Abstract
Most forms of apoptosis involve activation of caspases and it is likely that differences between cells in their ability to activate caspases contributes to the responsiveness of any given cell within a population to apoptotic stimuli. To study the molecular mechanisms that underlie such differences, it is necessary to measure caspase activity in individual cells. Here, we describe a method that allows the continuous monitoring of caspase activity in individual, living mammalian cells. This approach allows studies of the kinetics of caspase activation to be performed in individual cells within a population. We demonstrate that in a group of cells where some cells die and some cells survive in response to the same stimulus, the cells that die can be differentiated from those that survive based on the amount of caspase activity in each cell several hours before death occurs.
Similar content being viewed by others
Introduction
Most forms of apoptosis involve caspases, the proteases that are activated by apoptotic signals to digest cellular proteins and induce death.1,2 Caspases are cysteine-dependent proteases that recognize specific amino acid motifs containing aspartic acid residues in their target proteins. More than a dozen caspases have been identified in mammals. Caspases can be placed into three broad categories: those such as caspase 1 that are primarily involved in inflammatory responses; and two groups of caspases that are involved in apoptosis – initiator caspases such as caspase 8 and effector caspases such as caspase 3. Because defective apoptosis (both too much apoptosis and too little apoptosis) is critical in many diseases, there is considerable interest in understanding caspase regulation in more detail with the ultimate goal of manipulating caspase activity as a treatment for disease. Biochemical assays for caspase activity require relatively large numbers of cells that are harvested and lysed for analysis. Such analysis provides a measurement of the total caspase activity of a cell population but does not permit analysis of the variation in responses between individual cells within a population. Such differences in responsiveness may underlie differences in cell behavior. For example, a cell that harbors an oncogenic mutation might be less able to activate caspases than are its normal neighbors. To study the mechanisms that underlie such differences, it is useful to examine individual cells. Numerous methods exist to monitor apoptosis at the single cell level. These methods do not usually measure caspase activity directly and instead detect downstream consequences of caspase activity such as fragmented DNA, altered cell morphology, translocation of phosphatidylserine to the outer membrane or the appearance of a caspase-cleavage dependent epitope. Many of these methods require cell fixation and do not permit continuous monitoring of the apoptotic process over time thus precluding kinetic studies in individual cells. To further our understanding of caspase regulation, it would be useful to have a method that permits the continuous measurement of caspase activity in individual, living cells. Here we describe a method, based on fluorescence resonance energy transfer (FRET) between derivatives of green fluorescent protein (GFP) that allows caspase activity in individual cells to be measured over time. Using this method, we demonstrate that kinetic differences in caspase activity between cells in a population is correlated with whether the cells live or die within a particular time frame. These differences in caspase activity are apparent several hours before death occurs.
Results
FRET occurs when two fluorophores with overlapping emission and excitation spectra, such as GFP and its derivatives, are physically close to each other.3,4 The suitability of a GFP fusion protein for FRET-based measurements of a biological activity is determined by the spectral characteristics of the fluorophores and the sequence, length and structure of the linker peptide. For a caspase assay, the linker should be accessible to the enzyme while maintaining a structure that maximizes energy transfer in the intact fusion protein. We constructed and empirically tested several fusion proteins consisting of enhanced yellow fluorescent protein (YFP) fused via linker peptides that contained consensus caspase 3 cleavage sites (DEVD) to enhanced blue fluorescent protein (BFP). One of the intact fusion proteins emitted primarily yellow rather than blue fluorescence when excited with UV radiation at 380 nm indicating that FRET was occurring between the two fluorophores. Upon in vitro digestion of the purified recombinant protein with caspase 3, the ratio of yellow/blue fluorescence changed in a caspase activity-dependent manner (Figure 1a,b). The maximum change in fluorescence ratio that we detected upon caspase cleavage in vitro was approximately 14-fold. This change compares favorably with previous studies using trypsin cleavage of similar fusion proteins where 4–5-fold ratio changes were measured.3,5 We also tested a fusion protein where BFP was replaced with a cyan mutant, which resulted in a threefold change in fluorescence ratio after complete digestion. Similarly switching YFP for the green mutant resulted in only a 3.2 maximum change. Because the YFP-BFP combination led to the maximal ratio change in vitro, future experiments were performed with this combination. Caspase 3 did not cause fluorescence changes of a different fusion protein that contained a non-cleavable linker derived from the GluR3 glutamate receptor (Figure 1c). These data indicate that the caspase-dependent fluorescence changes required the presence of a caspase cleavage site. Caspases can be separated into three distinct groups that differ in their substrate specificity. When we tested whether the caspase 3-cleavable protein was affected by caspase 8, an initiator caspase, we found that incubation with caspase 8 caused only a minor change in fluorescence ratio. In this experiment, the emission ratios at 525 nm (i.e. yellow) and 445 nm (i.e. blue) were 3.24 for protein that was incubated in the absence of caspase, 0.42 for protein incubated in the presence of caspase 3 and 2.87 for protein incubated with caspase 8. Thus, our probe is a selective sensor for the activity of some but not all caspases.
