Abstract
Plant embryogenesis is intimately associated with programmed cell death. The mechanisms of initiation and control of programmed cell death during plant embryo development are not known. Proteolytic activity associated with caspase-like proteins is paramount for control of programmed cell death in animals and yeasts. Caspase family of proteases has unique strong preference for cleavage of the target proteins next to asparagine residue. In this work, we have used synthetic peptide substrates containing caspase recognition sites and corresponding specific inhibitors to analyse the role of caspase-like activity in the regulation of programmed cell death during plant embryogenesis. We demonstrate that VEIDase is a principal caspase-like activity implicated in plant embryogenesis. This activity increases at the early stages of embryo development that coincide with massive cell death during shape remodeling. The VEIDase activity exhibits high sensitivity to pH, ionic strength and Zn2+ concentration. Altogether, biochemical assays show that VEIDase plant caspase-like activity resembles that of both mammalian caspase-6 and yeast metacaspase, YCA1. In vivo, VEIDase activity is localised specifically in the embryonic cells during both the commitment and in the beginning of the execution phase of programmed cell death. Inhibition of VEIDase prevents normal embryo development via blocking the embryo-suspensor differentiation. Our data indicate that the VEIDase activity is an integral part in the control of plant developmental cell death programme, and that this activity is essential for the embryo pattern formation.
Similar content being viewed by others
Introduction
The roles of programmed cell death in development are highly conserved for both metazoans and plants and include targeted elimination of surplus, ‘no-longer-needed’, damaged and abnormal cells. In spite of the growing list of examples of developmental cell death in plants (reviewed in ref.1), very little is known about its mechanisms. Direct search for the plant orthologs of the core apoptotic cell-death proteins has shown that neither Bcl-2 family members, nor Apaf-1-related proteins or canonical caspases are present in plants.2,3 Furthermore, numerous cytological observations suggest that plants employ a nonapoptotic pathway of cell dismantling defined as autophagic cell death.1,4 The hallmark feature distinguishing this type of cell death from apoptosis is that complete cellular degradation takes place inside the dying cell itself without active participation of phagocytes.5
Notwithstanding a lack of direct plant homologs of mammalian caspases, the possibility that plants may possess enzymes with caspase-like function during programmed cell death has recently commanded the attention of cell death researchers. There are two lines of evidence supporting this possibility. First, Uren et al.6 have identified genes encoding ancestral caspase-like proteins, the metacaspases, which are present in plants, fungi and protozoa. Homology of metacaspases to caspases is not restricted to the primary sequence, including the catalytic diad of histidine and cysteine, but extends to the secondary structure as well. Although the implication of plant metacaspases in programmed cell death has not been shown so far, yeast metacaspase YCA1 has been demonstrated as an effective executor for cell death, displaying increased proteolytic activity against caspase-6-specific synthetic substrate VEID-7-amino-4-methylcoumarin (AMC).7 Second, there is a growing body of physiological and biochemical data suggesting that plant cells contain caspase-1- (i.e., YVADase) and/or 3-like (i.e., DEVDase) proteolytic activities,8 which correlate with the level of experimentally induced cell death,9,10,11,12,13 and that the latter can be abolished or delayed by synthetic9,10,11,12,13 or natural14,15 caspase-specific inhibitors. As most of these experiments dealt with pathogen- or drug-induced cell deaths, they cannot address the roles of caspase-related proteins (metacaspases) and caspase-like proteolytic activity in normal plant development. In animals, however, caspases (at least caspases-3, -7, -8 and -9) are well known to be indispensable for both embryonic and postembryonic development and survival.16,17,18,19
To examine the involvement of caspase-like activity in plant development, we have chosen Norway spruce somatic embryogenesis as a biological system. This system provides a unique opportunity to control the entire pathway of plant embryo development in laboratory conditions.20 Using time-lapse tracking technique, we have characterised previously all the successive stages in this developmental pathway, including the stages of proembryogenic mass, proembryogenic mass-to-embryo transition, early embryogeny and late embryogeny.21 We found that somatic embryo development of Norway spruce is similar to zygotic embryogeny of Pinaceae, even though embryo origin is different in each case (i.e., somatic cells in proembryogenic mass versus zygote).21,22 Our more recent studies suggest that autophagic programmed cell death regulates active shape remodeling during both somatic and zygotic embryo development,23,24 and that dysregulation of this cell death leads to embryonic aberrations.25 Using somatic embryos of Norway spruce, we characterized the cell biological pathway of autophagic death in the suspensor, a temporal structure, which is composed of the cells at successive steps of the death pathway and is finally eliminated during late embryogeny.23,25
In the present study, we address the question of whether caspase-like proteolytic activity is involved in the regulation of plant developmental cell death using Norway spruce somatic embryogenesis as the model system. We showed, for the first time, that VEIDase is a principal caspase-like activity implicated in plant embryogenesis. This activity is increased at the early stages of embryo development, and is directly involved in the terminal differentiation and death of the embryo suspensor.
