Abstract
The genetic tools available in Drosophila have facilitated our understanding of how apoptosis is regulated and executed in the context of the developing organism. All embryonic apoptosis is initiated by the activity of three genes, rpr, grim and hid. Each of these genes is independently regulated, allowing developmental apoptosis to be finely controlled. These initiators in turn activate the core apoptotic machinery, including the caspases. Drosophila counterparts to other conserved components of the apoptotic machinery have been recently identified, and we discuss how these may be integrated into the process of normal developmentally regulated cell death. We also outline the role that phagocytosis plays in the final stages of apoptosis and consider the molecular mechanisms guiding the elimination of apoptotic corpses. Cell Death and Differentiation (2000) 7, 1027–1034.
Similar content being viewed by others
In the beginning there was genetics
Many of the molecules important in the apoptotic process were first identified by genetic means. Studies in the worm, C. elegans, showed that three genes, ced3, ced4 and ced9, formed the core of the apoptotic machinery.1,2 The cloning of the ced3 gene led to the identification of the caspases as central effectors of the apoptotic process.3 Similarly, the Ced4 protein was found to be an important modulator of caspase activity,4,5,6,7 and the Apaf1 protein was recognized as a mammalian Ced4 homologue with similar activity.8 The third gene, ced9, was discovered to be a homologue of a gene already implicated as a regulator of apoptosis in humans, bcl-2.9
Human genetics has also contributed to the discovery of genes involved in apoptosis. The bcl-2 gene was first discovered at the breakpoint of a translocation found in B cell lymphomas.10 Further work demonstrated that cells overexpressing bcl-2 became less susceptible to apoptosis.11 The tumor suppressor p53 was also found to affect sensitivity to a variety of death inducers.12,13 The absence of p53, as seen in many tumors, made cells more resistant to death.
These discoveries demonstrated the value of genetics as a tool to investigate the regulation and mechanisms of apoptosis. In the early 1990's, Hermann Steller and colleagues tested the usefulness of Drosophila as a genetic model system to study this process. Although apoptosis had been observed to occur during Drosophila development, no systematic attempt had been made to identify genes that acted as central regulators of the process. Initial studies demonstrated that apoptosis in flies resembled apoptosis in other systems, both ultrastructurally and biochemically.14 Based on these findings, a genetic screen was undertaken to identify mutations affecting the apoptotic process.
All genetic screens are based on assumptions as to the selected phenotype. In C. elegans, mutations affecting apoptosis were identified as adults lacking apoptotic corpses.1 Some assumptions of the C. elegans screen were that cell death defective mutations would be viable, and that single gene mutations would detectably disrupt apoptosis. Related assumptions were made in the initial screen for Drosophila cell death defective mutations.15 As mutant embryos were screened for perturbations in the patterns of apoptosis, it was assumed that these embryos would live until the stage when apoptosis could be easily assayed. Rather than screening single gene mutations for defects in apoptosis, a collection of genomic deletions was assayed. It was considered likely that any apoptosis mutations detected would be the result of a single gene mutation.
Although the screen was a success, identifying a single genomic region that was essential for all apoptosis during embryogenesis, both of the above assumptions turned out to be incorrect. Mutants in the thread/dlAP1 gene, an important antiapoptotic regulator, die very early in embryogenesis and were missed in the initial screens.16,17,18 Additionally, multiple genes are responsible for the cell death defective phenotype seen in deletions overlapping the genomic region identified in the screen.15
After the identification of these deletions, attempts were made to generate single gene mutations that recapitulated the cell death defective phenotype. Several years of work led to the realization that the deletion of more than one gene was responsible for the defect. Eventually, three genes were identified that seem to act in a cooperative manner to regulate apoptosis in the embryo. These three genes, reaper (rpr), head involution defective (hid) and grim, share a short stretch of homology at their N-terminus, but are otherwise dissimilar.15,19,20 Deletions that remove only one of these genes have very weak effects on embryonic apoptosis, while the removal of two has stronger effects.15,19,20
Reaper, hid and grim are central regulators of apoptosis
What is the evidence that these three genes act as apoptotic regulators? First, as stated above, deletions that remove combinations of these genes have significant effects on the amount of cell death in the embryo.15,19,20 Second, each of these genes is capable of inducing apoptosis when ectopically expressed in the developing fly and in both Drosophila and mammalian cultured cells.19,20,21,22,23,24,25,26 Third, the expression of these genes corresponds to patterns of apoptosis. Rpr and grim appear to be exclusively and specifically expressed in cells that are doomed to die,15,19 while hid is more broadly expressed, both in cells that live and in cells that die.20 In addition hid is not expressed in the central nervous system, a site of significant levels of cell death in the embryo.
