetyltransferase gene during activation of T cells. Mol Cell Biol 6: 3042–3049
Smalley MJ, Iravani M, Leao M, Grigoriadis A, Kendrick H, Dexter T, Fenwick K, Regan JL, Britt K, McDonald S, Lord CJ, Mackay A, Ashworth A (2007) Regulator of G-protein signalling 2 mRNA is differentially expressed in mammary epithelial subpopulations and over-expressed in the majority of breast cancers. Breast Cancer Res 9: R85
Sobin LH, Wittekind CH (1997) UICC TNM Classification of Malignant Tumors, 5th edn. John Wiley & Sons, Inc.: New York
Steck PA, Pershouse MA, Jasser SA, Yung WK, Lin H, Ligon AH, Langford LA, Baumgard ML, Hattier T, Davis T, Frye C, Hu R, Swedlund B, Teng DH, Tavtigian SV (1997) Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat Genet 15: 356–362
Strober BE, Dunaief JL, Guha S, Goff SP (1996) Functional interactions between the hBRM/hBRG1 transcriptional activators and the pRB family proteins. Mol Cell Biol 16: 1576–1583
Sun A, Tawfik O, Gayed B, Thrasher JB, Hoestje S, Li C, Li B (2007) Aberrant expression of SWI/SNF catalytic subunits BRG1/BRM is associated with tumor development and increased invasiveness in prostate cancers. Prostate 67: 203–213
Takeuchi T, Tomida S, Yatabe Y, Kosaka T, Osada H, Yanagisawa K, Mitsudomi T, Takahashi T (2006) Expression profile-defined classification of lung adenocarcinoma shows close relationship with underlying major genetic changes and clinicopathologic behaviors. J Clin Oncol 24: 1679–1688
Tetsu O, McCormick F (1999) β-catenin regulates expression of cyclin D1 in colon carcinoma cells. Nature 398: 422–426
Virolle T, Adamson ED, Baron V, Birle D, Mercola D, Mustelin T, de Bella I (2001) The Egr-1 transcription factor directly activates PTEN during irradiation-induced signaling. Nature Cell Biol 3: 1124–1128
Wade PA, Wolffe AP (1999) Transcriptional regulation: SWItching circuitry. Curr Biol 9: R221–R224
Wang W, Xue Y, Zhou S, Kuo A, Cairns BR, Crabtree GR (1996) Diversity and specialization of mammalian SWI/SNF complexes. Genes Dev 10: 2117–2130
Weintraub H, Groudine M (1976) Chromosomal subunits in active genes have an altered conformation. Science 193: 4249–4253
Yamamichi N, Inada K, Ichinose M, Yamamichi-Nishina M, Mizutani T, Watanabe H, Shiogama K, Fujishiro M, Okazaki T, Yahagi N, Haraguchi T, Fujita S, Tsutsumi Y, Omata M, Iba H (2007) Frequent loss of Brm expression in gastric cancer correlates with histologic features and differentiation state. Cancer Res 67: 10727–10735
Yang JJ, Lee YJ, Hung HH, Tseng WP, Tu CC, Lee H, Wu WJ (2010) ZAK inhibits human lung cancer cell growth via ERK and JNK activation in an AP-1-dependent manner. Cancer Sci 101: 1374–1381
Acknowledgements
We thank Y Hashimoto for skilful technical assistance. This work was supported by Grant-in-Aids for Cancer Research from the Ministry of Health, Labor and Welfare of Japan (20-12) and for Scientific Research (C-19590347, C-20590341 and C-21590370) from the Japan Society for Promotion of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Watanabe, T., Semba, S. & Yokozaki, H. Regulation of PTEN expression by the SWI/SNF chromatin-remodelling protein BRG1 in human colorectal carcinoma cells. Br J Cancer 104, 146–154 (2011). https://doi.org/10.1038/sj.bjc.6606018
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.bjc.6606018
Keywords
This article is cited by
-
MiR-6511b-5p suppresses metastasis of pMMR colorectal cancer through methylation of CD44 by directly targeting BRG1
Clinical and Translational Oncology (2022)
-
PRMT1-mediated H4R3me2a recruits SMARCA4 to promote colorectal cancer progression by enhancing EGFR signaling
Genome Medicine (2021)
-
Hepatic SMARCA4 predicts HCC recurrence and promotes tumour cell proliferation by regulating SMAD6 expression
Cell Death & Disease (2018)
-
BRG1/SMARCA4 is essential for neuroblastoma cell viability through modulation of cell death and survival pathways
Oncogene (2016)
-
The chromatin remodelling component SMARCB1/INI1 influences the metastatic behavior of colorectal cancer through a gene signature mapping to chromosome 22
Journal of Translational Medicine (2013)