Results
Cell extracts of Norway spruce display caspase-like activity with a strong preference for VEID-containing substrate
To determine whether or not embryogenic cultures of Norway spruce contain any caspase-like proteolytic activity, we compared the ability of a large number (>50) of cell extracts prepared at different stages of embryogenesis to cleave six peptide substrates specific for distinct members of mammalian caspase family proteases. The substrates tested were YVAD-AMC (for caspase-1), VDVAD-AMC (for caspase-2), DEVD-AMC (for caspases-3 and -7), VEID-AMC (for caspase-6), IETD-AMC (for caspases-8 and –6) and LEHD-AMC (for caspase-9). We observed strikingly constant relative rates for proteolytic activities against six peptide substrates in all tested extracts (Figure 1). Pearson correlation coefficients (r) computed for mean proteolytic activities of cell extracts against different substrates, pairwise for each combination of two substrates, were in the range from 0.79 to 0.90. In the same time, the absolute cleavage rates of individual substrates varied significantly depending both on in vitro assay conditions and on the stage of embryogenesis when extract was prepared (see following results). VEID-AMC was found to be the most efficiently cleaved substrate (Figure 1). Next in order of cleavage rate to VEID-AMC was IETD-AMC (≈44% of the VEID-AMC cleavage rate) followed by four other peptides whose relative cleavage rates were 7–20 times lower than that of VEID-AMC (Figure 1). The similar pattern of relative cleavage rates of synthetic peptides, with the strongest preference for Val in P4 position (as in VEID) and tolerance for Ile in the same position (as in IETD), is a distinguishing characteristic of mammalian caspase-6.26
In vitro cleavage of peptide substrates specific for different mammalian caspases by a large number (>50) of cell extracts prepared at different stages of Norway spruce embryogenesis. Graph represents mean rates (±S.E.M.) of peptide cleavage relative to VEID-AMC (%). The assay was performed under standard conditions (Materials and Methods)
In general, caspase-like activities cannot be inhibited by protease inhibitors other than caspase-specific ones.8,27 Accordingly, we observed no inhibitory effect of a range of serine and cysteine protease inhibitors (calpain inhibitor I, aprotinin, leupeptin, phenylmethylsulphonyl fluoride (PMSF) and L-1-chloro-3-(4-tosylamido)-7-amino-2-heptanone (TLCK)), or of pepstatin (inhibitor for aspartic acid proteases) or lactacystine (inhibitor for proteasome), on the cleavage of VEID-AMC by cell extracts of Norway spruce (Figure 2a). In contrast, caspase-6 substrate-mimetic inhibitor Ac-VEID-CHO almost completely inhibited this activity at the concentration of 10 μM (Figure 2b). While inhibitory effects of aldehyde (CHO) caspase inhibitors other than Ac-VEID-CHO were also observed, they were substantially weaker than the one exerted by Ac-VEID-CHO (Figure 2a, data not shown).
Effects of protease inhibitors on Norway spruce VEIDase activity in vitro. (a) Cell extracts were incubated with lactacystine (20 μM), calpain inhibitor I (20 μM), pepstatin (1 μM), aprotinin (1 μM), leupeptin (100 μM), PMSF (250 μM), TLCK (50 μM), Ac-VEID-CHO (2 μM) or Ac-LEHD-CHO (2 μM), in the presence of substrate VEID-AMC. The data are expressed as a percent of mean VEIDase activity relative to control (no inhibitor; ±S.E.M., n=3). (b) The dose–response curve of the inhibition of VEIDase activity by Ac-VEID-CHO. The data are expressed as a percent of mean VEIDase activity at 0 μM Ac-VEID-CHO (±S.E.M., n=3). Each assay was performed under standard conditions (Materials and Methods) and repeated at least twice using different cell extracts
These results suggest that Norway spruce cells contain a single protease or a group of related proteases with the substrate preference and inhibitor specificity similar to those of mammalian caspase-6. Considering the large number of cell extracts analysed, it is also evident that there are no caspase-like proteases other than VEIDase (i.e., caspase-6-like) activated during embryogenesis in Norway spruce.
Biochemical characteristics of Norway spruce VEIDase activity
In the comparative study of biochemical characteristics of caspases-3, -6, -7 and -8, Stennicke and Salvesen28 observed high sensitivity of catalytic activity of purified mammalian caspase-6 towards subtle changes in pH, ionic strength and Zn2+ concentration. Therefore, we examined whether Norway spruce VEIDase has a similarly high sensitivity to the same factors.
To study the effect of pH, a sample of embryogenic culture was divided into parts, each part extracted and subsequently analysed for VEIDase activity at a fixed pH level using modified assay buffer containing 100 mM of either maleic acid (pH 2 or 2.5), formic acid (pH 3 or 3.5), acetic acid (pH 4–5) or MES (pH 5.5–6.5) instead of HEPES (pH 6.8–7.8). The double-bell-shaped VEIDase activity profile was obtained (Figure 3a). One, broader, bell was extended over a region of low pH (maximal activity at ≈pH 4.0), which were more acidic than normally found in plant vacuoles (Figure 3a). Another bell was similar to pH-dependence profile characteristic for caspase-6,28 occupying a very narrow pH region at around neutral pH, with the maximal activity at ≈pH 7.0 corresponding to acidified (by ≈0.3 pH units) cytoplasm of suspension cultured plant cells (Figure 3a).
Effects of pH (a), ionic strength (b), Zn2+ (c) and temperature (d) on Norway spruce VEIDase activity in vitro. (a) pH was adjusted both during preparation of cell extacts and during the assay. Bars show S.E.M. (n=3). The shaded areas illustrate the ranges of vacuolar and cytoplasmic pH found in suspension cultured plant cells.29 (b) Sodium chloride was added to reaction mixtures at the indicated concentrations just prior to addition of VEID-AMC. The data are expressed as a percent of mean VEIDase activity at 0 mM NaCl (±S.E.M., n=3). (c) Zinc chloride was added to reaction mixtures at the indicated concentrations just prior to addition of VEID-AMC. To avoid Zn2+ chelation, DTT in the assay buffer was replaced by equimolar amount of β-mercaptoethanol. The data are expressed as a percent of mean VEIDase activity at 0 μM Zn2+ (±S.E.M., n=3). (d) Different incubation temperatures were tested during the assay. The data are expressed as mean±S.E.M., n=3. *, P<0.01 versus 30°C. Each experiment was repeated at least twice using different cell extracts, and similar response profiles were obtained
To verify that the double-bell-shaped pH dependence was caused by catalytic activation/inactivation during proteolytic reaction, but not already during cell extraction, we monitored VEIDase activity in cell extracts prepared at pH 4 and 7 using acetic acid- and HEPES-containing buffers, respectively, and then adjusted to lower or higher pH in assay reactions by diluting with the appropriate buffers. The outcome of this experiment was identical to the pH-dependence profile shown in Figure 3a confirming that Norway spruce VEIDase catalysis is highly sensitive to pH.