Initial characterization of the phenotype of embryos homozygous for the smallest deletion that removes all three genes (H99) suggested that these genes are not the actual effectors of apoptosis. Apoptosis could be generated in these embryos by high doses of x-ray irradiation, indicating that the core apoptotic machinery was intact.15 Caspase activation was shown to be essential for killing by each of the genes.19,20,21 The baculovirus p35 protein, which acts as a broad-spectrum caspase inhibitor, inhibits death induced by the ectopic expression of each gene. The mechanisms by which rpr, grim and hid activate caspases are not fully understood. In vitro binding experiments suggest that at least part of their proapoptotic activity is the result of their ability to bind to and inactivate the antiapoptotic IAP proteins.
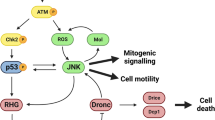
One attractive model for the regulation of cell death during development is that when the combined activity of Rpr, Hid and Grim in an individual cell exceeds a threshold, caspases are activated and the cell undergoes apoptosis. The activity of each of these genes is regulated by different upstream regulatory signals (Figure 1). Thus the decision of a cell to die is the integration of several regulatory pathways. Most of these regulatory pathways are currently unknown, but several have been identified and include developmental cues, cell to cell signaling mechanisms and internal sensing programs.
Reaper, Grim and Hid act in a cooperative manner to induce apoptosis. When the combined activity of the three proteins exceeds a threshold, apoptosis is induced. Each of these inducers can be regulated independently. Expression of rpr is induced by DNA damage,29,30 misregulation of the cell cycle,99 developmental signals15,27 and steroid hormone levels.28 grim and hid expression are also responsive to developmental signals.19,20 In addition, hid activity is negatively regulated by the EGF receptor pathway31,32
The transcription of rpr is clearly directed by a number of developmental cues, as it is expressed in many different cells that are fated to die during development. Rpr expression is also upregulated when cells die due to defects in the normal developmental program.27 The elaborate expression pattern of rpr suggests that its promoter may be responsive to a large number of developmental pathways. Responses to individual pathways may be directly integrated at the promoter, or may reflect the activity of a limited number of transcription factors, which in turn integrate several pathways.
Current data indicate that at least two pathways impact directly on the rpr promoter: the steroid hormone pathway and the p53 pathway. The expression of rpr is directly regulated by the steroid hormone ecdysone.28 This expression occurs in a number of obsolete larval tissues just prior to the onset of cell death. In addition, intracellular signaling in response to chromosomal damage leads to the expression of rpr and the subsequent death of the cell.27,29 Recent work has shown that the Drosophila homologue of p53, dmp53, is responsible for this signaling and acts through a p53-response element within the upstream regulatory region of rpr.29,30
The regulation of hid appears to be somewhat more complex than that of rpr. hid expression is observed in a variety of cells that do not undergo apoptosis, although ectopic over-expression of hid is a potent inducer of death.20 The activity of hid is modulated by growth factor signaling, acting through the Ras/Raf/MAPK pathway.31,32 MAPK activity inhibits hid both transcriptionally and post-translationally, probably through direct phosphorylation. In the developing eye and in the embryonic midline glia the epidermal growth factor receptor (EGFR) activates this antiapoptotic activity of Ras.31,32,33,34,35,36,37 Thus, cell survival is favored when cells are exposed to high levels of EGFR ligands, and cells undergo hid-induced apoptosis in the absence of this activity. Little is known about the regulation of grim expression, although it is also transcriptionally regulated during development, and rpr and grim expression appear quite similar.19
Rpr, Grim and Hid kill by inactivating IAPs
While it is clear that programmed cell death in the Drosophila embryo is initiated by the death regulators rpr, grim and hid, and that death itself is mediated through the proteolytic action of caspases, the mechanisms connecting the initiation of apoptosis to caspase activation are less clear. One class of anti-apoptotic molecules that has been shown to play a role in this process is the IAPs, or Inhibitors of Apoptosis Proteins. Originally identified in baculoviruses, the IAPs have been found in virtually all multicellular organisms that have been examined to date.38,39 IAPs are characterized by a well-conserved BIR domain (Baculovirus IAP Repeat). Many also have a well-conserved RING domain located near the C-terminus of the protein. Both of these structural domains consist of arrangements of cysteines and histidines that coordinate zinc ions.
