Abstract
ERCC1 (excision repair cross complementation group 1) is a subunit of the nucleotide excision repair complex, which can perform DNA strand incision correction of DNA damage. Association studies on the ERCC1 polymorphisms (C8092A and T19007C) in cancer had shown conflicting results. We performed a meta-analysis from all eligible case–control studies to assess the purported associations. Overall, the 19007C allele (3 853 patients and 4 349 controls) showed no significant effect on cancer risk compared to 19007T allele (P=0.39, odds ratio (OR)=0.95; 95% confidence interval (CI) 0.85–1.06, Pheterogeneity=0.001) in all subjects. Meta-analysis under other genetic contrasts did not reveal any significant association of T19007C to cancer in all subjects, Caucasians and Asians. The 19007C allele (2 279 patients and 2 808 controls) showed no significant effect on lung cancer risk compared to 19007T allele (P=0.72, OR=0.94, 95% CI 0.69–1.29, Pheterogeneity=0.0001) in all subjects. No significant effect of 8092A allele (3 865 patients and 3 750 controls) on cancer risk in all subjects (P=0.85, OR=1.01, 95% CI 0.94–1.08, Pheterogeneity=0.92) and in Caucasians and Asians compare to 8092C. No evidences of association of C8092A (501 patients and 620 controls) to squamous cell carcinoma were found. The accumulated evidence indicated ERCC1 T19007C and C8092A might not be risk factors for cancer. Significant between-study heterogeneity existed in T19007C, which arose from a study showing significant protecting effect of 19007C allele compare to 19007T allele in smokers. More studies based on larger, stratified case-control population should be required to further evaluate the role of ERCC1 C8092A and T19007C polymorphisms in different cancer, especially in smokers.
Similar content being viewed by others
Introduction
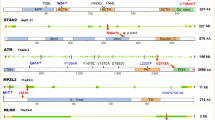
DNA in most cells is regularly damaged by chemical and physical agents in the environment such as smoking-related carcinogens and DNA lesions caused by shortwave UV component of sunlight.1 Unrepaired DNA damage may lead to the loss of genomic integrity, unregulated cell growth and ultimately cancer. Thus, genes coding for DNA repair proteins have been proposed as candidate cancer-susceptibility genes. NER (Nucleotide excision repair) is the component of the intricate network of DNA repair systems, which repairs bulk adducts lesions. ERCC1 (excision repair cross complementation group 1) is a subunit of the NER complex, which can recognize DNA damage, form heterodimers with XPF proteins, and then perform DNA strand incision.2, 3 Defect in ERCC1 had been reported to be associated with the most severe DNA repair deficiency among NER pathway genes.4
C8092A (rs3212986) and T19007C (rs11615) are two common polymorphisms of ERCC1. The C8092A polymorphism is located in the 3′-untranslated region of the gene and may affect ERCC1 messenger RNA stability.5 The synonymous T19007C polymorphism at codon 118 (Asn118Asn) may be associated with a reduction in expression of ERCC1 mRNA levels and subsequently a decreased level of ERCC1 protein.6 In recent years, many studies in cancer epidemiology focused on the associations between cancer risks and DNA-repair pathway gene SNPs, including genes in NER pathway. However, studies on the polymorphism of ERCC1 T19007C polymorphisms had shown conflicting results; studies on ERCC1 C8092A polymorphisms had shown different trends of risk in cancer, but none of the result was significant. Potential contribution of differences in patient populations (eg, age and years from onset, sex, disease severity, smoking, or diet status) might cause different results, and association can only be found in stratification analysis. Considering the possible small effect size of those two polymorphisms to cancer risk and the relatively small sample size in each study, it is important to perform a quantitative synthesis of the evidence with rigorous methods. Here, we performed a meta-analysis from all eligible case–control studies to address the association of ERCC1 C8092A and T19007C polymorphisms to cancer.