FRET-based assay for caspase activity. (a) Recombinant YFP-C-BFP fusion protein was incubated at 37°C for various times with purified caspase 3, controls included samples treated with caspase plus an inhibitor (z-VADfmk) or maintained for 90 min without caspase. Digestion was stopped by adding an excess of caspase inhibitor and the fluorescence emission spectra for each sample was analyzed in a photon counting fluorimeter with the excitation set at 380 nm. The intact protein fluoresces yellow and this fluorescence decreases with concomitant increase in blue fluorescence in a caspase-dependent manner. (b) Shows the corresponding Western blot indicating that the protein was indeed cleaved by caspase 3. (c) Shows fluorescence spectra with or without caspase 3 for a fusion protein that undergoes efficient FRET but contains a non-cleavable linker. (d) Shows fluorescence spectra for the YFP-C-BFP protein in the presence of equivalent amounts of caspase 3 or caspase 8, only caspase 3 causes significant changes in the fluorescence ratio
To determine whether the molecule was cleaved in apoptotic cells, we constructed a SHEP neuroblastoma cell line that expressed the protein. We induced apoptosis by treating the cells with 300 mM mannitol to produce hyperosmolar stress, which causes caspase activation and apoptosis in these cells.6 FACS was used to measure fluorescence (Figure 2a). The cells were excited with the UV portion of an argon laser (approximately 350 nm) and the amount of yellow (i.e. FRET) and blue (i.e. conventional) fluorescence was measured before or after overnight treatment with 300 mM mannitol. The control cells showed high yellow but low blue fluorescence while mannitol-treated cells were more blue but less yellow. Thus, the cell population switches from yellow to blue upon induction of apoptosis. When blue and yellow cells were sorted and analyzed by Western blotting, we found that only the blue cells contained cleaved fusion protein (Figure 2b). These experiments show that the fusion protein can be used to identify and sort populations of living cells that contain active caspases based on their fluorescence characteristics.
FRET measurements in living cells. (a) SHEP cells expressing the YFP-C-BFP were analyzed by FACS with excitation from the UV portion of an argon laser. Untreated control cells (left panel) are substantially yellow when excited by UV light, while cells treated with 300 mM mannitol (right panel) show a decrease in yellow fluorescence and an increase in blue fluorescence. (b) Stable YFP-C-BFP expressing SHEP cells were treated with 300 mM mannitol and sorted based on their fluorescence characteristics into ‘blue’ and ‘yellow’ groups. Whole-cell lysates of sorted cells were separated on a gel and Western blotted using an anti-GFP antibody. Lanes 1–3 show the lysates of 40 000, 80 000, and 160 000 cells, gated for low yellow and greater blue fluorescence (i.e. ‘blue’ cells). Lane 4 shows the lysate of 36 000 cells gated for high yellow and less blue fluorescence (‘yellow’ cells), indicating that only the blue cells contain cleaved fusion protein
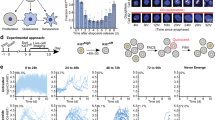
To determine whether this molecule would allow us to detect changes in caspase activity in individual cells, we performed time-lapse fluorescence microscopy on Hela cells transiently expressing the fusion protein after microinjection of an expression construct. Groups of cells on the same dish were injected with an expression construct encoding the caspase-cleavable YFP-C-BFP protein or a non-cleavable point mutant that replaced the aspartic acid at the cleavage site with an alanine. By injecting the two constructs into different groups of cells on the same dish, we compared the fluorescence changes in the same population of cells treated with the same apoptotic stimulus (staurosporin). The cells were incubated in an environmental chamber on the microscope and phase and fluorescence images were captured for each field at 30 min intervals throughout the experiment. Caspase activity was determined by quantifying fluorescence images after excitation at 380 nm and emission centered at both 545 nm (yellow fluorescence, i.e. FRET) and 450 nm (blue fluorescence, i.e. conventional excitation) were captured. To obtain an image of the yellow/blue ratio, a pixel by pixel ratio of the yellow and blue images was calculated. Figure 3a shows examples of a cell expressing the cleavable protein and a cell expressing the non-cleavable point mutant. The yellow/blue ratio decreased over time for the cleavable protein as indicated by the color change from primarily yellow and red to primarily blue and black in the images at different times. There was no reduction in ratio in cells that expressed the non-cleavable mutant. For quantitative measurement, the mean yellow fluorescence and blue fluorescence was calculated for individual cells at each time point. The relative change in fluorescence ratio was then determined by calculating the change in ratio for each cell between the start of the experiment and the 311 min time point. Figure 3b shows that the cleavable construct displayed a reduction in fluorescence ratio (blue lines) while there was no significant change for the non-cleavable construct (green lines). The mean negative slopes were calculated for 13 cells injected with the cleavable construct and 11 cells injected with the non-cleavable mutant (insert in Figure 3b). A t-test determined that these slopes were different with a highly significant P value (P<0.00001). Thus, we can follow caspase activity in individual cells by fluorescence microscopy and study differences between individual cells within a population in their caspase activities.
FRET measurement of caspase activity in individual cells. Hela cells were injected with an expression construct encoding YFP-C-BFP or a non-cleavable point mutant and treated with staurosporin to induce apoptosis. (a) Shows pseudocolored images of the pixel-by-pixel ratio of yellow/blue fluorescence of the same fields at time zero and 5 h 11 min after treatment. Phase and greyscale images were obtained for blue and yellow fluorescence after excitation at 380 nm, ratioed and expressed as RGB images, white and red indicate higher yellow/blue ratio (intact fusion protein), blue and black indicate lower yellow/blue ratio (cleaved fusion protein). The yellow/blue ratio decreases for the YFP-C-BFP protein (top panel) but not for the mutant (bottom panel). (b) Shows quantitative ratio measurements for individual cells expressing the two proteins at time zero and 5 h into the experiment. The ratio was reduced for all of the YFP-C-BFP expressing cells (blue lines) but not for those cells expressing the non-cleavable mutant (green lines). The insert shows the mean negative slopes (±S.E.M.) for both groups. Only the cleavable protein results in a change in fluorescence ratio
Apoptotic stimuli often displays variability in effectiveness between different cells; some cells are more resistant to a stimulus than others and this is reflected in asynchronous and variable amounts of death. This is true even when genetically identical cells are examined and it is common to find that a population of cells will contain some cells that die and some cells that survive an apoptotic stimulus. Two simple, non-exclusive, hypotheses can be suggested to explain such variability: surviving cells do not activate caspases as well in response to the stimulus, or, the surviving cells do activate caspases but are better able to repair the cellular damage. The ability to monitor caspase activity in individual cells allowed us to begin to test such hypotheses.
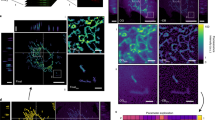
SHEP neuroblastoma cells undergo apoptosis in response to hyperosmolar stress induced by 300 mM mannitol.6 However, cells on the same plate die at very different times in an apparently random fashion when treated with this stimulus. Some cells continue to survive and divide up to 5 days after treatment, while many cells die between 13–24 h (data not shown). This variability in response allowed us to test whether there was a correlation between a particular cells' response and the level of caspase activity in that cell. A field of SHEP cells, grown from a single cell stably transfected with cleavable YFP-C-BFP, was chosen where approximately half the cells survived and half died during the experiment. Cells were scored as having died (outlined in red) or survived (outlined in yellow) by the end of the experiment (29 h) based on morphological criteria (cell rounding and membrane blebbing). Caspase activity was determined at 60 min intervals by quantitating yellow and blue fluorescence as before. Cells that were not treated with mannitol were not damaged when subjected to short bursts of UV light under these experimental conditions and continued to grow (data not shown). Figure 4a shows pseudocolored images of the pixel by pixel ratios at the beginning of the experiment and after approximately 22 h of mannitol treatment, when the first cell had died. At this time point, the cells that eventually died (marked in red) show a significant decrease in yellow/blue ratio as indicated by a change in the image from primarily white and yellow to primarily blue and black. The ratio change was much less for cells that survived as indicated by the modest alteration in color in those cells compared with the dying cells. These differences occurred despite the fact that both the cells that would survive and those that would eventually die displayed similar cell morphologies as demonstrated by the phase images. Figure 4b (top) shows quantitative data for six cells that died and seven cells that survived from the same field indicating the mean yellow/blue fluorescence ratio for each individual cell at approximately 13.3 h (800 min) after mannitol treatment. This time point was well before any death occurred in this field. Phase contrast images are shown for this time point and the end of the experiment in Figure 4a (lower panel). The slope of the lines corresponding to the change in fluorescence ratio during the first 13.3 h was calculated for each cell. Cells that died during the 29 h experiment (red lines) consistently had a steeper slope indicating more cleavage of the fusion protein than the cells that survived (yellow lines). The mean negative slopes for surviving cells and dying cells (insert in Figure 4b) are different with a significance value of P<0.001 by t-test. The lower panel in Figure 4b shows the normalized yellow/blue ratio at each time point for a dying and a living cell which best represented the average change for each respective group. At each time point, there was more caspase activity in the cell that eventually died. These data indicate that the cells that died displayed more caspase activity than the cells that survived, suggesting that the difference between surviving and dying cells depends upon their ability to activate caspases in response to the same stimulus.
Kinetic measurement of caspase activity in individual cells. (a) Shows a field of stable SHEP cells expressing the fusion protein before and 21.75 h after treatment with 300 nM mannitol followed using time-lapse microscopy at 37°C. The cells outlined in red died before the end of the experiment, cells outlined in yellow survived (bottom right). Ratio images were obtained as before. (b) (Top) shows quantitative data of the mean yellow/blue fluorescence for individual cells approximately 13.3 h (800 min) after mannitol treatment obtained from the original greyscale pictures. A phase contrast picture of this time point is shown in (a) (bottom left) along with the phase image when the experiment was stopped. Red lines indicate ratio changes from cells that died before the end of the experiment; yellow lines indicate ratio changes for cells that survived. The insert shows the mean negative slope for the red lines and yellow lines indicating that dying cells have a steeper slope (i.e. more caspase activity) than the surviving cells. The lower image shows the yellow/blue ratios at each time point for two sample cells. The vertical line marks the time of measurement for the upper graph (800 min)
Discussion
In this study, we describe a method that allows the continuous measurement of caspase activity in single living cells. This method is based on FRET measurements of GFP mutants.3 Alterations in the distance and orientation between fluorophores causes changes in FRET and such changes have been used to develop assays to measure various biological activities.3,5,7.8,9 A previous report described a GFP-BFP fusion protein that was cleaved by caspases resulting in changes in FRET that could be measured by FACS.10 In this case, ratio measurements were not made and changes in FRET were not detected in single cells. While we were completing our studies, Herman and colleagues11 described GFP-BFP fusion proteins that displayed caspase-dependent changes in FRET that could be detected in a fluorescence microscope. However, temporal measurements of caspase activity in individual cells were not performed in this study. Using our fusion protein, we obtained a more robust change in FRET emission upon in vitro digestion with caspase than that reported by Herman's group and we were able to measure both yellow and blue fluorescence in individual mammalian cells that could be followed for several hours. The method uses conventional fluorescence microscopy and should therefore be widely accessible.
The ability to measure caspase activity in individual, living cells should permit studies that would be otherwise difficult to perform. For example, we are currently performing experiments in primary human cells from surgical specimens that are only available in very small numbers (Morgan et al, submitted for publication). By combining fluorescence measurements with time-lapse microscopy, it is possible to continuously monitor caspase activity in individual cells and thus monitor temporal changes in caspase activity. Such changes can then be correlated with morphological changes in the cells or other cellular activities such as progression through the cell cycle.
In the experiments described here, we measured fluorescence after excitation at 380 nm so that all the yellow fluorescence was from FRET and we could measure a concomitant increase in blue fluorescence from the cleaved protein. Necrotic cell death where membrane integrity is lost early in the process would result in a reduction of both yellow and blue fluorescence because both cleaved and uncleaved proteins would leak out from the cell. This may be detectable by monitoring loss of fluorescence after direct excitation of YFP. In apoptotic cells and their membrane blebs, yellow fluorescence of the cleaved YFP can be obtained though direct excitation at 488 nm long after the loss of FRET-induced yellow fluorescence (data not shown). Therefore, it might be possible to identify both apoptotic and necrotic cells within a single population by measuring loss of FRET (in apoptotic cells) and loss of direct YFP excitation (in necrotic cells) at the same time. However, it should be noted that measurements of the loss of yellow fluorescence through leakage would be inherently less accurate than the ratio measurements that we used to measure caspase activity in apoptotic cells.
Our fusion protein could also be used in other applications. For example, by targeting the fusion protein to different sub-cellular compartments (cell membrane, nucleus, mitochondria, etc.), it should be possible to determine where caspase activity is located in cells that are treated with apoptotic stimuli. Recombinant fusion proteins and cell lines expressing the constructs may be useful for high throughput screening for drugs that moderate caspase activity and could be used to identify drugs that have organelle-specific effects. Finally, it should also be feasible to express the molecule in transgenic animals to permit single cell caspase activity measurements to be made in vivo.
Using our fusion protein, we showed that kinetic differences in caspase activation could be detected between individual cells within a group and could be correlated with each cells' eventual response to an apoptotic stimulus. Such differences occurred within clonal cells that were subjected to hyperosmolar stress, a non-selective physical stimulus that should have similar effects on every cell within a group. Importantly, these differences were apparent several hours before the cells died suggesting that the level of caspase activity in each cell determines its fate well before death actually occurs. These data suggest that some cells are inherently better able than others to activate caspases. Because the cells were clonal, it is unlikely that the differences are due to major changes in the levels of caspase regulators such as would occur when there is constitutive expression of a caspase inhibitor in tumor cells. It is possible that cells are variably able to respond to hyperosmolar stress at different stages of the cell cycle or that the differences we observe are due to some other variable. By measuring enzyme activities in single cells, it should now be possible to test such hypotheses and determine the molecular mechanisms that underlie the cell to cell variability in responsiveness.
Materials and Methods
Construction and digestion of YFP-BFP fusion protein
Several YFP-BFP fusion proteins were constructed and tested, maximum FRET was obtained with a peptide designated peptide C (TSTYGAGWNSRDEVDGNAYGRSDASLALPVAT) that contains a consensus cleavage site for caspase 3 and similar caspases (bold). A cDNA encoding enhanced YFP fused to enhanced BFP (Clontech, Palo Alto, CA, USA) linked via this peptide, was constructed in bacterial and eukaryotic expression plasmids by standard methods. The non-cleavable protein that was used in Figure 1c contained a highly structured linker peptide from the GluR3 glutamate receptor that contains no protease recognition sequences. The non-cleavable mutant fusion protein was constructed by site-directed mutagenesis to change the aspartic acid residue at the cleavage site to an alanine using a Quickchange kit (Stratagene, La Jolla, CA, USA). Recombinant GST-fusion proteins were purified from E. Coli and GST was removed by thrombin digestion. For in vitro digestion experiments, 10 μg protein was subjected to cleavage by purified active recombinant caspase (Pharmingen, La Jolla, CA, USA) in 150 μl assay buffer (20 nM PIPES, 100 mM NaCl, 10 nM DTT, 1 mM EDTA, 0.1% (w/v) CHAPS, 10% sucrose, pH 7.2) and incubated at 37°C then analyzed by gel electrophoresis and Western blotting or fluorescence spectroscopy.
FACS and fluorescence microscopy
The stable SHEP neuroblastoma cell line containing the YFP-C-BFP construct under the control of a CMV promoter was grown up from a single cell expressing the plasmid 2 weeks after transfection. Cells were harvested by scraping and analyzed on a FACS Vantage machine (Becton Dickinson), with excitation from the UV portion of an argon laser. Yellow and blue emission was detected. For transient expression studies, expression constructs were microinjected into the nucleus of Hela cells resulting in rapid expression of the fusion proteins.12 For microscopy, cells were maintained in an environmental chamber at 37°C and 5% CO2 on a Zeiss axiovert microscope. Phase and fluorescence images were captured using a Hamamatsu cooled CCD camera at 30 min intervals throughout the experiment. Greyscale images were obtained separately for both blue and yellow fluorescence using a filter wheel (Ludl) interchanging a 450 DF15 emission filter for blue fluorescence and a 535DF12.5 emission filter for yellow. Excitation light was provided by a 100 W mercury lamp in conjunction with a 380DF15 excitation filter and a 424DCLP BS dichroic mirror. Yellow/Blue ratio images were created from the primary data using the ratio module of the OpenLab (Improvision, Warwick, UK) software after normalization of the background using an area from the same field with no cells present. Quantitative data was obtained for each cell by creating a region of interest around the cell and then measuring the number of pixels of the greyscale images at each time point using the OpenLab software. For each cell, the average intensity for its defined area was determined and the average background intensity (determined from an equivalent adjacent area with no cell present) was subtracted. The Yellow/Blue ratio was then determined by dividing the mean intensity of the yellow picture divided by the mean intensity for the blue picture for each timepoint.
Abbreviations
- BFP:
-
blue fluorescent proteins
- FACS:
-
fluorescence activated cell sorting
- FRET:
-
fluorescence resonance energy transfer
- GFP:
-
green fluorescent protein
- YFP:
-
yellow fluorescent protein
References
Wolf BB and Green DR . 1999 Suicidal tendencies: apoptotic cell death by caspase family proteinases J. Biol. Chem. 274: 20049–20052
Thornberry NA and Lazebnik Y . 1998 Caspases: enemies within Science 281: 1312–1316
Pollok BA and Heim R . 1999 Using GFP in FRET-based applications Trends Cell Biol. 9: 57–60
Heim R . 1999 Green fluorescent protein forms for energy transfer Methods Enzymol. 302: 408–423
Heim R and Tsien RY . 1996 Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer Curr Biol 6: 178–182
Singleton JR, Dixit VM and Feldman EL . 1996 Type I insulin-like growth factor receptor activation regulates apoptotic proteins J. Biol. Chem. 271: 31791–31794
Miyawaki A, Llopis J, Heim R, McCaffrey JM, Adams JA, Ikuna M and Tsien RY . 1997 Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin Nature 388: 882–887
Romoser VA, Hinkle PM and Persechini A . 1997 Detection in living cells of Ca2+-dependent changes in the fluorescence emission of an indicator composed of two green fluorescent protein variants linked by a calmodulin-binding sequence. A new class of fluorescent indicators J. Biol. Chem. 272: 13270–13274
Nagai Y, Miyazaki M, Aoki R, Zama T, Inouye S, Hinose K, Iino M and Hagiwara M . 2000 A fluorescent indicator for visualizing cAMP-induced phosphorylation in vivo Nat. Biotechnol. 18: 313–316
Xu X, Gerard AL, Huang BC, Anderson DC, Payan DG and Luo Y . 1998 Detection of programmed cell death using fluorescence energy transfer Nucleic Acids Res. 26: 2034–2035
Mahajan NP, Harrison-Shostak DC, Michaux J and Herman B . 1999 Novel mutant green fluorescent protein protease substrates reveal the activation of specific caspases during apoptosis Chem. Biol. 6: 401–409
Alberts AS, Frost JA and Thorburn AM . 1993 Rapid transcription assay for the expression of two distinct reporter genes by microinjection DNA Cell Biol. 12: 935–943
Acknowledgements
We thank David Virshup and members of the Thorburn lab for helpful discussions and Wayne Green for help with FACS analysis. This work was supported by the Huntsman Cancer Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by DR Green
Rights and permissions
About this article
Cite this article
Morgan, M., Thorburn, A. Measurement of caspase activity in individual cells reveals differences in the kinetics of caspase activation between cells. Cell Death Differ 8, 38–43 (2001). https://doi.org/10.1038/sj.cdd.4400800
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4400800
Keywords
This article is cited by
-
Cell death stages in single apoptotic and necrotic cells monitored by Raman microspectroscopy
Scientific Reports (2014)
-
Features of programmed cell death in intact Xenopus oocytes and early embryos revealed by near-infrared fluorescence and real-time monitoring
Cell Death & Differentiation (2010)
-
An apoptosis signaling pathway induced by the death domain of FADD selectively kills normal but not cancerous prostate epithelial cells
Cell Death & Differentiation (2001)