In Drosophila, three IAPs, dIAP1, dIAP2 and deterin, have been characterized.40,41 Ectopic expression of either dIAP1 or dIAP2 suppresses killing by ectopically expressed rpr, grim or hid and also partially suppresses normal developmental cell death.41 When expressed in cultured cells deterin suppresses killing by over-expressed rpr and by cytotoxic insult.40
DIAP1 directly binds to and inhibits a number of Drosophila caspases.42,43,44 There are five characterized caspases in Drosophila. Two of these, Dredd and Dronc, appear to be apical caspases with long prodomains, while Dcp-1, drICE and Decay appear to be effector caspases. The inactive proforms of both Dronc and Dcp-1 are efficiently bound by DIAP1, while binding to drICE appears to occur only following cleavage of this caspase into active subunits. Thus, it seems that at least part of the function of DIAP1 is to complex with and inactivate previously activated caspases (i.e. drICE) or to maintain caspase proforms in their inactive states.
This caspase inhibitory activity is consistent with the observation that homozygous loss of function alleles of the th/dIAP1 gene are lethal very early in embryogenesis due to massive apoptosis.16,17,18 While the protective mechanism of DIAP1 can perhaps be entirely explained by its ability to interact with and inhibit caspases, this has not yet been shown for either DIAP2 or Deterin. It may be that these IAPs, as well as DIAP1, function through an as yet unidentified mechanism. In addition, Drosophila IAPs may have functions unrelated to apoptosis, as they do in other systems.45,46
When expressed in cultured cells, both DIAP1 and DIAP2 bind to the apoptotic inducers Rpr, Grim and Hid through their BIR domains.47,48 This has led to the model that Rpr, Grim and Hid kill by binding to and suppressing the protective function of IAPs. This model is strongly supported by the demonstration that DIAP1 directly binds to and inhibits the activity of the caspase DCP-1 and that the binding of Hid abrogates this caspase inhibition.18 Further support for such a mechanism comes from studies of mutant alleles of the th/dIAP1 gene which suppress the rough eye phenotype induced by ectopic expression of rpr and hid. These mutations result in DIAP1 molecules that can no longer be bound by either Rpr or Hid. Thus Rpr and Hid can no longer disrupt DIAP1-mediated inhibition of caspase activity.16
As the only conserved regions shared by Rpr, Grim and Hid are contained within the first 14 amino acids of each molecule, it would be reasonable to expect that binding to DIAP1 and the initiation of cell death would be mediated by these domains. This appears to be the case for Hid, as removal of the first 14 amino acids abolishes both the ability of Hid to interact with DIAP1 and to kill cultured cells.47 Oddly, deletion of the first 15 amino acids from either Rpr or Grim does not prevent either of these molecules from initiating cell death,49,50,51 and both N-terminally truncated Grim and Rpr are still able to co-precipitate with DIAP1. (Bangs and White, unpublished observations; and47) Coupled with the fact that the N-terminal portion of Grim, like that of Hid, is sufficient to both bind DIAP1 and induce killing, these data indicate that there are at least two domains in Grim that are capable of interacting with DIAP1 and inducing cell death.47 Further differences in the ways in which Rpr, Grim and Hid interact with DIAP1 are seen with certain gain of function mutations in the th/dIAP1 gene. One class of mutations suppresses killing by ectopically expressed rpr and grim, but has no effect on killing by hid. The other class of th/dIAP1 mutations instead suppresses killing by ectopically expressed hid, but has no appreciable impact on rpr or grim killing.17 One might expect that these mutations would also exhibit differential binding capabilities with respect to Rpr and Grim on the one hand and Hid on the other.
It is important to note that the recently identified SMAC/DIABLO protein in mammalian systems may be a functional homologue of Rpr, Hid and Grim.52,53 This protein is released from the mitochondria on receipt of a death-inducing signal, and potentiates caspase activation. SMAC/DIABLO has been shown to bind to IAPs, and thus may augment caspase activity by inactivating the caspase-inhibitory function of mammalian IAPs. SMAC/DIABLO does not have any sequence homology to Rpr, Hid and Grim, but it seems to promote apoptosis through the same mechanism.
Caspases are required for Rpr, Grim and Hid killing
Currently, five caspases have been characterized in Drosophila, and three others have been predicted based on genomic sequence information. Of these, the caspases Dredd/Dcp-254,55 and Dronc56 have the long N-terminal prodomains consistent with initiator caspases, while Dcp-157, drICE58 and Decay59 have the shorter prodomains associated with the executioner caspases. In addition, one of the three predicted, but uncharacterized, caspases has a long ‘initiator’ type prodomain, while the other two appear to be of the short prodomain ‘executioner’ class.60
The caspase Dronc mediates cell death initiated by rpr, grim and hid, as a reduction in the dosage of Dronc, or expression of a catalytically inactive Dronc, greatly suppresses the eye ablation phenotype due to ectopic rpr, grim or hid expression.44,61 Further, ectopically expressed Dronc itself can kill in yeast, cultured cells and cells in the developing Drosophila eye.44,56,61 Dronc effectively cleaves drICE44,61, as predicted for an apical caspase. Dronc is expressed highly in the embryo, and in tissues that will degenerate in response to the steroid hormone ecdysone. Thus it is expressed at the right time to be involved in apoptosis initiated by rpr, grim or hid.