Materials and methods
Identification and eligibility of relevant studies
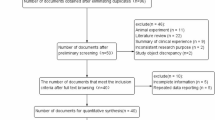
To identify all articles that examined the association of ERCC1 C8092A and T19007C polymorphisms with cancer, we conducted a literature search of PubMed database (from January 1991 to October 2006) using the following keywords and subject terms: ‘ERCC1’, ‘polymorphism’, and ‘cancer’. References of retrieved articles were screened. Abstracts, case reports, editorials, and review articles were excluded. Studies included in the current meta-analysis had to meet all the following criteria: (a) use an unrelated case–control design, (b) have available genotype frequency and (c) genotype distribution of control population must be in Hardy–Weinberg equilibrium (HWE).
Data extraction
Data were collected on the genotype of C8092A and T19007C according to different kinds of cancers. First author, year of publication, ethnicity of study population, and characteristics of cases and control were described.
Statistical analysis
Odds ratios (OR) corresponding to 95% confidence interval (CI) was applied to assess the strength of association of C8092A and T19007C with cancers as case–control studies were used, and OR was calculated according to the method of Woolf.7 A χ2-based Q statistic test was performed to assess the between – study heterogeneity.8 Heterogeneity was considered significant for P<0.10 because of the low power of the statistic. A fixed-effect model using the Mantel–Haenszel method and a random-effects model using the DerSimonian and Laird method were used to pool the results.9 The significance of the pooled OR was determined by the Z-test; a P-value of <0.05 was considered significant. For each genetic contrast, subgroup analysis according to ethnicity was only considered for Asian and Caucasian population to estimate ethnic-specific OR, subgroup analysis according to different kind of cancer was only considered for lung cancer and squamous cell carcinoma.
Publication bias was investigated by funnel plot, in which the standard error of log (OR) of each study was plotted against its OR. An asymmetric plot suggested possible publication bias. Funnel plot asymmetry was assessed by the method of Egger's linear regression test, a linear regression approach to measure funnel plot asymmetry on the natural logarithm scale of the OR.10 The significance of the intercept was determined by the t-test as suggested by Egger, and a P-value of <0.05 was considered significant.
HWE was tested by the χ2-test for goodness of fit using a web-based program (http://ihg.gsf.de/cgi-bin/hw/hwa1.pl). Analyses were performed using the software Stata version 7, ReviewManage 4.2. All P-values were two-sided.
Results
Selection of studies
The most popular SNP studied of ERCC1 were C8092A and T19007C; articles about SNPs other than these two were discarded in our selection of studies. A total of 21 articles were retrieved based on the searching criteria for cancer susceptibility related to these two SNPs, and 15 met our inclusion criteria. Out of these 15 eligible articles 10 described T19007C11, 12, 13, 14, 15, 16, 17, 18, 19, 20 and 9 of the 14 eligible articles described C8092A5, 11, 12, 17, 18, 21, 22, 23, 24 (Table 1). Among the 15 eligible articles included, 52% (11/21) stated that the age and sex status were matched between case and control population. All the articles used blood sample for genotyping.
In all the eligible articles, Wrensch et al23 provided data on two kinds of cancers: glioblastoma and nonglioblastoma; Matullo et al20 described on four different cancers: bladder, lung, leukemia, and upper aerodigestive (UADC)cancer, which include cancer from oral, pharyngeal, or laryngeal. Thus, each type of cancer in these two articles was treated as a separate study in the meta-analysis. Studies provide genotyping data of mixed population indicated as ‘mixed’ ethic (Table 1).
Summary statistics
A total of 3 853 cancer patients and 4 349 controls for T19007C, 3 865 cancer patients and 3 750 controls for C8092A were investigated. The allele frequencies were calculated for controls from the corresponding genotype distributions (Table 1). The T19007 allele had a lower representation among controls of Asian descent 24.7%, 95% CI (−24.9 to 74.3) than in controls of European descent 59.3%, 95% CI (55.8–62.8). The C8092 allele was equally represented among controls of Asian descent 74.2%, 95% CI (64.0–84.4) as in controls of European descent 75.0%, 95% CI (73.4–76.5). Overall, the prevalence of T19007 and C8092 allele was 52.2 and 74.6% in controls, respectively.
Overall effects for alleles
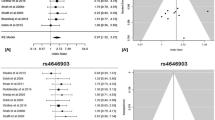
Significant heterogeneity existed in 13 studies when compared T19007C T vs C allele in different kinds of cancers (Table 2). The random effects model was used to pool the result (Figure 1a). There was no evidence that the C allele associated with the risk of cancer in all subjects. The summary OR was 0.95, 95% CI (0.85–1.06) by random effects (P=0.39) with Pheterogeneity=0.001 (Table 2). A sensitivity analysis showed that the study by Zienolddiny et al12 was the main cause of the heterogeneity. After exclusion of this study, the heterogeneity was no longer significant (Pheterogeneity=0.13), and the estimated overall effect remained insignificant (P=0.89, OR=1.00, 95% CI (0.94–1.06) by fixed effect).
Overall meta-analysis for T19007C polymorphism (C vs T allele) in cancers and lung cancer. The study is shown by a point estimate of the OR and the accompanying 95% CI under a random – effects model. (a) analyzed the comparison in cancers and (b) analyzed the comparison in lung cancer. n indicates the total number of C allele or T allele, N indicates the total number of C allele plus T allele.
Four out of 13 studies including 2 279 patients and 2 808 controls examined the association of 19007C to lung cancer risk. No evidence of association between 19007C allele and lung cancer risk (Figure 1b, P=0.72, OR=0.94, 95% CI (0.69–1.29), Pheterogeneity=0.0001, Table 2) was found. No between-study heterogeneity was existed (Pheterogeneity=0.16) after excluding the study of Zienolddiny et al,12 no association of 19007C allele to the risk of lung cancer (P=0.38, OR=1.04, 95% CI (0.95–1.15).
No association of 19007C allele to the risk of cancer in both subgroups was found, with 10 comparisons in Caucasian descent (P=0.48, OR=0.95, 95% CI (0.83–1.09) Pheterogeneity=0.0003) and two comparisons in Asian descent (P=0.99, OR=1.00, 95% CI (0.76–1.31), Pheterogeneity=0.89) (Table 2). The between-study heterogeneity remained after exclusion the study of Zienolddiny et al12 (P=0.93, OR=1.00, 95% CI (0.90–1.12), Pheterogeneity=0.05), which indicated other studies contribute to the heterogeneity in Caucasians.
No significant heterogeneity existed in 10 studies when compared C8092A A vs C allele in different kinds of cancers (Table 2). The fixed effect model was used to pool the result (Figure 2a). There was no evidence that the 8092A allele associated with the risk of cancer in all subjects, Caucasians, and Asians, with the summary OR=1.01, 95% CI (0.94–1.08) P=0.85, Pheterogeneity=0.92; OR=0.99, 95% CI (0.91–1.09) P=0.90, Pheterogeneity=0.55; OR=1.00, 95% CI (0.74–1.34) P=1.00, Pheterogeneity=0.87, respectively (Table 2).
Overall meta-analyses for C8092A polymorphism (A vs C allele) in cancers and squamous cell carcinoma. The study is shown by a point estimate of the OR and the accompanying 95% CI under a fixed – effect model in (a) when analyzed the comparison in cancers and (b) when analyzed the comparison in squamous cell carcinoma. n indicates the total number of A allele or C allele, N indicates the total number of A allele plus C allele.
Three out of 10 studies including 501 patients and 620 controls examined the association of C8092A to squamous cell carcinoma risk, but no association of 8092A to squamous cell carcinoma risk was found (Figure 2b, P=0.58, OR=0.95, 95% CI (0.78–1.15), Pheterogeneity=0.88, Table 2).
Other genetic contrasts
Meta-analysis under other genetic contrasts (dominant, recessive, and additive genetic models) suggested that the 19007C showed no association to cancer risk in all subjects, Caucasians, and Asians, as well as the association between T19007C and lung cancer in the genetic models in all subjects (Table 2). 8092A showed no association to cancer risk under different genetic models in all subjects, Caucasians and Asians. No evidence of association between 8092A and squamous cell carcinoma was discerned as we compared other genetic models (Table 2). No single study influenced the pooled ORs as reveal by sensitivity analysis (data not shown), which further confirmed that no association of the T19007C and C8092A polymorphisms to cancer risk.
Publication bias
Egger's test provided no evidence for funnel plot asymmetry in the comparison of 19007C allele vs 19007T allele in the OR analysis in overall cancer (t=−0.50, P=0.63). Similarly, no publication bias was detected for A vs C contrast of C8092A polymorphism in overall cancer (t=−0.80, P=0.45). Figure 3 showed the Begg's funnel plot of the Egger's test.
Discussion
In the analysis of ERCC1 T19007C polymorphisms to susceptibility of cancer, significant between-study heterogeneity was found in all the comparisons except in the recessive genetic model (CC vs (CT+TT) (Pheterogeneity=0. 15). Sensitivity analysis showed the study of Zienolddiny et al12 was the origin of the heterogeneity in all subjects. In the Caucasian subgroup analysis, the between-study heterogeneity remained after exclusion the study of Zienolddiny et al;12 further exclusion of either the Matullo_bladder data in Matullo et al20 or Skielbred et al15 make the between-study heterogeneity no longer exist, but the effect of exclusion Matullo_bladder data in Matullo et al (Pheterogeneity=0.14)20 is bigger than that of Skielbred et al (Pheterogeneity=0.11).15
Smoking is well known as a susceptibility factor for cancer, especially in lung cancer. Zienolddiny et al12 used matched smokers of cases and controls and reveal a significant protection effect of 19007C allele vs 19007T allele OR=0.58, 95% CI (0.45–0.75). In the study of Matullo et al,15, 20 different cancers including lung cancer were examined. The cases and controls were matched on sex, age, and smoking status, but the smoking status was only stratified for former-smokers (268 patients and 499 controls) and never-smokers (300 patients and 595 controls). The study provides overall genotyping data of cases and controls rather than stratified data for former-smokers and never-smokers. The results were different in different kinds of cancers: in lung cancer, a higher risk of C allele than T allele OR=1.34, 95%CI (1.02–1.76) was found; in bladder cancer, the C allele had a trend of protection compared to T allele OR=0.76, 95%CI (0.58–1.00). No association of C allele to the leukemia risk OR=0.98, 95% CI (0.77–1.23) and to UADC OR=1.16, 95% CI (0.84–1.60).
It is possible that the association between ERCC1 polymorphisms and cancer risk might be modified by smoking. However, no sufficient studies available to do such an analysis so far. More careful stratification analyses according to the smoking status and cancer types are needed; meta-regression analysis is then possible to reveal the risk of smoking in cancer.
Different studies showed different trend of effect of the C8092A polymorphism on the risk of different kinds of cancers, which reflected by the OR value (below 1 or above 1), but none of them had a significant effect. No between-study heterogeneity existed among different cancer types. Potential contribution of differences in patient populations (eg, age and years from onset, gender difference, and cancer types) did not affect the overall effect of C8092A polymorphism.
DNA repair is known as a ‘double-edged sword’ in cancer. Impaired DNA repair capacity may increase risk of cancer, but meanwhile induce resistance to the antitumor activity of platinum-DNA adducts.25 Platinum-based chemotherapy induced DNA damage (inter-/intrastrand cross-linking) by forming platinum-DNA adducts and causes cell death.26 Increased tolerance to DNA damage, resulting from a highly efficient DNA-repair capacity,27 is one of the mechanisms of drug resistance of platinum-based chemotherapy. Bulky DNA adducts by cisplatin are mainly repaired by the NER pathway. Enhanced NER ability for excision of bifunctional DNA adducts28, 29 may effectively reduce the anticancer effect of platinum-based chemotherapy, leading to continued cancer growth and metastasis.30 Thus, Alterations in the function of ERCC1 may affect DNA repair proficiency and influence cancer patients' response to cisplatin. ERCC1 T19007C and C8092A polymorphisms may not confer a significant effect in the susceptibility of cancer, but they may play an important role in the drug sensitivity of platinum-based chemotherapy.
In conclusion, our meta-analysis suggests that the ERCC1 T19007C and C8092A polymorphisms had no association to cancer risk. However, significant between-study heterogeneity did exist in the analysis T19007C allele, which results from a study using smokers of matched cases and controls in lung cancer.12 More studies or large case–control studies, especially studies stratified for smoking–genotyping interaction should be performed to clarify possible roles of T19007C and C8092A in different kinds of cancer.
References
de Laat WL, Jaspers NG, Hoeijmakers JH : Molecular mechanism of nucleotide excision repair. Genes Dev 1999; 13: 768–785.
Goode EL, Ulrich CM, Potter JD : Polymorphisms in DNA repair genes and associations with cancer risk. Cancer Epidemiol Biomarkers Prev 2002; 11: 1513–1530.
van Duin M, de Wit J, Odijk H et al: Molecular characterization of the human excision repair gene ERCC-1: cDNA cloning and amino acid homology with the yeast DNA repair gene RAD10. Cell 1986; 44: 913–923.
Wilson MD, Ruttan CC, Koop BF, Glickman BW : ERCC1: a comparative genomic perspective. Environ Mol Mutagen 2001; 38: 209–215.
Chen P, Wiencke J, Aldape K et al: Association of an ERCC1 polymorphism with adult-onset glioma. Cancer Epidemiol Biomarkers Prev 2000; 9: 843–847.
Yu JJ, Lee KB, Mu C et al: Comparison of two human ovarian carcinoma cell lines (A2780/CP70 and MCAS) that are equally resistant to platinum, but differ at codon 118 of the ERCC1 gene. Int J Oncol 2000; 16: 555–560.
Woolf B : On estimating the relation between blood group and disease. Ann Hum Genet 1955; 19: 251–253.
Lau J, Ioannidis JP, Schmid CH : Quantitative synthesis in systematic reviews. Ann Intern Med 1997; 127: 820–826.
Petitti D : Meta-analysis, Decision Analysis, and Cost-effectiveness Analysis. New York: Oxford University Press, 1994.
Egger M, Davey Smith G, Schneider M, Minder C : Bias in meta-analysis detected by a simple, graphical test. BMJ 1997; 315: 629–634.
Zhou W, Liu G, Park S et al: Gene-smoking interaction associations for the ERCC1 polymorphisms in the risk of lung cancer. Cancer Epidemiol Biomarkers Prev 2005; 14: 491–496.
Zienolddiny S, Campa D, Lind H et al: Polymorphisms of DNA repair genes and risk of non-small cell lung cancer. Carcinogenesis 2006; 27: 560–567.
Yin J, Vogel U, Guo L, Ma Y, Wang H : Lack of association between DNA repair gene ERCC1 polymorphism and risk of lung cancer in a Chinese population. Cancer Genet Cytogenet 2006; 164: 66–70.
Hirata H, Hinoda Y, Matsuyama H et al: Polymorphisms of DNA repair genes are associated with renal cell carcinoma. Biochem Biophys Res Commun 2006; 342: 1058–1062.
Skjelbred CF, Saebo M, Nexo BA et al: Effects of polymorphisms in ERCC1, ASE-1 and RAI on the risk of colorectal carcinomas and adenomas: a case–control study. BMC Cancer 2006; 6: 175.
Matullo G, Guarrera S, Sacerdote C et al: Polymorphisms/haplotypes in DNA repair genes and smoking: a bladder cancer case–control study. Cancer Epidemiol Biomarkers Prev 2005; 14: 2569–2578.
Moreno V, Gemignani F, Landi S et al: Polymorphisms in genes of nucleotide and base excision repair: risk and prognosis of colorectal cancer. Clin Cancer Res 2006; 12: 2101–2108.
Weiss JM, Weiss NS, Ulrich CM et al: Interindividual variation in nucleotide excision repair genes and risk of endometrial cancer. Cancer Epidemiol Biomarkers Prev 2005; 14: 2524–2530.
Vogel U, Olsen A, Wallin H et al: Effect of polymorphisms in XPD, RAI, ASE-1 and ERCC1 on the risk of basal cell carcinoma among Caucasians after age 50. Cancer Detect Prev 2005; 29: 209–214.
Matullo G, Dunning AM, Guarrera S et al: DNA repair polymorphisms and cancer risk in non-smokers in a cohort study. Carcinogenesis 2006; 27: 997–1007.
Sturgis EM, Dahlstrom KR, Spitz MR, Wei Q : DNA repair gene ERCC1 and ERCC2/XPD polymorphisms and risk of squamous cell carcinoma of the head and neck. Arch Otolaryngol Head Neck Surg 2002; 128: 1084–1088.
Sugimura T, Kumimoto H, Tohnai I et al: Gene-environment interaction involved in oral carcinogenesis: molecular epidemiological study for metabolic and DNA repair gene polymorphisms. J Oral Pathol Med 2006; 35: 11–18.
Wrensch M, Kelsey KT, Liu M et al: ERCC1 and ERCC2 polymorphisms and adult glioma. Neuro-oncol 2005; 7: 495–507.
Yang M, Kim WH, Choi Y et al: Effects of ERCC1 expression in peripheral blood on the risk of head and neck cancer. Eur J Cancer Prev 2006; 15: 269–273.
Wei Q, Frazier ML, Levin B : DNA repair: a double-edged sword. J Natl Cancer Inst 2000; 92: 440–441.
Fox M, Roberts JJ : Drug resistance and DNA repair. Cancer Metastasis Rev 1987; 6: 261–281.
Johnson SW, Stevenson JP, O'Dwyer PJ : Cisplatin and Its Analogues. Philadelphia, PA: Lippincott Williams and Wilkins, 2001.
Reed E : Platinum-DNA adduct, nucleotide excision repair and platinum based anti-cancer chemotherapy. Cancer Treat Rev 1998; 24: 331–344.
Zeng-Rong N, Paterson J, Alpert L et al: Elevated DNA repair capacity is associated with intrinsic resistance of lung cancer to chemotherapy. Cancer Res 1995; 55: 4760–4764.
Wei Q, Cheng L, Xie K et al: Direct correlation between DNA repair capacity and metastatic potential of K-1735 murine melanoma cells. J Invest Dermatol 1997; 108: 3–6.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Li, Y., Gu, S., Wu, Q. et al. No association of ERCC1 C8092A and T19007C polymorphisms to cancer risk: a meta-analysis. Eur J Hum Genet 15, 967–973 (2007). https://doi.org/10.1038/sj.ejhg.5201855
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201855
Keywords
This article is cited by
-
ERCC1 polymorphisms as prognostic markers in T4 breast cancer patients treated with platinum-based chemotherapy
Journal of Translational Medicine (2014)
-
Pharmacogenetic analysis of adjuvant FOLFOX for Korean patients with colon cancer
Cancer Chemotherapy and Pharmacology (2013)
-
A C118T polymorphism of ERCC1 and response to cisplatin chemotherapy in patients with late-stage non-small cell lung cancer
Journal of Cancer Research and Clinical Oncology (2012)