Activity of mammalian caspase-6 was shown to be completely abolished with 1 M concentration of NaCl.28 In vitro VEIDase activity of Norway spruce seems to be even more sensitive to ionic strength, since it declined faster than caspase-6 activity as ionic strength increased, becoming hardly detectable at 100 mM (Figure 3b).
Zinc is known as a potent inhibitor of apoptosis; consistently, zinc chelation has an adverse effect.30,31,32 These effects of Zn2+ on apoptosis are believed to be due to the inactivation or activation of effector caspases.31,32 Among different mammalian caspases, caspase-6 is most readily inhibited by Zn2+, being completely inactivated in vitro by 100 μM.28 Since DTT chelates Zn2+, we analysed sensitivity of spruce VEIDase activity to this ion using assay buffer with DTT replaced by equimolar amount of β-mercaptoethanol. Two-fold inhibition of VEIDase was observed at 10 μM Zn2+ (Figure 3c). As in the case of caspase-6, 100 μM Zn2+ completely abrogated VEIDase in spruce cell extracts (Figure 3c).
Catalytic activity of most animal proteases, including caspases, has an optimum at 37°C. As Norway spruce is growing in moderate and subarctic zones, under temperatures substantially lower than 37°C, we were tempted to verify whether spruce VEIDase is tolerating increased temperatures and, at the same time, to determine the optimal temperature range of this activity. We found that spruce VEIDase was significantly inhibited in vitro by the temperatures beyond the range from 28 to 33°C (Figure 3d), reflecting ecological adaptation of plant caspase-like enzyme(s) to environmental conditions.
To exclude the possibility that there are spruce caspase-like proteases other than VEIDase, activated by the changes of pH, ionic strength, [Zn2+] or temperature, we assayed the cleavage of five other peptide substrates (Figure 1) in the same cell extracts under the same conditions as tested for VEID-AMC cleavage (Figure 3). All response profiles obtained for the cleavage of these peptides were similar in shape to the ones generated for VEIDase activity (Figure 3), with the same relative rates for proteolytic activities as observed under standard assay conditions (Figure 1; data not shown).
VEIDase activity is associated with the shaping of developing embryos
To begin to examine the role for the VEIDase in spruce embryogenesis, we first analysed kinetics of this activity over the whole developmental pathway starting with proembryogenic masses to fully developed mature embryos. The activity increased about three-fold, as proembryogenic masses developed to early embryos under depletion and subsequent withdrawal of plant growth regulators (PGR), with the maximal activity observed during early embryogeny at the time of active elongation of the embryo suspensor (at 4 days after withdrawal of PGR; Figure 4a). Thereafter, the activity began to decline, becoming barely detectable in mature embryos (after 35 days with abscisic acid (ABA); Figure 4a), at the developmental stage when embryo shaping is no longer required.23
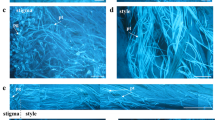
Developmental regulation of VEIDase activity during Norway spruce embryogenesis. (a) Kinetics of VEIDase activity during embryogenesis. Cell extracts were prepared at the indicated time points during embryogenesis: 3 and 7 days after addition of PGR (3d +PGR and 7d +PGR, respectively), 1, 4 and 7 days after withdrawal of PGR (1d −PGR, 4d −PGR and 7d −PGR, respectively) and 7 and 35 days after the onset of ABA treatment (7d ABA and 35d ABA, respectively). VEIDase activity was measured in vitro under standard assay conditions. On the top of each bar, a schematic drawing of the corresponding developmental stage is shown (not drawn to scale), including proliferating proembryogenic mass (3d+PGR), proembryogenic mass competent to form embryos (7d+PGR), the beginning of early embryogeny (1d −PGR), active growth of the embryo-suspensor (4d −PGR), embryo at the end of early embryogeny with the fully elongated suspensor (7d −PGR), embryo in the beginning of late embryogeny showing elimination of the suspensor (7d ABA) and mature embryo (35d ABA).21,23,25 Blue colour corresponds to vacuolated cells, which undergo programmed cell death and stain in vivo blue with Evan's blue. The meristematic living cells are shown in red, which stain in vivo red with acetocarmine.23 Note a 10–40 times lower VEIDase activity in the mature embryos (35d ABA) compared to the less-developed embryos exhibiting active shape remodeling. Data are expressed as mean±S.E.M., n=3. This experiment was repeated twice on separate occasions with similar results. (b) Embryo-specific pattern of in vivo localisation of VEIDase activity. Early embryos sampled 4 days after withdrawal of PGR were triple-stained: by DAPI, red fluorescent marker and caspase-6-specific fluorogenic substrate (see Materials and Methods). Blue fluorescence illustrates nuclear DAPI staining. Red fluorescence shows nonspecific cytoplasmic staining. Green fluorescence indicates VEIDase activity, which is shown by yellow colour on the merged image. Apical–basal embryonic pattern formation is illustrated on the merged image by double-side arrows delineating embryonal mass (EM), embryonal tubes (ET) and embryo-suspensor (ES). Note that VEIDase activity is restricted to the embryonal tubes and embryo-suspensor cells. Asterisks mark large vacuoles (unstained) in the embryo-suspensor. Scale bar=50 μm. In all, more than 100 embryos similar to the one shown in the figure were analysed, and all embryos exhibited the same staining pattern
To verify that the VEIDase activity is localised specifically in those embryonic cells that undergo programmed cell death, we stained living early embryos collected at 4 days after withdrawal of PGR (the time for the maximal VEIDase activity; Figure 4a) with the cell permeable fluorogenic substrate containing the -VEIDN- sequence (caspase-6 cleavage site in lamin A) for fluorescent microscopy analysis (see Materials and Methods). The spruce embryo at this stage exhibits a ‘gradient’ of cell death along its apical–basal axis, starting with living meristematic cells in the embryonal mass followed by embryonal tube cells that are committed to die and, finally, embryo-suspensor cells showing a greater extent of cell dismantling towards the basal part of the embryo.25 VEIDase activity was detected in the embryonal tube cells and in the first adjacent layer of suspensor cells, whereas neither embryonal mass cells nor cells in the distal layer of embryo suspensor exhibited VEID cleavage in vivo (Figure 4b). Together, these results demonstrated that increase in the VEIDase activity was associated with the shaping of the spruce embryo at the early stages of embryogenesis, and localised specifically in the cells during both the commitment (embryonal tube cells) and in the beginning of the execution phase of programmed cell death (proximal embryo-suspensor cells).