Dredd appears to be an apical caspase by virtue of its relatively long prodomain and its association with the ced4/Apaf-1 homologue Ark.55,62 Reductions in the dosage of dredd have a suppressive effect on killing by ectopically expressed rpr and grim, indicating that Dredd, like Dronc, mediates killing by these death activators. Intriguingly, Dredd may not play a significant role when cell death is initiated by hid, as reduction of dredd does not appear to appreciably affect killing by ectopically expressed hid. Among the Drosophila caspases that have been characterized thus far, only dredd expression is linked specifically to cell death, and transcripts accumulate in cells that are specified to die. This accumulation in doomed cells is apparently coupled to the activity of the cell death inducers rpr, grim and perhaps hid, as embryos homozygous for the H99 deletion, thus lacking rpr, grim and hid, do not show wild-type accumulation of dredd mRNA. The mechanism by which cells accumulate dredd mRNA in response to rpr, grim and hid is presently unknown.
There is some evidence that specific executioner caspases act downstream of individual death inducers, as ectopic expression of rpr and grim seems to promote the activation of Dcp-1.63 In addition some caspases may play unique roles in specific developmental processes. In the absence of dcp-1, oogenesis is abnormal.64 However, no mutant alleles have been characterized for many of these caspases. In general, the characterization of Drosophila executioner caspases lags significantly behind the mammalian studies.
The mitochondrial pathway
Until recently, no Drosophila homologues for either ced4/Apaf-1 or ced9/bcl-2 had been found. The discovery of Ark, a ced4/Apaf1 homologue62,65,66 and debcl and Buffy, ced9/bcl-2 family members67,68,69,70, has solidified the concept that apoptotic mechanisms are well conserved across species. In both the worm and in mammalian systems, ced4/Apaf1 activation promotes caspase activation.71 Pro- and anti-apoptotic members of the ced9/bcl2 family modulate this activation. In mammalian systems Apaf1 activity is upregulated by binding to cytochrome c, which is released from the mitochondria upon induction of death. This ‘mitochondrial pathway’ seems to enhance killing by a number of inducers, and may act as an amplifier of an apoptotic signal.
The Drosophila ced4/Apaf1 homologue dark/Dapaf-1/HAC-1 is listed in sequence databases as Ark.62,65,66 Like its counterparts in worms and mammalian cells, Ark has an apparent nucleotide binding site in the N-terminal Ced-4 domain, and like Apaf-1, but distinct from Ced4, the C-terminal domain contains two series of WD repeats. This WD domain seems to act as a negative regulator of Ark function. Cytochrome c binding releases this inhibition. An alternatively spliced isoform of Ark lacking this inhibitory WD domain has also been reported, although it is unclear whether it is expressed at significant levels.65
Ectopic expression of Ark in Drosophila S2 cells induces a moderate amount of cell death at best, but expression of a truncated version lacking WD repeats results in a substantial increase in cell killing.62,65 Genetic evidence indicates that Ark is an important component of death pathways activated by rpr, grim and hid.62,65,66 The normally robust eye ablation phenotypes observed for ectopically expressed rpr, grim and hid are substantially suppressed by Ark mutations.62,65 Decreased Ark also suppresses death in eyes that ectopically express dcp-1.66 Interestingly, a strong synergistic effect between ectopically expressed rpr and dcp-1 is not suppressed to any significant degree in Ark heterozygotes. Taken together, these latter two observations suggest that Dcp-1 activation by Ark and Rpr may occur through independent pathways.
Embryos mutant for Ark exhibit reduced cell death, particularly in the CNS and epidermal regions, and the larval nervous systems are substantially larger than wild-type and have excess cells.62,65,66 Adult flies have a number of relatively mild defects including abnormal wings, a few extra bristles, occasional melanotic tumors and extra photoreceptors and/or pigment cells.62,65 In addition, approximately 47% of the males homozygous for a null mutation are sterile.62 These relatively mild and weakly penetrant phenotypes and the observations of at least some developmental cell death in these mutants, indicate that Ark is not absolutely necessary for normal cell death to occur, but may instead contribute greatly to the efficiency of the process.
Considering the conservation of the apoptotic machinery from worms to humans, it comes as no surprise that Drosophila would have Bcl-2 family members as part of its cell death repertoire. Several laboratories have identified and characterized the proapoptotic Debcl/dBorg-1/dRob-1.67,68,69 This protein contains the BH1, BH2 and BH3 domains as well as a C-terminal transmembrane domain that localizes it to intracytoplasmic membranes, primarily the mitochondria. Ectopic expression of Debcl increases cell death in a variety of tissues and in the eye produces a rough to severely ablated phenotype in a dose-dependent manner. This phenotype is dramatically suppressed by the caspase inhibitor p35 demonstrating that Debcl functions in a caspase-dependent manner in the fly.67,69
In cultured cells, Debcl has a potent killing activity that is conserved across several species. Interestingly, several cell types, including Drosophila S2 cells and COS cells are relatively refractory to killing by Debcl, and in these cells Debcl may in fact be protective when expressed at levels that do not induce death.67 When Debcl does induce killing in S2 cells, caspase activation is also observed.68 In contrast to what is seen in Drosophila eyes, p35 does not appear to prevent killing by Debcl in S2 cells although caspase activation is abolished. This suggests that in some situations, Debcl may induce cell death by both caspase-dependent and caspase-independent mechanisms.
As is the case for other members of the Bcl2 family, the mechanism of Debcl function is unclear. One possibility is that Debcl binds to and neutralizes the activity of anti-apoptotic Bcl2 homologues. This hypothesis, hampered somewhat by the fact that no pro-survival Bcl-2 homologues have yet been reported in Drosophila, is supported by the observation that Debcl co-precipitates with a number of the anti-apoptotic Bcl2 family members from other species.69 Genetic studies show that Debcl interacts with the Drosophila Apaf1/ced4 homologue Ark and that reducing the dosage of Ark significantly suppresses the rough eye phenotype caused by ectopic expression of Debcl. Debcl also interacts with the apoptotic inhibitor th/dIAP1 as shown by the fact that when the gene dosage of th/dIAP1 is halved, the rough eye phenotype produced by Debcl is strongly enhanced. This does not appear to be true for interactions between Debcl and dIAP2. Reducing the dosage of the cell death-inducing genes rpr, grim and hid does not appear to have any effect on killing by Debcl, suggesting that the activity of these genes is not rate limiting in Debcl-induced death.69
A second Drosophila bcl-2 family member, Buffy, has also been reported, although it has not been thoroughly characterized.67,69,70 Other family members may also be found in the newly completed genome sequence.
How do the mitochondrial pathway and the Rpr, Grim, Hid/DIAP pathway interact?
The mechanisms by which Rpr, Grim and Hid kill are not fully understood, but a model can be made based on the existing data (Figure 2). Assuming that experiments done in vitro accurately reflect the situation in vivo, it is likely that rpr, hid and/or grim expression or Hid activation, above a threshold level, initiates a cascade of caspase activity by binding to the protective molecule DIAP1 and disrupting its caspase-inhibitory activity. Given the fact that Dronc is among those caspases that are regulated by DIAP1, it is expected that Dronc activation would occur fairly early in the process, presumably by self-cleavage, and that an active Dronc would subsequently process other caspases as well.44 Circumstantial evidence suggests that Dredd, whose expression is up-regulated following the expression of rpr, grim or hid, would be an early target of the active Dronc.55
A model for the interactions between the inducers Rpr, Hid and Grim, and the apoptotic effector machinery. The interactions between Rpr, Grim and Hid and DIAP1 and the caspases are supported by experimental data (represented by solid lines, see text). Dotted lines represent purely speculative interactions between these inducers and the mitochondrial/Ark pathway, including the bcl-2 family homologues and the scythe homologue. The pro- or anti-apoptotic activity of Buffy has not yet been reported
It is unclear whether the Bcl2-related Debcl or the Apaf-1 homologue Ark are absolutely necessary for the preceding scenario to occur or whether they are utilized instead to improve the efficiency or timing of the process. The connection between the cell death regulators Rpr, Grim and Hid on the one hand and Debcl and/or Ark on the other is also murky. One possibility is that the pathways are initiated independently by the death stimuli, and only interact at the level of caspase activation. Another possibility is that the Drosophila homologue of Xenopus Scythe (CG7546 in the GadFly genome annotation database) connects Rpr, and possibly Grim and Hid to caspase activation by Debcl and Ark. Xenopus Scythe has been shown to bind to Rpr.72 As a consequence of the interaction of Rpr with Xenopus Scythe, an unidentified proapoptotic factor is released from Scythe.73 This unidentified factor possesses a cytochrome c releasing activity when combined with mitochondria, and could therefore be responsible for cytochrome c dependent activation of an Apaf-1-like molecule in Xenopus. A possible scenario is that Drosophila Scythe sequesters a proapoptotic factor that is released when one of the cell death activators interacts with Scythe.
Mammalian pro-apoptotic Bcl2-family members such as Bid and Bim are sequestered from mitochondria until activated by cleavage or signaling.74 Debcl (and Buffy) might also be sequestered in living cells, as they are broadly expressed. It is possible that Drosophila Scythe might play a role in this sequestration, or that caspase activation downstream of IAP inactivation might result in the release of the proapoptotic Bcl-2 proteins. These proteins could then mediate either a release of cytochrome c from the mitochondria or participate in the altered display of cytochrome c observed with mitochondria in the early stages of programmed cell death in Drosophila.75 Once cytochrome c was available, cytochrome c-dependent interactions between Ark and either Dredd or Dronc could result in the activation of these caspases and an amplification of the caspase cascade.
While the preceding scenario is obviously speculative, it attempts to tie many of the known components of the Drosophila apoptotic machinery with the apoptotic pathways of other systems. Clearly, more work needs to be done to reach a decent understanding of the molecular mechanisms that govern and execute this important process. The genetic tools available in Drosophila will be invaluable for the identification of both pro- and anti-apoptotic molecules and for the delineation of the apoptotic pathways in the living organism.
Phagocytosis is a crucial final step in apoptosis
Cells that are programmed to die must be cleared rapidly. This is an essential element of the apoptotic program, as persistent corpses may release cytotoxic substances and damage neighboring developing tissues.76,77 In all multicellular organisms, apoptotic cells are removed through phagocytosis. Phagocytosis is the process by which cells engulf and digest large particles that they recognize as ‘non-self’ or effete self. In vertebrates, phagocytosis can be carried out either by ‘professional’ or by ‘non-professional’ phagocytes, i.e. by cells whose major function is engulfment or by cells with other functions that are capable of engulfment.78 In C. elegans, apoptotic corpses are phagocytosed by neighboring cells.79 In the Drosophila embryo there is some evidence of phagocytosis by glial cells.80 However, the most efficient embryonic phagocytes are professional ‘macrophages’, the phagocytic hemocytes or blood cells.
Drosophila embryonic macrophage precursors originate from the procephalic mesoderm approximately 2 h after gastrulation.81 These 40 precursors give rise to approximately 700 hemocytes. This complement of hemocytes remains constant throughout embryogenesis, independent of the amount of apoptosis in the embryo. Emerging hemocytes are scattered round or irregularly shaped cells that migrate freely in the hemocoel (plasma) and spread throughout the embryo.
Most hemocytes become phagocytic macrophages as they encounter apoptotic cells.81 Recent work has shown that there are in fact two major classes of hemocytes in the embryo: the macrophage lineage (also called plasmatocytes), and the crystal cell lineage.82 Crystal cells participate in the sequestration of large particles by melanization.83 It is interesting to note that the differentiation of these lineages requires the activity of specific transcription factors, similar to those in mammalian systems. The GATA homologue serpent is required for all hemocyte differentiation84, while the AML1 homologue Lozenge is required for the crystal cell lineage.82 The gcm transcription factor plays a role in macrophage differentiation.85
The macrophage lineage can be distinguished by the expression of Peroxidasin, an extracellular matrix protein produced by these cells.86 By the end of embryogenesis approximately 90–95% of the Peroxidasin-positive hemocytes are phagocytic.81 This number can rise to 100% in the presence of excess apoptosis, suggesting that the presence of apoptotic corpses might be important for the terminal differentiation of macrophages and/or to trigger their phagocytic activity. Although macrophage-induced apoptosis is evident in the developing mouse eye,87 there is no evidence that Drosophila macrophages play an active role in apoptosis. Indeed apoptosis seems to proceed normally in mutants that lack all head mesoderm and are devoid of hemocytes.81
A critical step in phagocytosis is the recognition of the target to be engulfed. This requires receptors that recognize specific biochemical patterns at the surface of the target.88 Croquemort (Crq), a Drosophila member of the CD36 family of receptors, is one such receptor.89 Crq is expressed exclusively on macrophages during embryogenesis. It is most prominently expressed at the membrane surface of subcellular vesicles that contain apoptotic corpses. Moreover, crq expression is sufficient to trigger the recognition and engulfment of apoptotic cells in a heterologous system. Importantly, in the absence of crq, macrophages do not efficiently remove apoptotic corpses.90 However, crq-deficient macrophages are able to carry out all other known macrophage functions, including the engulfment of bacteria. These results argue that crq specifically plays a role in phagocytosis of apoptotic corpses in the embryo.
The homology between Crq and mammalian CD36 suggests conservation in the mechanisms underlying phagocytosis of apoptotic corpses throughout evolution.90,91,92 CD36 was shown to act in concert with the vitronectin receptor, an integrin, in the recognition of apoptotic corpses via a bridge of thrombospondin, an extracellular matrix protein.92 Whether Crq requires other partners to assemble in a phagocytic complex remains to be explored. Perhaps most interesting will be the elucidation of the molecular nature of the pattern recognized by this phagocytic complex. In addition, the role of crq in the phagocytosis of apoptotic corpses at later stages of development is unknown. However, Crq is expressed on plasmatocytes, the phagocytic hemocytes of larval and pupal stages.82,90
Other receptors that may play a role in the recognition of apoptotic cells in the embryo are the Drosophila phosphatidylserine (PS) receptor,93 and the Drosophila scavenger receptor, DSR-CI.94 PS exposure is a common feature of apoptotic death,95 and a recently identified mammalian PS receptor appears to be both necessary and sufficient to promote the engulfment of apoptotic corpses in some cell types.93 The Drosophila homologue is highly conserved, but its function has not yet been characterized. Although the physiological significance of scavenger receptor activity remains unknown in the fly, a number of scavenger receptors have been proposed to participate in phagocytosis of apoptotic cells in mammals.96,97,98
Genetic strategies can clearly be used to evaluate the requirement of specific molecules in apoptosis as well as phagocytosis of apoptotic corpses in Drosophila. The elaborate regulation of apoptosis during development and the role of apoptosis in a variety of developmental processes are just beginning to be examined. In addition it is apparent that considerable work is still to be done to understand how the various components of the death machinery interact to kill cells. Finally, the genetic dissection of phagocytosis in the fly will provide important information on the recognition and clearance of apoptotic cells, as well as pathogens. It is clear that the apoptotic process is highly conserved, and insights gained in such model systems will have implications in other organisms.
References
Ellis H and Horvitz HR . 1986 Genetic control of programmed cell death in the nematode C. elegans. Cell. 44: 817–829
Hengartner MO, Ellis RE and Horvitz HR . 1992 Caenorhabditis elegans gene ced-9 protects cells from programmed cell death. Nature 356: 494–499
Yuan J, Shaham S, Ellis HM and Horvitz HR . 1993 The C. elegans cell death gene ced-3 encodes a protein similar to mammalian interleukin - 1β -converting enzyme. Cell. 75: 641–652
Chinnaiyan AM, Chaudhary D, O'Rourke K, Koonin EV and Dixit VM . 1997 Role of CED-4 in the activation of CED-3. Nature 388: 728–729
Irmler M, Hofmann K, Vaux D and Tschopp J . 1997 Direct physical interaction between the Caenorhabditis elegans ‘death proteins’ CED-3 and CED-4. FEBS Lett. 406: 189–190
Wu D, Wallen HD, Inohara N and Nunez G . 1997 Interaction and regulation of the Caenorhabditis elegans death protease CED-3 by CED-4 and CED-9. J. Biol. Chem. 272: 21449–21454
Seshagiri S and Miller LK . 1997 Caenorhabditis elegans CED-4 stimulates CED-3 processing and CED-3- induced apoptosis. Curr. Biol. 7: 455–460
Zou H, Henzel W, Liu X, Lutschg A and Wang X . 1997 Apaf-1, a human protein homologous to C. elegans CED-4, participates in cytochrome c-dependent activation of caspase-3. Cell. 90: 405–413
Hengartner MO and Horvitz HR . 1994 C. elegans cell survival gene ced-9 encodes a functional homolog of the mammalian proto-oncogene bcl-2. Cell 76: 665–676
Tsujimoto Y, Ikegaki N and Croce CM . 1987 Characterization of the protein product of bcl-2, the gene involved in human follicular lymphoma. Oncogene 2: 3–7
Vaux DL, Cory S and Adams JM . 1988 Bcl-2 gene promotes heamopoietic cell survival and cooperates with c-myc to immortalize pre-B cells. Nature 335: 440–442
Lowe SW, Ruley HE, Jacks T and Housman DE . 1993 p53-dependent apoptosis modulates the cytotoxicity of anticancer agents. Cell. 74: 957–967
Tanaka N, Ishihara M, Kitagawa M, Harada H, Kimura T, Matsuyama T, Lamphier MS, Aizawa S, Mak TW and Taniguchi T . 1994 Cellular commitment to oncogene-induced transformation or apoptosis is dependent on the transcription factor IRF-1. Cell. 77: 829–839
Abrams JM, White K, Fessler LI and Steller H . 1993 Programmed cell death during Drosophila embryogenesis. Development 117: 29–43
White K, Grether ME, Abrams JM, Young L, Farrell K and Steller H . 1994 Genetic control of programmed cell death in Drosophila. Science 264: 677–683
Goyal L, McCall K, Agapite J, Hartwieg E and Steller H . 2000 Induction of apoptosis by Drosophila reaper, hid and grim through inhibition of IAP function. EMBO J. 19: 589–597
Lisi S, Mazzon I and White K . 2000 Diverse domains of THREAD/DIAP1 are required to inhibit apoptosis induced by REAPER and HID in Drosophila. Genetics 154: 669–678
Wang S, Hawkins C, Yoo S, Müller H-A and Hay B . 1999 The Drosophila caspase inhibitor DIAP1 is essential for cell survival and is negatively regulated by HID. Cell 98: 453–463
Chen P, Nordstrom W, Gish B and Abrams JM . 1996 grim, a novel cell death gene in Drosophila. Genes Dev. 10: 1773–1782
Grether ME, Abrams JM, Agapite J, White K and Steller H . 1995 The head involution defective gene of Drosophila melanogaster functions in programmed cell death. Genes Dev. 9: 1694–1708
White K, Tahaoglu E and Steller H . 1996 Cell killing by the Drosophila gene reaper. Science 271: 805–807
Haining W, Carboy-Newcomb C, Wei C and Steller H . 1999 The proapoptotic function of Drosophila Hid is conserved in mammalian cells. Proc. Natl. Acad. Sci. USA 96: 4936–4941
Claveria C, Albar J, Serrano A, Buesa J, Barbero J, Martinez-A C and Torres M . 1998 Drosophila grim induces apoptosis in mammalian cells. EMBO J. 17: 7199–7208
McCarthy J and Dixit V . 1998 Apoptosis induced by Drosophila Reaper and Grim in a human system. J. Biol. Chem. 273: 24009–24015
Pronk GJ, Ramer K, Amiri P and Williams LT . 1996 Requirement of an ICE-like protease for induction of apoptosis and ceramide generation by REAPER. Science 271: 808–810
Evans E, Kuwana T, Strum S, Smith J, Newmeyer D and Kornbluth S . 1997 Reaper-induced apoptosis in a vertebrate system. EMBO J. 16: 7372–7381
Nordstrom W, Chen P, Steller H and Abrams JM . 1996 Activation of the reaper gene during ectopic cell killing in Drosophila. Dev. Biol. 180: 213–226
Jiang CA, Lamblin AFJ, Steller H and Thummel CS . 2000 A steroid-triggered transcriptional hierarchy controls salivary gland cell death during Drosophila metamorphosis. Mol. Cell 5: 445–455
Brodsky MH, Nordstrom W, Tsang G, Kwan E, Rubin GM and Abrams JM . 2000 Drosophila p53 binds a damage response element at the reaper locus. Cell 101: 103–113
Ollmann M, Young LM, Di Como CJ, Karim F, Belvin M, Robertson S, Whittaker K, Demsky M, Fisher WW, Buchman A, Duyk G, Friedman L, Prives C and Kopczynski C . 2000 Drosophila p53 is a structural and functional homolog of the tumor suppressor p53. Cell. 101: 91–101
Kurada P and White K . 1998 Ras promotes cell survival in Drosophila by downregulating hid expression. Cell. 95: 319–329
Bergmann A, Agapite J, McCall K and Steller H . 1998 The Drosophila gene hid is a direct molecular target of Ras-dependent survival signaling. Cell. 95: 331–341
Stemerdink C and Jacobs JR . 1997 Argos and Spitz group genes function to regulate midline glial cell number in Drosophila embryos. Development 124: 3787–3796
Sawamoto A, Taguchi A, Hirota Y, Yamada C, Jin M and Okano H . 1998 Argos induces programmed cell death in the developing Drosophila eye by inhibition of the ras pathway. Cell. Death Differ. 5: 262–270
Scholz H, Sadlowski E, Klaes A and Klambt C . 1997 Control of midline glia development in the embryonic Drosophila CNS. Mech. Dev. 64: 137–151
Miller DT and Cagan RL . 1998 Local induction of patterning and programmed cell death in the developing Drosophila retina. Development 125: 2327–2335
Freeman M . 1996 Reiterative use of the EGF receptor triggers differentiation of all cell types in the Drosophila eye. