Abstract
The tumor suppressor p53 has two alternative effects, causing either cell cycle arrest or apoptosis. These different effects are supposed to be mediated by the transcriptional activation of different target genes. perp, encoding a transmembrane protein of the Pmp22 family, is a transcriptional p53 target exclusively upregulated in apoptotic cells. However, its role during normal development had remained largely unclear. Here, we report the isolation and characterization of a zebrafish perp homolog. Upon overexpression in early zebrafish embryos, perp induces apoptosis. In addition, it contributes to p53-dependent and UV-induced cell death. However, during normal zebrafish development, perp displays a p53-independent and spatially restricted expression in specific cell types and tissues. Antisense-mediated loss of Perp function leads to increased apoptosis in perp-expressing cells of the developing skin and notochord. We conclude that, in contrast to its proapoptotic function in stressed cells, Perp plays an antiapoptotic role during normal zebrafish development to regulate tissue-specific cell survival.
Similar content being viewed by others
Introduction
The transcription factor p53 plays a crucial role during tumor suppression, and loss of p53 is associated with most human cancers. Upon DNA damage or other cellular stress, p53 becomes activated and induces either cell cycle arrest to allow DNA repair, or apoptosis, to eliminate cells with elevated carcinogenic potential. The choice between G1 arrest and apoptosis is not well understood, but is proposed to be mediated by the activation of different p53 target genes. Thus, the upregulation of cyclin-dependent kinase inhibitors like p21 can drive cells into G1 arrest, while factors like Bax, Killer, or Perp are implicated in p53-induced apoptosis (reviewed in Wu et al1).
Bax and Killer activate apoptosis by two distinct pathways, both of which converge at the level of caspase activation (reviewed in Hengarter2). Killer1 is a death receptor inserted in the plasma membrane, with procaspases associated to its cytoplasmic domain, which are released into the cytoplasm upon receptor activation.2 Bax3 and other proapoptotic members of the Bcl2 family like Puma and Noxa4, 5, 6 are involved in the mitochondrial pathway, promoting the mitochondrial release of cytochrome c, which in turn promotes procaspase cleavage to generate active caspases.2 In contrast, the mechanisms of Perp action in apoptosis are less well understood, but might be separate from the classical cell death pathways (reviewed in Ihrie et al7).
Perp (p53 apoptosis effector related to PMP22) was initially discovered in mouse embryonic fibroblasts (MEFs), in which it is upregulated during apoptosis, but not upon induced G1 arrest.8 Perp is a direct p53 target gene, with several, highly conserved, p53-binding sites in its promoter region.8 Furthermore, it is sufficient to induce apoptosis upon overexpression in MEFs,8 while loss of Perp function in Perp-deficient mice leads to reduced responsiveness of cells to extrinsic apoptotic stimuli.7 This effect is restricted to specific cell types, like thymocytes and neurons of the central nervous system, while the responsiveness of other cell types is not affected.7
Clues to the mechanisms of Perp function might come from data obtained for related proteins (Ihrie et al7; see also Discussion). Perp is a member of the peripheral myelin protein (PMP)-22 family, which also includes the epithelial membrane proteins EMP1-3, and the lens membrane protein MP-20. All family members are putative tetraspan transmembrane proteins, which localize to the plasma membrane and secretory pathway rather than the mitochondria.8 Similar to Perp, PMP22 and the EMPs can induce caspase-dependent apoptosis upon overexpression in cell culture.9, 10 However, members of the PMP22 family have also been implicated with other cellular processes, such as membrane trafficking and cell adhesion.
Detailed loss-of-function data about the in vivo role of PMP22-like proteins are only available for PMP22 itself. PMP22 is a major myelin component of Schwann cells, and both gain and loss of PMP22 function leads to peripheral neuropathies in mouse and man. Specifically, the hereditary neuropathy with liability to pressure palsies (HNPP) is caused by a deletion of the PMP22 gene,11, 12 while the demyelinating hereditary motor sensory neuropathy Charcot-Marie-Tooth disease type 1A (CMT1A) is due to a heterozygous duplication of the same DNA segment.13 In rare cases, CMT1A can also be caused by point mutations in the PMP22 gene, and in mice, similar mutations are associated with the Trembler and Trembler-J phenotype.14, 15, 16 In sum, these data suggest that both elevated and reduced levels of PMP22 can cause apoptosis and tissue degeneration.
Here, we describe a similar phenomenon for the PMP22-related Perp protein in zebrafish embryos. Like its mammalian homologs, zebrafish Perp has an apoptosis-inducing potential, and contributes to ultraviolet (UV) irradiation-induced and p53-dependent apoptosis in early zebrafish embryos. In contrast, Perp plays an antiapoptotic role during later stages of normal zebrafish development. In mouse, the phenotype caused by loss of Perp function has not been described in detail, it causes postnatal lethality.7 In zebrafish, antisense-based loss of Perp function leads to strong apoptosis specifically in the skin and the notochord of developing larvae, indicating that Perp is required for the survival of specific cell types during normal zebrafish development.
Results
Cloning and temporal expression pattern of zebrafish perp
Database searches for zebrafish Perp-homologs revealed several overlapping EST clones, one of which contains a full-length open reading frame encoding a single protein of 182 amino-acid residues (Genebank Accession numbers BM101781 and BM101553). Perp proteins represent a subclass of the Pmp22 family of putative transmembrane proteins. They are significantly conserved within the four putative membrane-associated regions, whereas the putative extracellular domains are very diverse.17, 18 Comparison of the deduced amino-acid sequence of the zebrafish clone with mammalian members of the Perp/PMP22 family revealed the highest overall similarity to the Perp subclass of the family (Figure 1a, 43.5 and 42.5% amino-acid identity to human and mouse Perp, respectively). In EST and genomic databases screens, we did not find any other zebrafish genes with higher similarities to Perp than to other subclasses of the family, suggesting that only one perp gene is present in zebrafish.
Zebrafish Perp is a member of the PMP22 family and expressed throughout zebrafish development. (a) Phylogenetic tree between different zebrafish, mouse and human members of the PMP22 family, assembled by the Jotun Hein method (DNASTAR Inc.), revealing that the zebrafish protein encoded by cloned cDNA is the Perp ortholog. (b) Semiquantitative RT-PCR analysis of zebrafish perp expression during embryonic development. As control, transcripts of the ef1α housekeeping gene19 were amplified. perp transcripts can be detected during all investigated stages, indicating maternal and zygotic expression of the perp gene
We next investigated the temporal expression pattern of perp during zebrafish development via semiquantitative RT-PCR. perp cDNA was detected already before midblastula transition, indicating a maternal contribution of perp mRNA. After the onset of zygotic transcription, elevated levels of perp cDNA could be seen, and equally high levels were found throughout all investigated stages of further development (Figure 1b).
Proapoptotic effects of zebrafish Perp upon overexpression in the zebrafish embryos
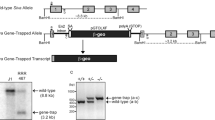
Perp is sufficient to induce apoptosis when overexpressed in MEFs.8 To investigate whether zebrafish Perp has a similar proapoptotic effect in zebrafish embryos, we injected in vitro-synthesized perp mRNA into one- to two-cell stage zebrafish embryos. Injected embryos proceed normally through gastrulation, but display a severely malformed body shape at 24 h after fertilization (hpf; Figure 2c). A comparable phenotype is seen upon p53 overexpression (data not shown). Employing TUNEL assays at the end of gastrulation (10 hpf), we found elevated levels of apoptosis in perp mRNA-injected embryos (Figure 2a, b), indicating that the morphological malformations result from enhanced cell death during gastrulation and segmentation stages. The phenotype can be suppressed by coinjecting an antisense morpholino oligonucleotide (perp MO) (Figure 2d). This MO efficiently blocks the translation of perp mRNA, as revealed in Western blots with protein extract from embryos injected with mRNA encoding a Perp-Myc fusion protein, probed with an anti-Myc antibody (Figure 2e). In contrast, an MO containing four nucleotide exchanges (perp 4 mm MO) does not block translation, showing that the perp MO is specific. We conclude that the apoptosis obtained upon perp mRNA injection is specifically induced by exogenous Perp protein encoded by the injected mRNA.
Overexpression of perp in zebrafish embryos leads to cell death. (a,b) TUNEL stainings of embryos at tailbud stage (end of gastrulation, 10 hpf), lateral views, anterior up, dorsal to the right; (a) embryo injected with perp mRNA (300 pg); (b) uninjected sibling. (c,d) The perp overexpression phenotype can be rescued by perp MO. Embryos were injected with 75 pg mRNA encoding a Perp-Myc fusion alone (c), or together with perp MO (0.45 pM) (d), or perp 4 mm control MO (0.45 pM); 24 hpf, lateral views, anterior to the left, dorsal up. perp mRNA-injected embryos show multiple defects, such as a short and bent-up axis, and a small head, resulting from severe apoptosis during earlier stages (c; 22/27), while perp-mRNA, perp-MO-injected embryo displays wild-type morphology (d; 27/28). Coinjection of perp mRNA with 4 mm perp MO yielded 25/31 affected embryos (not shown). (e) The perp MO used in panel (d) and in Figures 6, 7 and 8 efficiently targets perp mRNA. Western blot of protein extracts from shield stage embryos (6 hpf), injected either with mRNA encoding a Perp-Myc fusion and perp MO (left lane), or perp 4 mm control MO (right lane), probed with anti-Myc antibody. Concentrations are as in (c,d). The full-match but not the four mismatch control MO blocks translation of the injected mRNA
perp expression is increased upon UV-irradiation, and is required for UV-induced apoptosis
Several experiments were carried out addressing the role of perp and the correlation between p53 and perp in stressed zebrafish cells. In specific tissues of the mouse embryo, Perp is required to mediate stress-induced and p53-dependent apoptosis.7 An efficient way to induce p53-dependent apoptosis in zebrafish embryos is UV irradiation.20 Thus, irradiation of zebrafish embryos at a late blastula stage (sphere; 4 hpf) results in a significantly enhanced number of apoptotic cells at 10 hpf, as revealed by TUNEL staining (Figure 3d, e). This apoptosis can be suppressed by injection of a p53-specific MO that blocks the generation of endogenous p53 protein (Figure 3c, e). A similar, slightly weaker suppression of UV-induced apoptosis was obtained by injection of the perp MO (Figure 3a, e), whereas the perp 4 mm MO had no suppressing effect (Figure 3b, e). These data indicate that like p53, endogenous Perp protein is involved in UV-induced apoptosis in the early zebrafish embryo.
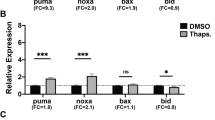
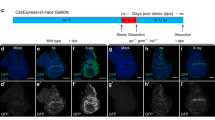
Perp contributes to UV- and p53-induced apoptosis in the zebrafish embryo. (a–g) UV treatment of zebrafish embryos leads to elevated perp expression levels, while UV-induced apoptosis is reduced in perp-deficient embryos. (a–d) TUNEL stainings of embryos at tailbud stage, after UV irradiation at sphere stage (late blastula, 4 hpf); animal views; injected MOs are indicated in lower right corners; due to the UV treatment, embryos fail to form a proper body axis;21 (a) embryo injected with perp MO; (b) embryo injected with perp 4 mm MO; (c) embryo injected with p53 MO; (d) uninjected sibling. (e) Diagram illustrating average numbers of TUNEL-positive cells shown in embryos as shown in (a–d). Number of evaluated embryos per lane: n=4. Standard deviations are indicated by error bars. (f) Semiquantitative RT-PCR to determine relative transcript levels of perp, and, as control, ef1α, in p53 morphant (lanes 1, 3) and wild-type siblings embryos (lanes 2, 4), after UV irradiation (lanes 3, 4), or nonirradiated (lanes 1, 2); tailbud-stage (10 hpf). (g) Semiquantitative time course RT-PCR to determine relative transcript levels of perp, mdm2, and, as control, ef1α after UV irradiation of p53 MO injected embryos (lanes 1, 3, 5, 7) and uninjected siblings (lanes 2, 4, 6, 8) at different time points after the UV irradiation: directly after the irradiation (0 h, 4 hpf, lanes 1, 2) and in 2 h intervals thereafter (2 h, lanes 3, 4; 4 h, lanes 5, 6; 6 h, lanes 7, 8). (h–l) p53 overexpression in zebrafish embryos leads to elevated perp expression levels, while p53-induced apoptosis is reduced in perp-deficient embryos. (h–j) TUNEL stainings of embryos at tailbud stage (10 hpf), animal views; injected mRNA and/or MOs are indicated in the lower right corners. Due to strong p53 overexpression, embryos fail to form a proper body axis. (k) Diagram illustrating average numbers of TUNEL-positive cells shown in embryos as shown in (h–j). Number of evaluated embryos per lane: n=3. (l) Semiquantitative time course RT-PCR to determine relative transcript levels of perp, p21, and, as control, ef1α, at different time points after the p53 mRNA injection. Injected embryos (lanes 1, 3, 5, 7) and uninjected control siblings (lanes 2, 4, 6, 8) were harvested at indicated stages: 256 cells stage (2hpf, lane 1, 2), sphere stage (4hpf, lane 3, 4), shield stage (6 hpf, lane 5, 6), and tailbud stage (10 hpf, lane 7, 8)
Accordingly, the increase in UV-induced and p53- and Perp-dependent apoptosis is correlated with an increase in perp transcription, as revealed by semiquantitative RT-PCR (Figure 3f). At 10 hpf, UV-treated embryos display several-fold higher perp mRNA levels than untreated embryos (Figure 3f, lanes 2 and 4). This increase in perp expression is partly dependent on p53, as it is less pronounced in p53-deficient embryos (Figure 3f, lanes 3 and 4). However, perp mRNA levels in irradiated p53 morphants are still higher than in nonirradiated morphant embryos (Figure 3f, lanes 1 and 3). Together, these findings point to both a p53-dependent and a p53-independent activation of perp expression in UV-irradiated zebrafish cells. Time course analyses further show that UV-induced transcriptional activation of perp occurs with the same kinetics as the transcriptional activation of the p53 target gene mdm2.20, 22 In both cases, higher transcript levels become apparent approximately 4 h after the irradiation (Figure 4g). However, in contrast to perp, UV-induced mdm2 expression strictly depends on p53, as no increase in mdm2 levels was obtained upon UV irradiation of p53 morphant embryos (Figure 4g, compare lanes 5 and 7 with lane 1).
perp is expressed in specific cells of the zebrafish embryo. In situ hybridization with perp antisense riboprobe at different stages of development. (a,b) Shield stage (early gastrula; 6 hpf). (a) Whole-mount embryo, (b) Sagital section. (c–e) 80% epiboly (midgastrula; 8 hpf). (c) Whole mount embryo. (d,e) Longitudinal sections, anterior up, arrow in (e) indicates single deep cells at the dorsal margin. (f) Tailbud stage (end of gastrulation; 10 hpf), transverse section through dorsal midline, dorsal up. perp is expressed in the notochord anlage (nc). (g–k) 25-somite stage (24 hpf). (g) Lateral overview over entire embryo; positions of section (h), magnified region (i), and optical cross-section (j,k) are indicated. (h) Transverse section through the tail of embryo stained for perp mRNA (in blue) and p63 protein (in brown), close-up on the skin. (i) Magnified view on the tail tip. perp shows prominent expression in the notochord. (j) Optical cross-section at level of eyes. (j) Optical cross-section at the level of the otic vesicles. (k,l) 36 hpf. (l) Dorsal view on the head. (m) Transverse section through trunk. (n,o) 48 hpf. (n) Higher magnification of pectoral fin. (o) Dorsal view on trunk. Abbreviations: bae, branchial arch epithelium; bel, basal epidermal layer; EVL, enveloping layer; epi, epidermis, skin; gcl, ganglion cell layer of retina; gt, gut; hed, hepatic duct; le, lense; li, liver; nc, notochord; oel, outer epidermal layer; olf, olfactory epithelium of nasal pits; ov, otic vesicle; pan, pancreas; pcp, prechordal plate; pfc, pectoral fin cartilage; pn, pronephric duct; vdi, ventral diencephalon; vhb, ventral hindbrain
perp expression is increased upon p53 overexpression, and is required for p53-induced apoptosis
In order to more directly assess the correlation between Perp and p53 during the induction of apoptosis, we performed p53 overexpression studies, injecting p53 mRNA into 1–4 cell stage zebrafish embryos (0 hpf). Similar to UV irradiated embryos, embryos injected with p53 mRNA display significantly elevated numbers of TUNEL-positive cells at 10 hpf (Figure 3j, k). This increase in cell death rates is strongly suppressed in embryos coinjected with p53 mRNA and perp MO (Figure 3h, k), but not in embryos coinjected with p53 mRNA and perp 4 mm MO (Figure 3i, k), indicating that perp is required to mediate p53-induced apoptosis. Along the same lines, p53 mRNA-injected embryos display higher levels of perp mRNA at the tailbud stage (10 hpf) (Figure 3l, lanes 7, 8). Time course analyses further show that this increase in perp expression does not occur right after p53 mRNA injection (Figure 3l, lanes 1–4), but only 5–6 h later, from late blastula/early gastrula stages onwards (Figure 3l, lanes 5, 6), when zygotic perp expression is initiated during normal development (Figure 3l, lanes 2, 4, 6, 8; compare with Figure 1b). A similar time course is also seen for the p53 target gene p21, which shows elevated transcript levels in p53 mRNA-injected embryos from mid-blastula stages onwards, coincident with the onset of p21 transcription in untreated embryos (Figure 3l). Together, these data indicate that in stressed zebrafish cells, perp and p21 are transcriptional targets of p53. However, p53 is not sufficient to activate their expression; rather, additional factors are required, which appear to be absent during cleavage and early blastula stages of zebrafish development.
Spatial expression pattern of perp during zebrafish development
We next investigated the spatial expression pattern of perp during normal zebrafish development by whole-mount in situ hybridization. The earliest restricted expression could be detected at the onset of gastrulation (shield stage, 6 hpf), when perp is expressed in a monolayer of cells covering the embryo (Figure 4a, b). These most likely represent the enveloping layer (EVL) cells that will later be shed. During later gastrulation, perp is also expressed by cells of the prechordal plate mesoderm, and in single mesodermal cells in dorsal marginal regions giving rise to the notochord (Figure 4c–e; 80% epiboly stage; 8 hpf). At the end of gastrulation (tailbud stage; 10 hpf), perp expression is prominent throughout the entire notochord anlage along the dorsal midline of the embryo (Figure 4f). During segmentation stages, the perp expression pattern becomes more complex. At 24 hpf, strong expression is detected in the notochord, eye vesicles, lenses, neural tissues of the brain (ventral diencephalon and hindbrain), otic vesicles, and skin (Figure 4g–k). In the skin, perp is expressed in both epidermal cell layers, the basal layer characterized by the coexpression of the putative keratinocyte stem cell factor p63,23, 24 and the outer keratinocyte layer (Figure 4h), while the EVL cells covering the embryo during earlier stages of development have already been shed.25 At 36 hpf, perp shows expression in the skin, nasal pits, gut, liver, and pronephric ducts (Figure 4i, m). At 48 hpf, perp remains expressed in the skin, the otic vesicle, and gut. In addition, the hepatic duct, pancreas, and cartilage-forming chondrocytes in the pectoral fins have started to express perp (Figure 4n, o).
During normal development, zebrafish perp is expressed independently of p53 and other members of the p53/63/73 family
Despite their upregulation in stressed cells, many of the p53 target genes display an unaltered expression in p53-deficient mouse embryos (see Discussion). The same appears to apply to perp expression during the early stages of zebrafish development, as revealed via comparative RT-PCR analyses of p53 morphant and wild-type sibling embryos (see Figure 3f, lanes 1, 2 for tailbud stage, 10 hpf). To examine whether the same is also true for perp expression during later stages of zebrafish embryogenesis, when perp displays a spatially restricted expression (see Figure 4), we first compared the expression pattern of perp with those of p53 and its relatives p63 and p73 in zebrafish embryos. At 48 hpf, p53 shows prominent expression in the brain, eyes, and gut22 (Figure 5a). p63 is expressed in the skin23, 24 (Figure 5b), overlapping with the epidermal expression of perp, but not in the other perp expression domains. However, even within the skin, the expression patterns of perp and p63 differ. While p63 expression is restricted to basal cells of the two-layered skin tissue, perp is expressed in both layers (see Figure 4h). p73 is expressed in specific cells of the telencephalon, in the pharyngeal endoderm, and in the pronephric ducts, showing coexpression with perp in the pronephric ducts only26, 27 (data not shown). Together, these data indicate that the three members of the p53/63/73 family are only expressed in a subset of the perp expression domains.
During normal zebrafish development, perp is expressed independently of p53 and the related proteins ΔNp63 and TAp73. (a,b,d–f) show whole-mount in situ hybridizations at 36 hpf; (a) p53 expression, which is most prominent in the brain, eyes, and gut. (b) p63 expression, most prominent in the skin. (c–e) perp expression in uninjected control embryo (c), and in p53 morphant (d), and ΔNp63 morphant embryo (e). perp expression in eyes, midbrain–hindbrain boundary region (mhb), and gut (gt) of p53 morphant, and in the skin (epi) of ΔNp63 morphant are indicated. (f) Comparative semiquantitative RT-PCR analysis of perp transcript levels in p53 morphants (lane 2), ΔNp63 morphants (lane 3), TAp73 morphants (lane 4), p53/TAp73 double-morphants (lane 5) and p53/Tap73/ΔNp63 triple-morphants (lane 6) at 48 hpf, with ef1α cDNA amplification as control. Efficiency of used p53 MO was confirmed by parallel coinjections with mdm2 MO, leading to a rescue of the mdm2 morphant phenotype, as described elsewhere.20 Efficiency of ΔNp63 and TAp73 MOs was confirmed by the skin and craniofacial phenotypes of injected sibling embryos at later stages of development23, 26
To investigate whether in such coexpressing domains, perp expression might depend on p53, p63, or p73 activity, we studied perp expression in embryos injected with the corresponding antisense MO. However, p53 morphants show unaltered perp expression in the eyes, hindbrain, and gut (Figure 5d). In contrast to p53, p63 and p73 can form different N-terminal isoforms, driven by alternative promoters. TA isoforms (TAp63, TAp73) contain an N-terminal transactivation domain similar to that of p53, and also act as transcriptional activators, whereas the N-terminally truncated isoforms (ΔNp63, ΔNp73) lack this domain, and are proposed to act as transcriptional repressors.28 During zebrafish embryogenesis, only ΔNp63 and TAp73 transcripts could be detected, while the other isoforms appear to be absent or expressed at much lower levels.23, 24, 27, 26 However, both ΔNp63 and TAp73 morphant embryos, although displaying specific developmental defects,23, 27, 26 show an unaltered perp expression pattern, including coexpressing tissues like skin and pronephric ducts (Figure 5e; and data not shown). Also, there was no indication that loss of any of the p53/63/73 family members affects the intensity of perp expression. Thus, semiquantitative RT-PCR analyses revealed indistinguishable amounts of perp transcripts in p53, ΔNp63, and TAp73 deficient animals, as well as in double or triple morphant embryos in which all three genes are knocked down (Figure 5f). In sum, we conclude that perp expression during normal development is largely independent of p53 family members.
Loss of Perp function leads to enhanced apoptosis rates in the skin and notochord
To study the function of perp during normal zebrafish development, we analyzed embryos that had been injected with perp antisense morpholino oligonucleotides (perp MOs). Morphologically, perp morphant embryos look normal until 30 hpf, when cells in the skin and fin folds appear round, rather than acquiring or maintaining a flattened shape (data not shown; but see below, Figure 8). At 48 hpf, the disorganization of epithelial cells of fins and skin becomes very severe. The tail fin is collapsed (Figure 6c, d), and the body skin appears very rough (Figure 6e, f). In addition, the pectoral fins are shortened and malformed (Figure 6g, h). Similarly, cells of the notochord, normally large and tightly packed in a ‘stack of pennies’-like fashion,29 appear round and disorganized (Figure 6c, d). A very similar notochord phenotype has been previously described for several zebrafish mutants with notochord degeneration.30, 31 Most likely resulting from this degeneration of the notochord, perp mutant embryos display an overall reduction of their body length. Particularly, the tail region does not extend properly (Figure 6a, b).
Loss of Perp causes caspase-dependent apoptosis. Acridine orange (a–f) and FITC-VAD-fmk (h,i) stainings of live embryos at 30 hpf; (a,c,e,h) injected with perp MO, (b) injected with perp MO and treated with caspase inhibitor zVAD-fmk, (c,f,i) injected with perp 4 mm MO, (d) uninjected control; (a–d,h,i) lateral view on tip of tail; (e,f) magnified lateral view on notochord. Note that in the notochord of perp morphants (e), all cells display altered, roundish shapes, while only some of the cells have become acridine orange positive. (g) Diagram illustrating average numbers of acridine orange-positive cells in embryos as shown in (a–d) within the tail region posterior of the anus. Number of evaluated embryos: control n=5; perp 4 mm MO-injected n=5; perp MO-injected n=6; perp 4 mm MO-injected+zVAD-FMK n=8
Loss of Perp causes tissue degeneration. All images show live embryos at 48 hpf, either injected with a perp-specific antisense MO (perp MO) (a,c,e,g), or the four mismatch control MO (4 mm) (b,d,f,h). (a,b) Overviews over entire embryos. (c–f) Higher magnifications of the tail region, (c,d) lateral views, (e,f) ventral views. The morphology of cells of the median fin fold and the notochord are changed in morphants (c; arrow points to roundly shaped cell), as compared to control embryos (d). The same is true for the trunk skin (e,f). (g,h) Dorsal view on the pectoral fins. Abbreviations: epi, skin; pf, pectoral fin; tf, tail fin
The defects in the skin, pectoral fins, and notochord of perp MO-injected embryos are consistent with the expression of perp in skin keratinocytes, pectoral fin chondrocytes, and notochord cells during normal development (Figure 4). Other areas of perp expression, however, appear unaffected in perp morphants, as assessed by the general morphology at 48 hpf and later stages, and in situ hybridization with the tissue-specific markers foxa3 (liver, pancreas32), pax2.1 (pronephric ducts33; otic vesicle34), and crystalline B1 (lens35) (data not shown).
Several experiments were performed to examine the cellular defects underlying the altered morphology of the affected tissues. Anti-Fibronectin and anti-pan Cadherin immunostainings25 revealed no obvious differences in the levels and distributions of these cell adhesion proteins (data not shown). Furthermore, immunostainings with an antiphosphorylated Histone antibody marking mitotic cells, and BrdU (5′ Bromo-2′-desoxyuridine) incorporation studies labelling replicating cells were normal, indicating that cell proliferation is not affected (data not shown). However, TUNEL stainings revealed a significant increase in the number of dying cells in the notochord, the pectoral fins, and the skin of perp morphant embryos (Figure 7a–f). Elevated cell death in the notochord and skin is already visible at 36 hpf (Figure 7e, f), when cells still show normal expression of marker genes, such as cytokeratin 8 in the skin,36 or shh37 and No tail/Brachyury38 in the posterior notochord (data not shown).
Loss of Perp causes cell death in specific embryonic tissues. TUNEL staining of embryos injected with perp MO (a, c, e), or the four mismatch control MO (4 mm MO) (b, d, f). (a–d) 48 hpf; (e, f) 36 hpf. (a,b) Whole-mount embryos; (c,d) magnified view on pectoral fins. (a) (e,f) 6 μm JB-4 transverse section through trunk at 36 hpf
Since TUNEL stainings of fragmented DNA do not discriminate between necrosis, apoptosis, and other forms of programmed cell death,39, 40 we also carried out apoptosis-specific stainings, using acridine orange (AO), a membrane-permeable aromatic derivative that becomes fluorescent in acidic lysosomal vesicles, thereby preferentially staining apoptotic cells41 (Figure 8a–g), and FITC-VAD-fmk, which stains activated caspases42 (Figure 8h, i). Already at 30 hpf, and coincident with changes in cell morphology, perp morphant embryos display severely increased numbers of AO-positive cells in the skin and notochord (Figure 8a, e, g), whereas embryos injected with the perp 4 mm control MO show normal numbers (Figure 8c, f, g). In perp morphants, the number of AO-positive cells can be significantly reduced by treating the embryos with the pan-caspase inhibitor zVAD-fmk,43 indicative for caspase-dependent apoptosis (Figure 8b, g). Consistently, perp morphant embryos display a dramatic increase in the number of FITC-VAD-fmk-positive cells (Figure 8h), whereas only few these cells were detected in embryos injected with perp 4 mm MO (Figure 8i). We conclude that during normal development, Perp is required for the survival of these specific cell types, leading to caspase-dependent apoptosis in specific tissues of perp-deficient embryos.
Discussion
In recent years, much progress has been made in understanding how the tumor-suppressing effect of p53 is mediated. Many transcriptional target genes of p53 have been found to be involved in p53-dependent stress responses to inhibit cell growth either through cell cycle arrest or induction of apoptosis.44 However, maybe discouraged by the finding that p53-deficient mice do not show striking developmental defects,45 less effort has been spent in exploring the role of these p53-inducible genes during normal development. In the case of the apoptosis-specific p53 target Perp, knockout mice have been generated and reported to be postnatally lethal.7 However, functional studies have only addressed the requirement of Perp to mediate stress-induced apoptosis in specific cell types, rather than its role during normal development.7 Here we report loss-of-function studies of the zebrafish perp ortholog, demonstrating that in contrast to its proapoptotic effect in stressed cells, it is required for the survival of notochord and skin cells during normal zebrafish development.
The perp expression pattern hints at functions during cell adhesion and cell differentiation in specific cell types
Thus far no detailed description of perp expression during the development of other vertebrate organisms has been reported. In zebrafish, perp is expressed at all developmental stages investigated. During later embryogenesis, its expression pattern is complex, which makes it difficult to speculate what perp-expressing cells might have in common, and what the role of Perp in these cell types might be. Clearly, perp expression has a preference for epithelium-forming tissues, such as the enveloping cell layer (EVL) during early stages of development, the skin during later stages, the olfactory epithelium of the nasal pits, the ear vesicles, the pronephric ducts, and the gut. In addition, it is expressed in tissues with a special mode of cell package and matrix secretion, such as the notochord, forming a rod-like structure stabilizing the body axis, the lenses, forming transparent structures bundling the light, and the cartilage-forming chondrocytes. Thus, Perp might be involved in conferring special adherence properties to expressing cells. Furthermore, it is expressed in the prechordal plate mesoderm during its migration from the marginal to the animal region of the gastrulating embryo, suggesting a potential role in cell migration and/or cell recognition, similar to that discussed for its relative Pmp22 in neural crest and sclerotomal cells of the zebrafish embryo.18
In most cell types, perp expression is only transient, coinciding with tissue differentiation. Thus, the EVL, the only tissue showing perp expression in early embryos, is the first tissue to differentiate, while the inner cell mass remains undifferentiated for several more hours.46 In the notochord, perp expression ceases at 36 hpf, in parallel with many other markers of differentiating notochord cells, such as the secreted protein Calymmin.47 In the lens, perp starts to be expressed around 24 hpf and vanishes again during the second day of development, very similar to the dynamics of connexin43 and 44, markers for differentiating lens fiber cells.48 Similarly, in the gastrointestinal tract,49, 50 pronephric ducts,51 and cartilage,52, 53 perp expression coincides with cell differentiation. In sum, it appears that in most tissues, the perp gene is expressed during cell differentiation processes, while it is switched off in terminally differentiated cells.
Zebrafish Perp has a conserved p53-dependent cell death-inducing potential, but is not a developmental target of the p53 family
The zebrafish Perp protein has a conserved apoptosis-inducing potential, as forced expression is sufficient to induce apoptosis in early zebrafish embryos, consistent with its proapoptotic effect in MEFs.8 In addition, like in mouse, zebrafish perp transcription can be activated by overexpression of p53, or by stressing cells via UV-irradiation, while the death-inducing effects of both treatments are significantly reduced in the absence of Perp, indicating that Perp is a transcriptional target and an essential mediator of the proapoptotic effect of p53 (Figure 3).
However, the epistatic relationship between p53 and perp is more complex. Thus, even in stressed cells, perp can be activated independently of p53, as indicated by our observation that UV irradiation of early zebrafish embryos can induce perp expression in the absence of p53, although at lower levels (Figure 3). Furthermore, p53 is not sufficient to induce perp expression, as indicated by our observation that elevated levels of perp mRNA were only obtained several hours after the p53 injection, most likely when thus far other unidentified regulatory factors have started to be expressed (Figure 3). During normal zebrafish development, perp expression appears to be completely independent of p53 and other members of the p53 family, as even the simultaneous loss of all currently known zebrafish p53 family members changesneither the spatial perp expression pattern nor its expression levels (Figures 3, 5). In sum, it appears that perp expression during normal development is governed by other factors, which might act in concert with p53 to achieve maximum, unphysiologically high levels of perp transcripts in stressed cells. The same appears to apply to other stress-induced p53 target genes, as revealed by comparative expression analysis in wild-type and p53-deficient mice, demonstrating both p53-dependent and p53-independent expression of Bax, Fas, Killer, and Mdm2 in different cell types of adult mice.54, 55
Perp and its PMP22 relatives: possible mechanisms of action
During embryonic development, several important regulators, such as members of the family of Bone Morphogenetic proteins, have both pro- and antiapoptotic effects, depending on the cellular context (see for example Liu et al56 and Israsena and Kessler57). However, the molecular mechanisms underlying these contrary effects are not fully understood. Here, we describe a similar phenomenon for the transmembrane protein Perp, which appears to promote or prevent cell death in a dose-dependent manner. Overexpression of zebrafish perp in early zebrafish embryos leads to enhanced apoptosis, similar to results obtained upon forced Perp expression in cell culture.8 In contrast, our loss-of-function studies clearly demonstrate that during normal development, Perp is essential for the survival of specific cell types. Thus, it appears that at a certain protein level, Perp is required for cell survival, whereas both lower and higher levels can lead to cell death. A similar dosage-dependent pro- or antiapoptotic effect has previously been described for the Perp-related protein PMP22, which can induce degenerative defects in Schwann cells of the peripheral nervous system upon both gain- and loss-of-function mutations (see Introduction). In the cell culture, massive forced expression of PMP22 induces apoptosis,9 whereas lower amounts of PMP22 interfere with the cell cycle, prolonging the G1 phase.58 Together, these point to a role of PMP22 at the choice point between G1 arrest and apoptosis, similar to that described for p53 (see Introduction).
A similar cell-cycle regulating function to induce G1 arrest and promote cell differentiation might also be attributed to Perp, which appears to be mainly required during tissue differentiation, but less for tissue maintenance, as perp expression is usually switched off after cells have differentiated (see above). Whether the apoptosis observed in perp morphant embryos is a default consequence of failed cell differentiation, or whether Perp plays a direct role in cell survival/death pathways, is currently unclear. However, both in the notochord and skin of perp morphants, apoptotic cells show normal expression of all differentiation markers, arguing for a direct cell survival function.
Currently, we can only speculate about the mode of such an antiapoptotic action. However, it is interesting to note that PMP22 is likely to function as an adhesion molecule.12, 59 Along these lines, it is interesting to note that in the affected tissues of zebrafish embryos, perp-deficient cells display alterations in cell shape coincident with or even prior to apparent signs of apoptosis (Figure 8). Proper cell adhesion can be essential for cell survival60 and differentiation.61 Thus, matrix detachment is known to activate specific intracellular pathways leading to apoptosis, also called anoikis.60, 62, 63, 64 In this light, Perp might block the anoikis-machinery to prevent cell death and allow transient matrix detachments, while cells change their shapes and relative positions during tissue differentiation. Such a function would be consistent with the observed expression of perp in prechordal plate cells, while they undergo active cell migration.65
Perp is only required in specific cells of the zebrafish embryo
Antisense-based knockdown of Perp function in the zebrafish embryo causes a rather late, but very specific phenotype, affecting only a subset of the cell types displaying perp expression throughout the different stages of embryonic development. The late appearance of the loss-of-function phenotype might be due to maternal contribution of Perp protein, which could, for example, compensate for the loss of zygotic protein in the cells of the EVL during gastrula stages. Another reason for the restricted effect of Perp loss of function, particularly during later stages of development, might be partial functional redundancy of Perp with other members of the PMP22 family. In mouse embryos, many of them are widely expressed.66, 67 In zebrafish, only pmp22 has been described thus far.18 Comparing the expression patterns of zebrafish perp and pmp22, obvious areas of overlapping expression are in the nasal pits, otic vesicles, and gut. However, embryos coinjected with pmp22 and perp MOs show no additional defects compared to perp single morphants (MN and MH, unpublished results), suggesting that further members of the family might be expressed in the unaffected cell types.
In conclusion, our findings demonstrate a dual role of Perp as a stress-dependent mediator of cell death as well as a tissue-specific survival factor. The biological significance of its function is underlined by the lethality of knockout mice, and the degenerative loss-of-function phenotype in the zebrafish described here. In the future, it will be important to further elucidate the mechanisms of Perp action that underlie its dose-dependent pro- and antiapoptotic effects.
Materials and Methods
Generation of constructs, mRNA synthesis, and microinjection
For expression constructs, the perp open reading frame was amplified via RT-PCR with the primers 5′-GGA ATT CAC CAT GTT CCG GTG TGG GAT C-3′ (sense) and 5′-GCT CTA GAC TTA GCC AGA CTG GTA GAT G-3′ (antisense) with Pfu DNA polymerase (Stratagene), and cloned into the EcoRI and XbaI sites of pCS2+.68 To test the efficiency of perp MOs (see below), perp cDNA containing parts of the 5′ UTR was amplified with primers 5′-CGG GAT CCC TTA ACA CTC GAC CAG AAC ACAC-3′ (sense) and 5′-CAC TAG TGC CAG ACT GGT AGA TGT ATT TGG TC-3′ (antisense), and cloned into the BamHI and SpeI sites of pCS2-6MT,68 encoding tagged Perp protein with six C-terminal Myc epitopes. Capped RNA was prepared with the Message Machine kit (Ambion), and injected into 1–2 cell stage embryos, as previously described (1.5 nl per embryo69). Amounts of injected RNA are given in the respective figures. p53 RNA was prepared and injected as described.20
Morpholino injections
Antisense MOs (Gene Tools) were dissolved in water to 1 mM. For injection, MOs were diluted in 1 × Danieu's buffer,70 and 1.5 nl per embryo was injected into embryos at the 1–2 cell stage, as described.69 MO sequences and injected concentrations were: perp MO 5′-CGG AAC ATT ATG CTT CTG ATA GAG T-3′, 0.3 pM/nl; perp 4 mm control MO 5′-CGG AtC ATT tTG CTT gTG ATt GAG T-3′, 0.3 pM/nl; p53 MO 5′-AGA ATT GAT TTT GCC GAC CTC CTC T-3′, 0.14 pM/nl. ΔNp63 MO and TAp73 MO were used as described.23, 26
Immunoblotting
Embryos were injected with 75 pg perp mRNA together with either perp MO or 4 mm-perp control MO, grown until shield stage (early gastrula, 6 hpf), and embryonic protein extracts were generated as described.71 Protein samples were separated via SDS-PAGE on 12% acrylamid/bis-acrylamid gels, and blotted on Hybond P membranes (Amersham). Immunoblotting analyses were performed using the anti-Myc antibody 9E10 (Roche Diagnostics, Mannheim, Germany), or an anti-BAP32 antibody72 as a positive control.
UV irradiation
Embryos were dechorionated in agarose-coated dishes and grown until sphere stage. Embryos were placed at 2.5 cm distance from a UV bulb (TUV 15W/G15 T8, Philips), irradiated for 20 s with 50 mJ/cm2, and fixed 6 h later.
Inhibition of caspase activity
To block caspase activity and thereby caspase-dependent apoptosis, zebrafish embryos were treated with the membrane-permeable pan-caspase inhibitor zVAD-fmk (carbobenzoxy-valyl-alanyl-aspartyl-[O-methyl]-fluoromethylketone, Promega, Madison, WI, USA),43 as described elsewhere.73
In situ hybridization, immunohistochemistry, and cell death detection
Whole-mount in situ hybridizations and immunohistochemistry were carried out as previously described.74 For perp in situ probe synthesis, plasmid pCS2-perp was linearized with BamHI and transcribed with T7 RNA polymerase. In addition, riboprobes of zebrafish p5322 and zebrafish p6323 were used. p63 protein was stained, using the p63-specific mouse monoclonal antibody 4a4 (Santa Cruz Biotechnology, CA, USA).24 Mitotic cells were stained with an polyclonal rat antiphosphorylated histone H3 antibody (Upstate Biotechnology, NY, USA). Immunostainings with anti-Fibronectin (Sigma, F3648, 1 : 200) and anti-pan Cadherin antibodies were carried out using reported concentrations and protocols.25, 75
For detection of apoptotic cells, terminal desoxynucleotidyl transferase-mediated biotinylated UTP nick end labelings (TUNEL) were performed, using the In Situ Cell Death Detection Kit (Roche Diagnostics, Mannheim, Germany), essentially as described elsewhere.76 For detection of apoptotic cells in live embryos, AO and FITC-VAD-fmk stainings were performed. For AO stainings, dechorionated embryos were incubated in 5 μg/ml AO (Sigma Chemical Co., St. Louis, MO, USA) in E3 medium for 30 min and analyzed by fluorescent microscopy. For FITC-VAD-fmk stainings, dechorionated embryos were placed in 10 μM FITC-VAD-fmk (Promega, Madison, WI, USA) solution in E3 at sphere stage (4 hpf). At 24 hpf, embryos were transferred to fresh solution, and analyzed by fluorescent microscopy. Fluorescent images were taken on a Zeiss Axiophot microscope (Carl Zeiss, Jena, Germany), using an ORCA ER C4742-95 digital camera (Hamamatzu, Bridgewater, NJ, USA). Bright-field and fluorescent images were superimposed with Openlab software (Improvision, Lexington, MA, USA).
Histology
Embryos younger than 24 hpf were embedded in paraffin wax and 6-μm-thick sections were cut with Feather S35 microtome blades on a Leica RM 2155 Microtome. Sections were dewaxed using Rotihistol and mounted in Rotihistokit (Roth, Germany). Embryos of 24 hpf or older were embedded in JB4 plastic (Polysciences, Inc.). In total, 6 μm sections were cut using Leica TC-65 blades on a Leica RM 2155 Microtome. Sections were directly mounted in Permount (Fisher Chemicals).
RT-PCRs
Zebrafish embryos were collected at specific time points after fertilization. Total RNA from whole embryos was isolated using Trizol-LS reagent (Gibco) and cDNA was generated using Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA, USA), followed by amplification of perp cDNA with the primers listed above. Primers used to amplify cDNAs were as follows: perp, sense 5′-CTT AGC CAG ACT GGT AGA TG-3′, antisense 5′-TTA GCC AGA CTG GTA GAT G-3′; p21,77 sense 5′-AGC TGC ATT CGT CTC GTA GC-3′, antisense 5′-TGA GAA CTT ACT GGC AGC TTC A-3′; mdm2,77 sense 5′-TTG GGA GTT GCC TTG TGG-3′; antisense 5′-AGA CAG TCT TCG CAG CAG-3′; ef1α78 sense 5′-TCA CCC TGG GAG TGA AAC AGC-3′, antisense 5′-ACT TGC AGG CGA TGT CAG CAG-3′.
Abbreviations
- AO:
-
acridine orange
- BrdU:
-
5′ Bromo-2′-desoxyuridine
- ΔNp63:
-
amino-terminally truncated isoform of p63
- EST:
-
expressed sequence tag
- EVL:
-
enveloping layer
- hpf:
-
hours postfertilization
- mdm2:
-
mouse double minute 2
- MO:
-
morpholino oligonucleotide
- RT-PCR:
-
polymerase chain reaction after reverse transcription of RNA
- SDS-PAGE:
-
sodiumdodecylsulfate polyacrylamide gel electrophoresis
- TAp73:
-
transactivating N-terminally full-length isoform of p73
- TUNEL:
-
terminal deoxynucleotidyl transferase-mediated biotinylated UTP nick end labeling
- UV:
-
ultraviolet
- 4 mm:
-
four mismatch control
References
Wu GS, Burns TF, McDonald ER, Jiang W, Meng R, Krantz ID, Karo G, Gan DD, Zhou JY, Muschel R, Hamilton SR, Spinner NB, Markowitz S, Wu G and El-Deiry WS (1997) KILLER/DR5 is a DNA damage-inducible p53-regulated death receptor gene. Nat. Genet. 17: 141–143
Hengarter MO (2000) The biochemistry of apoptosis. Nature 407: 770–776
Mayashita T and Reed JC (1995) Tumor suppressor p53 is a direct transcriptional activator of the human BAX gene. Cell 80: 293–299
Jeffers JR, Paraganas E, Lee Y, Yang C, Wang J, Brennan J, MacLean KH, Jan J, Chittenden T, Ihle JN, McKinnon PJ, Cleveland JL and Zambetti GP (2003) Puma is an essential mediator of p53-dependent and -independent apoptotic pathways. Cancer Cell 4: 321–328
Villunger A, Michalak EM, Coultas L, Mullauer F, Bock G, Ausserlechner MJ, Adams JM and Strasser A (2003) p53- and drug-induced apoptotic responses mediated by the BH3-only proteins puma and noxa. Science 302: 1036–1038
Shibue T, Takeda K, Oda E, Tanaka H, Murasawa H, Takaoka A, Marishita Y, Akira S, Tanigushi T and Tanaka N (2003) Integral role of Noxa in p53-mediated apoptosis. Genes Dev. 17: 2233–2238
Ihrie RA, Reczek E, Horner JS, Khachatrian L, Sage J, Jacks T and Attardi LD (2003) Perp is a mediator of p53-dependent apoptosis in diverse cell types. Curr. Biol. 13: 1985–1990
Attardi LD, Reczek EE, Cosmas C, Demicco EG, McCurrach ME, Lowe SW and Jacks T (2000) PERP, an apoptosis-associated target of p53, is a novel member of the PMP-22/gas3 family. Genes Dev. 14: 704–718
Fabretti E, Edomi P, Brancolini C and Schneider C (1995) Apoptotic phenotype induced by overexpression of wild-type gas2/PMP22: its correlation to the demyelinating peripheral neuropathy CMTA1. Genes Dev. 9: 1846–1856
Wilson HL, Wilson SA, Surprenant A and North RA (2002) Epithelial membrane proteins induce membrane blebbing and interact with the P2X7 C terminus. J. Biol. Chem. 277: 34017–34023
Chance PF, Alderson MK, Leppig KA, Lensch MW, Matsunami N, Smith B, Swanson PD, Odelberg SJ, Disteche CM and Bird TD (1993) DNA deletion associated with hereditary neuropathy with liability to pressure palsies. Cell 72: 143–151
Adlkofer K, Martini R, Aguzzi A, Zielasek J, Toyka KV and Suter U (1995) Hypermyelination and demyelinating peripheral neuropathy in Pmp22-deficient mice. Nat. Genet. 11: 274–280
Valentijn LJ, Bolhuis PA, Zorn I, Hoogendijk JE, van den Bosch N, Hensels GW, Stanton Jr VP, Housman DE, Fischbeck KH and Ross DA (1992) The peripheral myelin gene PMP-22/GAS-3 is duplicated in Charcot-Marie-Tooth disease type 1A. Nat. Genet. 1: 166–170
Suter U, Moskow JJ, Welcher AA, Snipes GJ, Kosaras B, Sidman RL, Buchberg AM and Shooter EM (1992) A leucine-to-proline mutation in the putative first transmembrane domain of the 22-kDa peripheral myelin protein in the trembler-J mouse. Proc. Natl. Acad. Sci. USA 89: 4382–4386
Suter U, Welcher AA, Ozcelik T, Snipes GJ, Kosaras B, Francke U, Billings-Gagliardi S, Sidman RL and Shooter EM (1992) Trembler mouse carries a point mutation in a myelin gene. Nature 356: 241–244
Valentijn LJ, Baas F, Wolterman RA, Hoogendijk JE, van den Bosch NH, Zorn I, Gabreels-Festen AW, de Visser M and Bolhuis PA (1992) Identical point mutations of PMP-22 in Trembler-J mouse and Charcot-Marie-Tooth disease type 1A. Nat. Genet. 2: 288–291
Naef R and Suter U (1998) Many facets of the peripheral myelin protein PMP22 in myelination and disease. Microsc. Res. Tech. 41: 359–371
Wulf P, Bernhardt RR and Suter U (1999) Characterization of peripheral myelin protein 22 in zebrafish (zPMP22) suggests an early role in the development of the peripheral nervous system. J. Neurosci. Res. 57: 467–478
Nordness S, Krauss S and Johansen T (1994) cDNA sequence of zebrafish (Brachydanio rerio) translation elongation factor-1α: molecular phylogeny of eukaryotes based on elongation factor-1α protein sequences. Biochim. Biophys. Acta 1219: 529–532
Langheinrich U, Hennen E, Stott G and Vacun G (2002) Zebrafish as a model organism for the identification and characterization of drugs and genes affecting p53 signaling. Curr. Biol. 12: 2023–2028
Jesuthasan S and Strähle U (1997) Dynamic microtubules and specification of the zebrafish embryonic axis. Curr. Biol. 7: 31–42
Thisse C, Neel H, Thisse B, Daujat S and Piette J (2000) The Mdm2 gene of zebrafish (Danio rerio): preferential expression during development of neural and muscular tissues, and absence of tumor formation after overexpression of its cDNA during early embryogenesis. Differentiation 66: 61–70
Bakkers J, Hild M, Kramer C, Furutani-Seiki M and Hammerschmidt M (2002) Zebrafish ΔNp63 is a direct target of Bmp signaling and encodes a transcriptional repressor blocking neural specification in the ventral ectoderm. Dev. Cell 2: 617–627
Lee H and Kimelman D (2002) A dominant-negative from of p63 is required for epidermal proliferation in zebrafish. Dev. Cell 2: 607–616
Crawford BD, Henry CA, Clason TA, Becker AL and Hille MB (2003) Activity and distribution of Paxillin, focal adhesion kinase, and cadherin indicate cooperative roles during zebrafish embryogenesis. Mol. Biol. Cell 14: 3063–3081
Rentzsch F, Kramer C and Hammerschmidt M (2003) Specific and conserved roles of TAp73 during zebrafish development. Gene 323: 19–30
Pan H, Dung H-N, Hsu H-M, Hsiao K-M and Chen L-Y (2003) Cloning and developmental expression of p73 cDNA in zebrafish. Biochem. Biophys. Res. Commun. 307: 395–400
Yang A and McKeon F (2000) P63 and P73: P53 mimics, menaces and more. Nat. Rev. Mol. Cell Biol. 1: 199–207
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B and Schilling TF (1995) Stages of embryonic development of the zebrafish. Dev. Dyn. 203: 253–310
Odenthal J, Haffter P, Vogelsang E, Brand M, Van Eeden FJM, Furutani-Seiki M, Granato M, Hammerschmidt M, Heisenberg C-P, Jiang Y-J, Kane DA, Kelsh RN, Mullins MC, Warga RM, Allende ML, Weinberg ES and Nüsslein-Volhard C (1996) Mutations affecting the formation of the notochord in the zebrafish, Danio rerio. Development 123: 103–115
Stemple DL, Solnica-Krezel L, Zwartkruis F, Neuhauss SCF, Schier AF, Malicki J, Stanier DYR, Abdelilah S, Rangini Z, Mountcastle-Shah E and Driever W (1996) Mutations affecting development of the notochord in zebrafish. Development 123: 117–128
Odenthal J and Nüsslein-Volhard C (1998) forkhead domain genes in zebrafish. Genes Dev. Evol. 208: 245–258
Majumdar A, Lun K, Brand M and Drummond IA (2000) Zebrafish no isthmus reveals a role for pax2.1 in tubule differentiation and patterning events in the pronephric primordia. Development 127: 2089–2098
Pfeffer PL, Gerster T, Lun K, Brand M and Busslinger M (1998) Characterization of three novel members of the zebrafish Pax2/5/8 family: dependency of Pax5 and Pax8 expression on the Pax2.1 (noi) function. Development 125: 3063–3074
Chen JY, Chang BE, Chen YH, Lin CJ, Wu JL and Kuo CM (2001) Molecular cloning, developmental expression, and hormonal regulation of zebrafish (Danio rerio) beta crystallin B1, a member of the superfamily of beta crystallin proteins. Biochem. Biophys. Res. Commun. 285: 105–110
Imboden M, Goblet C, Korn H and Vriz S (1997) Cytokeratin 8 is a suitable epidermal marker during zebrafish development. CR Acad Sci III 320: 689–700
Krauss S, Concordet J-P and Ingham PW (1993) A functionally conserved homolog of the Drosophila segment polarity gene hh is expressed in tissues with polarizing activity in zebrafish embryos. Cell 75: 1431–1444
Schulte-Merker S, Eeden FV, Halpern ME, Kimmel CB and Nüsslein-Volhard C (1994) No tail (ntl) is the zebrafish homologue of the mouse T (Brachyury) gene. Development 120: 1009–1015
Abraham MC and Shabam S (2004) Death without caspases, caspases without death. Trends Cell Biol. 14: 184–193
Guimaraes CA and Linden R (2004) Programmed cell death: apoptosis and alternative deathstyles. Eur. J. Biochem. 271: 1638–1650
Abrams JM, White K, Fessler LI and Steller H (1993) Programmed cell death during Drosophila embryogenesis. Development 117: 29–43
Pozarowski P, Huang X, Halicka DH, Lee B, Johnson G and Darzynkiewicz Z (2003) Interaction of fluorochrome-labeled caspase inhibitors with apoptotic cells: a caution in data interpretation. Cytometry 55A: 50–60
Li M, Ona VO, Guégan C, Chen M, Jackson-Lewis V, Andrews LJ, Olszewski AJ, Stieg PE, Lee J-P, Przedborski S and Friedlander RM (2000) Functional role of caspase-1 and caspase-3 in an ALS transgenic mouse model. Science 288: 335–338
Vousden KH and Lu X (2002) Live or let die: the cell's response to p53. Nat. Rev. Cancer 2: 594–604
Donehower LA, Harvey M, Slagle BL, McArthur MJ, Montgomery Jr CA, Butel JS and Bradley A (1992) Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 356: 215–221
Kimmel CB, Kane DA and Ho RK (1991) In Cell–Cell Interactions in Early Development Gerhart J, ed (New York: Wiley-Liss) pp. 203–225
Cerda J, Grund C, Franke WW and Brand M (2002) Molecular characterization of Calymmin, a novel notochord sheath-associated extracellular matrix protein in the zebrafish embryo. Dev. Dyn. 224: 200–209
Cheng S, Christie T and Valdimarsson G (2003) Expression of connexin48.5, connexin44.1, and connexin43 during zebrafish (Danio rerio) lens development. Dev. Dyn. 228: 709–715
Biemar F, Argenton F, Schmidtke R, Epperlein S, Peers B and Driever W (2001) Pancreas development in zebrafish: early dispersed appearance of endocrine hormone expressing cells and their convergence to form the definitive islet. Dev. Biol. 230: 189–203
Field HA, Ober EA, Roeser T and Stainier DY (2003) Formation of the digestive system in zebrafish. I. Liver morphogenesis. Dev. Biol. 253: 279–290
Drummond I (2003) Making a zebrafish kidney: a tale of two tubes. Trends Cell Biol. 13: 357–365
Grandel H and Schulte-Merker S (1998) The development of the paired fins in the zebrafish (Danio rerio). Mech. Dev. 79: 99–120
Kimmel CB, Miller CT and Moens CB (2001) Specification and morphogenesis of the zebrafish larval head skeleton. Dev. Biol. 233: 239–257
Bouvard V, Zaitchouk T, Vacher M, Duthu A, Canivet M, Choisy-Rossi C, Nieruchalski M and May E (2000) Tissue and cell-specific expression of p53-target genes: bax, fas, mdm2 and waf1/p21, before and following ionising irradiation in mice. Oncogene 19: 649–660
Burns TF, Bernhardt EJ and El-Deiry WS (2001) Tissue specific expression of p53 target genes suggest a key role for KILLER/DR5 in p53-dependent apoptosis in vivo. Oncogene 20: 4601–4612
Liu J, Wilson SI and Reh T (2003) BMP receptor 1b is required for axon guidance and cell survival in the developing retina. Dev. Biol. 256: 34–47
Israsena N and Kessler JA (2002) Msx2 and p21CIP1/WAF1 mediate the proapoptotic effects of Bone Morphogenetic Protein-4 on the ventricular zone progenitor cells. J. Neurosci. Res. 69: 803–809
Zoidl G, D'Urso D, Blass-Kampmann S, Schmalenbach C, Kuhn R and Müller HW (1997) Influence of elevated expression of rat wild-type PMP22 and its mutant PMP22Trembler on cell growth of NIH3T3 cells. Cell Tissue Res. 287: 459–470
Yoshikawa H and Dyck PJ (1991) Uncompacted inner myelin lamellae in inherited tendency to pressure palsy. J. Neuropathol. Exp. Neurol. 50: 649–657
Grossman J (2002) Molecular mechanisms of ‘detachment-induced apoptosis – anoikis’. Apoptosis 7: 247–260
Danen EH and Sonnenberg A (2003) Integrins in regulation of tissue development and function. J. Pathol. 201: 632–641
Giancotti FG (1997) Integrin signaling: specificity and control of cell survival and cell cycle progression. Curr. Opin. Cell Biol. 9: 691–700
Frisch SM and Ruoslahti E (1997) Integrins and anoikis. Curr. Opin. Cell Biol. 9: 701–706
Stupack DG and Cheresh DA (2004) A bit-role for integrins in apoptosis. Nat. Cell Biol. 6: 388–389
Winklbauer R and Selchow A (1992) Motile behaviour and protrusive activity of migratory mesoderm cells from Xenopus gastrula. Dev. Biol. 150: 335–351
Taylor V, Welcher AA, Program AE and Suter U (1995) Epithelial membrane protein-1, peripheral myelin protein 22, and lens membrane protein 20 define a novel gene family. J. Biol. Chem. 270: 28824–28833
Taylor V and Suter U (1996) Epithelial membrane protein-2 and epithelial membrane protein-3: two novel members of the peripheral myelin protein 22 gene family. Gene 175: 115–120
Rupp RAW, Snider L and Weintraub H (1994) Xenopus embryos regulate the nuclear localization of XMyoD. Genes Dev. 8: 1311–1323
Hammerschmidt M, Blader P and Strähle U (1999) In Methods in Cell Biology Detrich HW, Westerfield M, Zon LI, eds (San Diego: Academic Press) pp. 87–115
Nasevicius A and Ekker SC (2000) Effective targeted gene ‘knockdown’ in zebrafish. Nat. Genet. 26: 216–220
Westerfield M (1994) The Zebrafish Book: A Guide for the Laboratory Use of Zebrafish. Eugene, Oregon: University of Oregon Press
Adachi T, Schamel WW, Kim KM, Watanabe T, Becker B, Nielsen PJ and Reth M (1996) The specific association of the IgD molecule with the accessory proteins BAP31/BAP29 lies in the IgD transmembrane sequence. EMBO J. 7: 1534–1541
Sanders LH and Whitlock KE (2003) Phenotype of the zebrafish masterblind (mbl) mutant is dependent on genetic background. Dev. Dyn. 227: 291–300
Hammerschmidt M, Pelegri F, Mullins MC, Kane DA, van Eeden FJM, Granato M, Brand M, Furutani-Seiki M, Haffter P, Heisenberg C-P, Jiang Y-J, Kelsh RN, Odenthal J, Warga RM and Nüsslein-Volhard C (1996) dino and mercedes, two genes regulating dorsal development in the zebrafish embryo. Development 123: 95–102
Bakkers J, Kramer C, Pothof J, Quaedvlieg NEM, Spaink HP and Hammerschmidt M (2004) Hyaluronan synthase 2 (Has2) is required upstream of Rac1 to govern dorsal migration of lateral cells during zebrafish gastrulation. Development 131: 525–537
Cole LK and Ross LS (2001) Apoptosis in the developing zebrafish embryo. Dev. Biol. 240: 123–142
Liu TX, Howlett NG, Deng M, Langenau DM, Hsu K, Rhodes J, Kanki JP, D'Andrea AD and Look AT (2003) Knockdown of zebrafish Fancd2 causes developmental abnormalities via p53-dependent apoptosis. Dev. Cell 5: 903–914
Nordness S, Krauss S and Johansen T (1994) cDNA sequence of zebrafish (Brachydanio rerio) translation elongation factor-1 alpha: molecular phylogeny of eukaryotes based on elongation factor-1 alpha protein sequences. Biochim. Biophys. Acta 1219: 529–532
Acknowledgements
We thank A Vortkamp for fruitful discussions, J Odenthal, C-M Kuo, P Ingham, ML Martorana, S Schulte-Merker for sending reagents, and V Taylor for critical reading of the manuscript. Work in the lab of MH was supported by the Max-Planck society and the National Institute of Health (NIH grant 1R01-GM63904). MN was supported by a long-term predoctoral fellowship by the Boehringer Ingelheim Fonds, Stuttgart.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by V De Laurenzi
Rights and permissions
About this article
Cite this article
Nowak, M., Köster, C. & Hammerschmidt, M. Perp is required for tissue-specific cell survival during zebrafish development. Cell Death Differ 12, 52–64 (2005). https://doi.org/10.1038/sj.cdd.4401519
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401519
Keywords
This article is cited by
-
The PMP22 Gene and Its Related Diseases
Molecular Neurobiology (2013)
-
PERP expression stabilizes active p53 via modulation of p53-MDM2 interaction in uveal melanoma cells
Cell Death & Disease (2011)
-
An essential function for the centrosomal protein NEDD1 in zebrafish development
Cell Death & Differentiation (2010)
-
Transgenic expression of prothymosin alpha on zebrafish epidermal cells promotes proliferation and attenuates UVB-induced apoptosis
Transgenic Research (2010)
-
The zebrafish as a model organism for the study of apoptosis
Apoptosis (2010)