Inhibition of VEIDase activity disrupts embryonic pattern formation
Developmental regulation of VEIDase activity during spruce embryogenesis suggested that this activity might be critical for correct embryonic pattern formation. Therefore, we next examined what the consequences of inhibition of this activity would be for embryonic pattern formation and associated programmed cell death. In these experiments, we used the pan-caspase inhibitor, zVAD-fluoromethyl ketone (fmk), and caspase-6-specific inhibitor, zVEID-fmk, whose effects on embryogenesis were compared with both a control (DMSO) and zLEHD-fmk (caspase-9-specific inhibitor).
The level of terminal deoxynucleotidyl transferase (TdT)-mediated dUTP nick end labelling (TUNEL) in the whole culture was significantly (P<0.01) reduced by zVAD-fmk and zVEID-fmk at the concentrations 2 μM and 200 nM, respectively, whereas the treatment with zLEHD-fmk at much higher concentration (20 μM) had no effect (Figure 5a). The frequency of embryos in zVAD-fmk- and zVEID-fmk-treated cultures was reduced too (Figure 5b), strongly supporting our previous finding of the vital role for programmed cell death in plant embryo development.23,25,33
Developmental effects of the inhibition of VEIDase activity. Embryogenic cell culture was treated with cell-permeable caspase inhibitors zVAD-fmk, zVEID-fmk and zLEHD-fmk, simultaneously with withdrawal of PGR. (a) At 4 days after the treatments, TUNEL was assessed. Results indicate percent of average frequency of TUNEL-positive nuclei in the control (DMSO-treated) culture, and expressed as mean (±S.E.M.) of five samples, with at least 200 nuclei observed for each sample. *, P<0.01 versus control. This experiment was repeated twice on separate occasions with similar results. (b) At 7 days after the treatment, the frequency of embryos in the culture was analysed. The data are expressed as a percent of mean embryo frequency in the control (DMSO-treated) culture (±S.E.M. of three independent experiments, with at least 500 cell aggregates observed per experiment). *, P<0.01 versus control. (c) Typical developmental aberrations observed 7 days after the treatment with either zVAD-fmk or zVEID-fmk at 2 μM and 20 μM (shown for zVEID-fmk only). The embryos were stained with DAPI and TUNEL. Both concentrations of inhibitors affect individual embryo development, as the embryonal masses (EM) are aggregated in clumps. The embryo-suspensor (ES) is prevented from normally elongating, and its development is completely blocked upon treatments with 2 and 20 μM of inhibitor, respectively. DNA fragmentation in the distal part of the suspensor in the control embryo is illustrated by a group of TUNEL-positive nuclei (red fluorescence inside the box). Note a lack of TUNEL in the abnormal embryos developed upon inhibitor treatments. Scale bars=50 μm
Pattern formation of spruce embryos treated with either zVAD-fmk or zVEID-fmk deviated from the normal morphotype, with the severity of aberrations correlating well with the concentration of inhibitors. First developmental aberrations were reproducibly observed when inhibitors were applied at 2 μM, the response being that the embryonal masses were unable to separate from each other and proliferated in clumps possessing short suspensors with no signs of cell death as revealed by in vivo VEIDase staining (data not shown) and TUNEL (Figure 5c). At higher concentration of inhibitors (20 μM), the differentiation of embryo-suspensor was blocked (Figure 5c). We conclude that the VEIDase activity is an integral part of the developmental cell death program, and that this activity is necessary for embryo pattern formation.
Discussion
We report that Norway spruce embryo development implicates activation of a protease or a group of proteases with the preferential cleavage of VEID sequence-containing substrate. VEID amino-acid sequence corresponds to the site of lamin A cleaved by mammalian caspase-6 during apoptosis.34,35 The pattern of relative cleavage rate of different peptides by spruce cell extracts in vitro is highly stable (Figure 1) and similar to the one specific for mammalian caspase-6, which shows favorable interaction with peptides containing β-branched amino acids in P4 position.26 Therefore, we further conclude that there is no other caspase-like protease activated during spruce embryo pattern formation.
Apart from similar substrate specificity, both spruce VEIDase and caspase-6 can be inhibited by Ac-VEID-CHO (Figure 2) and are highly sensitive to changes in pH, ionic strength and the presence of Zn2+ (Figure 3).28 Noteworthy, in vitro activation of spruce VEIDase at a pH substantially lower than the vacuolar pH of plant cells (Figure 3a) may be indicative of the involvement of this proteolytic activity in the degradation of cytoplasm inside Golgi- and plastid-derived acidic vesicles, the earliest event of the execution phase of autophagic cell death in embryo-suspensor.23,25,36,37,38
This is the first study of caspase-like activity in a gymnosperm plant. Neither caspase-1- nor 3-like-activities previously reported for angiosperm cell death systems8,9,11,13 were found to be involved in spruce embryonic cell death. It is possible that the observed differences in substrate specificity between angiosperm and gymnosperm caspase-like proteases are attributed to a long phylogenetic distance between the two groups of higher plants. Being phylogenetically older group, gymnosperms might have evolved only a single, central cell-death caspase-like protease, as in the case of Caenorhabditis elegans39 and yeast.7
Proper regulation of caspases is crucial in animal development. Knockout of caspases-3 and -9 in mice results in prenatal or perinatal lethality, mainly due to the failure of apoptosis during brain development.16,17 Likewise, mice with knockout of caspase-8 die during embryogenesis with abnormalities in heart development.18 Drosophila flies with mutations in the caspase dcp-1 die as larvae owing to tumorogenesis.40 The latter study, together with the observation that caspase inhibitor p35 prevents autophagy in Drosophila salivary glands,41 suggests that caspases can also be critically involved in the regulation of autophagic cell death.5 In our present work, VEIDase activity was increased at the early stages of Norway spruce embryo development and then abrogated, as embryo shaping was accomplished (Figure 4a). Consistent with this, the results of in vivo inhibitor experiments (Figure 5) further support the idea that VEIDase activity is crucial in autophagic cell death during plant embryo pattern formation.
Although the definite place of VEIDase in Norway spruce autophagic cell death pathway remains to be elucidated, in vivo studies suggest that it is already activated in the embryonal tube cells, that is, during commitment to cell death (Figure 4b).25 The localisation of VEIDase activity in the uppermost cells of embryo-suspensor (Figure 4b) indicates an additional role for this activity at the beginning of the execution phase of cell death when disruption of microtubule network and slow engulfment of the cytoplasm by growing and fusing vacuoles are the major events.23,25 Consistent with this, the embryos treated with VEIDase inhibitors exhibited a block in suspensor organ formation (Figure 5c).
Any direct comparison of the roles for mammalian caspase-6 and plant VEIDase activity in programmed cell death is difficult, as very little is currently known about caspase-6 in apoptosis. In fact, the only substrate presently thought to be cleaved exclusively by caspase-6 is lamin A/C.35,42 Cleavage of lamin A by caspase-6 was shown to be critical for chromatin condensation and nuclear breakdown.43 Interestingly, although plants do not have homologs of animal lamins,44 and programmed cell death in spruce embryos does not involve strong chromatin condensation,23 we observed decreased frequency of DNA fragmentation in zVAD-fmk- and zVEID-fmk-treated cultures (Figure 5a). Apart from the role of caspase-6 as one of the downstream caspases, a recent study of Cowling and Downward45 revealed an additional, upstream role of caspase-6 in direct activation of initiator procaspase-8 in Fas-triggered apoptosis.
Our present efforts are directed towards isolating the Norway spruce protein responsible for VEIDase activity during developmental cell death. Interestingly, overexpression of yeast metacasaspase gene YCA1 revealed a pattern of caspase-like activity strikingly similar to the one shown in the present study (Figure 1), with the clear preference of the cleavage of VEID-containing substrate.7 Cloning and characterisation of the Norway spruce metacaspase(s) will unambiguously show whether VEIDase activity is caused by a metacaspase.
Materials and Methods
Cell culture and treatments
Embryogenic cell line 95.88.22 of Norway spruce (Picea abies L. Karst.) was used in this study. The entire embryo development pathway was regulated as described,23,25,33 including repeated addition of PGR, auxin and cytokinin, to maintain proliferation of proembryogenic masses, followed by withdrawal of these PGR to stimulate early embryogeny and, finally, ABA treatment to promote late embryogeny and embryo maturation. In brief, the cryopreserved cell line was thawed from liquid nitrogen and established as a suspension culture in liquid PGR-containing medium. Cell density was adjusted to 6% (vol/vol) at every weekly subculture. At 7 days after the last addition of PGR, cell aggregates were first washed in PGR-free medium and then subcultured into fresh PGR-free medium, with the cell density adjusted to 6%. At 7 days after withdrawal of PGR, early embryos were subjected to ABA treatment. Mature embryos developed within 5 weeks after the start of ABA treatment.
In in vivo caspase inhibitor experiments, PGR-free medium was supplemented with cell-permeable caspase inhibitors (in DMSO) at 200 nM, 2 μM and 20 μM, which were added just prior to inoculation of cell aggregates. In all the treatments and control, the concentration of DMSO in the medium was adjusted to 0.2% (vol/vol). Pan-caspase inhibitor zVAD-fmk and caspases-6- and -9-specific inhibitors zVEID-fmk and zLEHD-fmk, respectively, were tested (all from Enzyme Systems Products, Dublin, CA, USA). Each treatment was carried out with two Erlenmeyer flasks and repeated twice. The frequency of embryos in the culture was determined after 7 days through observation of at least 500 individual cell aggregates for each treatment.23
In vitro caspase substrate cleavage assay
Cell extracts were prepared at successive stages of embryogenesis. The samples were ground in liquid nitrogen, and the cell powder was resuspended in assay buffer (100 mM HEPES/10% sucrose/0.1% CHAPS/5 mM DTT/10−6% Nonidet P-40, pH 7.0, if not otherwise stated) at 4°C. Cell debris was pelleted by centrifugation at 16 000 × g for 10 min at 4°C and the supernatant (cell extract) was collected and stored at −70°C. Protein concentration in the defrosted extracts was estimated by Bradford assay (BioRad, Hercules, CA, USA). Proteolytic activity was measured in 150-μl reaction mixtures containing 22.5 μg protein and 50 μM of individual AMC-conjugated peptide (Peptide Institute, Inc., Osaka, Japan), specific for individual mammalian caspase, in the assay buffer. Triplicate reactions were incubated for 1 h at 30°C (if not otherwise stated), and fluorescence readings of ice-cooled samples were taken against a blank containing assay buffer and peptide alone (i.e., no protein) using an excitation wavelength of 360 nm and emission wavelengths of 460 nm (Versa-Fluor Fluorometer, BioRad, Hercules, CA, USA). Kinetics of substrate hydrolysis was tested to be linear throughout the 1-h reactions. Standards containing 0–600 pmol of AMC (in the assay buffer) were utilised to determine the amount of fluorochrome released. Proteolytic activity was expressed in pmol AMC liberated per minute.
In the experiments with individual protease inhibitors, the latter were added to reaction mixtures just prior to addition of VEID-AMC. Most of the inhibitors were obtained from Roche (Mannheim, Germany), except for lactacystine, Ac-VEID-CHO and Ac-LEHD-CHO (all from Peptide Institute, Inc., Osaka, Japan). Dimethylsulphoxide concentration was less than 1% throughout all assays.
In vivo staining of VEIDase activity
Specific for caspase-6, VEIDase activity can be detected in living cells by using the fluorogenic cell-permeable peptidic substrate containing -VEIDN- sequence (caspase-6 cleavage site in lamin A) and included in CyToxiLux kit (OncoImmunin, Inc.,. Gaithersburg, MD, USA). This substrate is composed of two fluorophores covalently linked to caspase-6-specific peptide. In the uncleaved substrate, fluorescence is quenched owing to the formation of intramolecular excitonic dimers. Upon cleavage of the peptide, the fluorophore–fluorophore interaction is abolished, leading to an increase in fluorescence.46,47 In the present study, we adjusted CyToxiLux protocol for use on Norway spruce embryos.
At 4 days after withdrawal of PGR, 500-μl samples of densely packed early embryos were collected and stained in 2 ml of fresh PGR-free culture medium containing 1 μl of red fluorescent marker (from the kit) for 1 h at 25°C in the dark. Stained embryos were washed for 30 min in three 2-ml aliquots of culture medium and then incubated with 500 μl of caspase substrate solution (from the kit) for 1 h at 25°C (optimal temperature) in the dark. After subsequent 45-min washing with three 2-ml aliquots of wash buffer (from the kit), the embryo samples were stained with 1 μg/ml 4′-6-diamino-2-phenylindole (DAPI) (Roche, Mannheim, Germany) in the wash buffer and the same microscopic fields were examined with a Microphot-FXA fluorescence microscope (Nikon, Tokyo, Japan) using distinct cubes for DAPI (excitation filter, 340–380 nm; dichroic mirror, 400 nm; barrier filter, 435–485 nm), red fluorescent marker (excitation filter, 541–551 nm; dichroic mirror, 580 nm; barrier filter, 590 nm) and caspase substrate (excitation filter, 490 nm; dichroic mirror, 505 nm; barrier filter, 525 nm). Images were captured with a Fujix digital camera HC-300Zi (Nikon, Tokyo, Japan). Specificity of VEIDase activity staining was confirmed by changing incubation time with caspase substrate solution. Staining intensity gradually increased over the first hour of incubation, whereas prolonged (>90 min) incubations were found to be toxic for the Norway spruce embryos leading to collapse of the cells in both embryonal mass and suspensor.
In situ detection of DNA fragmentation
Terminal deoxynucleotidyl transferase (TdT)-mediated dUTP nick end labelling and DAPI counterstaining of whole mount proembryogenic masses and embryos were performed as described,23 the only difference being that a TMR red in situ cell death detection kit (Roche, Mannheim, Germany) was used in this study. Application of TMR red-labeled-dUTP without TdT was used as the negative control. To assess the frequency of nuclei with fragmented DNA in control and caspase inhibitor-treated cultures, at least 1000 nuclei were observed in DAPI/TUNEL-stained samples for each treatment.
Statistics
Numerical data are shown as mean±S.E.M. of not less than three individual experiments. Pearson correlation coefficients (r) were computed for mean proteolytic activities of Norway spruce cell extracts against different fluorogenic peptides, pairwise for each combination of two peptides. Comparisons between means were made with Student's t test (P<0.01). Statistical analyses were performed by using JMP 3.1 software (SAS Institute Inc.).
Abbreviations
- ABA:
-
abscisic acid
- AMC:
-
7-amino-4-methylcoumarin
- DAPI:
-
4′-6-diamino-2-phenylindole
- fmk:
-
fluoromethyl ketone
- PGR:
-
plant growth regulators
- PMSF:
-
phenylmethylsulphonyl fluoride
- TLCK:
-
L-1-chloro-3-(4-tosylamido)-7-amino-2-heptanone
- TUNEL:
-
terminal deoxynucleotidyl transferase (TdT)-mediated dUTP nick end labelling
References
Beers EP and McDowell JM (2001) Regulation and execution of programmed cell death in response to pathogens, stress and developmental cues. Curr. Opin. Plant Biol. 4: 561–567
Ameisen JC (2002) On the origin, evolution and nature of programmed cell death: a timeline of four billion years. Cell Death Differ. 9: 367–393
Koonin EV and Aravind L (2002) Origin and evolution of eukaryotic apoptosis: the bacterial connection. Cell Death Differ. 9: 394–404
Zhivotovsky B (2002) From the nematode and mammals back to the pine tree: on the diversity and evolution of programmed cell death. Cell Death Differ. 9: 867–869
Baehrecke EH (2002) How death shapes life during development. Nat. Rev. Mol. Cell Biol. 3: 779–787
Uren AG, O'Rourke K, Aravind L, Pisabarro MT, Seshagiri S, Koonin EV and Dixit VM (2000) Identification of paracaspases and metacaspases: two ancient families of caspase-like proteins, one of which plays a key role in MALT lymphoma. Mol. Cell 6: 961–967
Madeo F, Herker E, Maldener C, Wissing S, Lächelt S, Herlan M, Fehr M, Lauber K, Sigrist SJ, Wesselborg S and Fröhlich K-U (2002) A caspase-related protease regulates apoptosis in yeast. Mol. Cell 9: 911–917
Korthout HAAJ, Berecki G, Bruin W, van Duijn B and Wang M (2000) The presence and subcellular localization of caspase 3-like proteinases in plant cells. FEBS Lett. 475: 139–144
Del Pozo O and Lam E (1998) Caspases and programmed cell death in the hypersensitive response of plants to pathogens. Curr. Biol. 8: 1129–1132
D'Sylva I, Poirier GG and Heath MC (1998) Activation of cysteine proteases in cowpea plants during the hypersensitive response – a form of programmed cell death. Exp. Cell Res. 245: 389–399
Clarke A, Desikan R, Hurst RD, Hancock JT and Neill SJ (2000) NO way back: nitric oxide and programmed cell death in Arabidopsis thaliana suspension cultures. Plant J. 24: 667–677
Elbaz M, Avni A and Weil M (2002) Constitutive caspase-like machinery executes programmed cell death in plant cells. Cell Death Differ. 9: 726–733
Woltering EJ, van der Bent A and Hoeberichts A (2002) Do plant caspases exist? Plant Physiol. 130: 1764–1769
Hansen G (2000) Evidence for Agrobacterium-induced apoptosis in maize cells. Mol. Plant-Microbe Interact. 6: 649–657
Lincoln JE, Richael C, Overduin B, Smith K, Bostock R and Gilchrist DG (2002) Expression of antiapoptotic baculovirus p35 gene in tomati blocks programmed cell death and provides broad-spectrum resistance to desease. Proc. Natl. Acad. Sci. USA 99: 15217–15221
Kuida K, Zheng TS, Na SQ, Kuan CY, Yang D, Karasuyama H, Rakic P and Flavell RA (1996) Decreased apoptosis in the brain and premature lethality in CPP32-deficient mice. Nature 384: 368–372
Kuida K, Haydar TF, Kuan CY, Gu Y, Taya C, Karasuyama H, Su MSS, Rakic P and Flavell RA (1998) Reduced apoptosis and cytochrome c-mediated caspase activation in mice lacking caspase 9. Cell 94: 325–337
Varfolomeev EE, Schuchmann M, Luria V, Chiannilkulchai N, Beckmann JS, Mett IL, Rebrikov D, Brodianski VM, Kemper OC, Kollet O, Lapidot T, Soffer D, Sobe T, Avraham KB, Goncharov T, Holtmann H, Lonai P and Wallach D (1998) Targeted disruption of the mouse Caspase 8 gene ablates cell death induction by the TNF receptors, Fas/Apo1, and DR3 and is lethal prenatally. Immunity 9: 267–276
Zheng TS and Flavell RA (2000) Divinations and surprises: genetic analysis of caspase function in mice. Exp. Cell Res. 256: 67–73
Von Arnold S, Sabala I, Bozhkov P, Dyachok J and Filonova L (2002) Developmental pathways of somatic embryogenesis. Plant Cell Tiss Org. Cult. 69: 233–249
Filonova LH, Bozhkov PV and von Arnold S (2000) Developmental pathway of somatic embryogenesis in Picea abies as revealed by time-lapse tracking. J. Exp. Bot. 51: 249–264
Singh H (1978) Embryology of Gymnosperms. (Borntrager, Berlin)
Filonova LH, Bozhkov PV, Brukhin VB, Daniel G, Zhivotovsky B and von Arnold S (2000) Two waves of programmed cell death occur during formation and development of somatic embryos in the gymnosperm, Norway spruce. J. Cell Sci. 113: 4399–4411
Filonova LH, von Arnold S, Daniel G and Bozhkov PV (2002) Programmed cell death eliminates all but one embryo in a polyembryonic plant seed. Cell Death Differ. 9: 1057–1062
Smertenko AP, Bozhkov PV, Filonova LH, von Arnold S and Hussey PJ (2003) Re-organisation of cytoskeleton during developmental programmed cell death in Picea abies embryos. Plant J. 33: 813–824
Talanian VT, Quinlan C, Trautz S, Hacket MC, Mankovich JA, Banach D, Ghayur T, Brady KD and Wong WW (1997) Substrate specificities of caspase family proteases. J. Biol. Chem. 27: 9677–9682
Lockshin RA and Zakeri Z (2002) Caspase-independent cell deaths. Curr. Opin. Cell Biol. 14: 727–733
Stennicke HR and Salvesen GS (1997) Biochemical characteristics of caspases-3, -6, -7, and -8. J. Biol. Chem. 272: 25719–25723
Kurkdjian A and Guern J (1989) Intracellular pH: measurement and importance in cell activity. Annu. Rev. Plant Physiol. Plant Mol. Biol. 40: 271–303
Sunderman Jr FW (1995) The influence of zinc on apoptosis. Ann. Clin. Lab. Sci. 25: 134–142
Jiang S, Zhivotovsky B, Burgess DH, Gahm A, Chow SC and Orrenius S (1997) The role of proteolysis in T cell apoptosis triggered by chelation of intracellular Zn2+. Cell Death Differ. 4: 39–50
Chimienty F, Seve M, Richard S, Mathieu J and Favier A (2001) Role of cellular zinc in programmed cell death: temporal relationship between zinc depletion, activation of caspases, and cleavage of Sp family transcription factors. Biochem. Pharmacol. 62: 51–62
Bozhkov PV, Filonova LH and von Arnold S (2002) A key developmental switch during Norway spruce somatic embryogenesis is induced by withdrawal of growth regulators and is associated with cell death and extracellular acidification. Biotechnol. Bioeng. 77: 658–667
Orth K, Chinnaiyan AM, Garg M, Froelich CJ and Dixit VM (1996) The CED-3/ICE-like protease Mch2 is activated during apoptosis and cleaves the death substrate lamin A. J. Biol. Chem. 271: 16443–16446
Takahashi A, Alnemri ES, Lazebnik YA, Fernandes-Alnemri T, Litwack G, Moir RD, Goldmen RD, Poirier GG, Kaufmann SH and Earnshaw WC (1996) Cleavage of lamin A by Mch2α but not CPP32: multiple ICE-related proteases with distinct substrate recognition properties are active in apoptosis. Proc. Natl. Acad. Sci. USA 93: 8395–8400
Nagl W (1976) Ultrastructural and developmental aspects of autolysis in embryo-suspensors. Ber. Deutsch. Bot. Ges. Bd. 89: 301–311
Nagl W (1977) Plastolysomes – plastids involved in autolysis of the embryo-suspensor in Phaseolus. Z. Pflanzenphysiol. 85: 45–51
Gärtner P-J and Nagl W (1980) Acid phosphatase activity in plastids (plastolysomes) of senescing embryo-suspensor cells. Planta 149: 341–349
Ellis RE and Horvitz RH (1986) Genetic control of programmed cell death in the nematode, C. elegans. Cell 44: 817–829
Song ZW, McCall K and Steller H (1997) DCP-1, a Drosophila cell death protease essential for development. Science 275: 536–540
Lee CY and Baehrecke EH (2001) Steroid regulation of autophagic programmed cell death during development. Development 128: 1443–1455
Slee EA, Adrain C and Martin SJ (2001) Executioner caspase-3, -6 and –7 perform distinct, non-redundant roles during the demolition phase of apoptosis. J. Biol. Chem. 276: 7320–7326
Ruchaud S, Korfali N, Villa P, Kottke TJ, Dingwall C, Kaufmann SH and Earnshaw WC (2002) Caspase-6 gene disruption reveals a requirement for lamin A cleavage in apoptotic chromatin condensation. EMBO J. 21: 1967–1977
Meier I (2001) The plant nuclear envelope. Cell Mol. Life Sci. 58: 1774–1780
Cowling V and Downward J (2002) Caspase-6 is the direct activator of caspase-8 in the cytochrome c-induced apoptosis pathway: absolute requirement for removal of caspase-6 prodomain. Cell Death Differ. 9: 1046–1056
Komoriya A, Packard BZ, Brown MJ, Wu M-L and Henkart PA (2000) Assessment of caspase activities in intact apoptotic thymocytes using cell-permeable fluorogenic caspase substrates. J. Exp. Med. 191: 1819–1828
Liu L, Chahroudi A, Silvestri G, Wernett ME, Kaiser WJ, Safrit JT, Komoriya A, Altman JD, Packard BZ and Feinberg MB (2002) Visualization and quantification of T cell-mediated cytotoxicity using cell-permeable fluorogenic caspase substrates. Nat. Med. 8: 185–189
Acknowledgements
We thank David Clapham and Beverly Packard for helpful discussion and correspondence. This work was supported by grants from the Carl Tryggers Foundation (to PVB), the Swedish Foundation for International Cooperation in Research and Higher Education (to PVB and APS), the Spanish Ministry of Education (to MFS), the Swedish National Research Council (to BZ), the Swedish Cancer Society (to BZ) the European Commission FP5 (to BZ), and the Forest Tree Breeding Association in Sweden (to SVA).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by B. Osborne
Rights and permissions
About this article
Cite this article
Bozhkov, P., Filonova, L., Suarez, M. et al. VEIDase is a principal caspase-like activity involved in plant programmed cell death and essential for embryonic pattern formation. Cell Death Differ 11, 175–182 (2004). https://doi.org/10.1038/sj.cdd.4401330
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401330
Keywords
This article is cited by
-
Ac-DEVD-CHO (caspase-3/DEVDase inhibitor) suppresses self-incompatibility–induced programmed cell death in the pollen tubes of petunia (Petunia hybrida E. Vilm.)
Cell Death Discovery (2024)
-
Genetics Behind Sexual Incompatibility in Plants: How Much We Know and What More to Uncover?
Journal of Plant Growth Regulation (2023)
-
RNA-Seq analysis reveals potential regulators of programmed cell death and leaf remodelling in lace plant (Aponogeton madagascariensis)
BMC Plant Biology (2021)
-
Caspase-like proteases and the phytohormone cytokinin as determinants of S-RNAse–based self-incompatibility–induced PCD in Petunia hybrida L.
Protoplasma (2021)
-
Cylindrospermopsin induces biochemical changes leading to programmed cell death in plants
Apoptosis (2017)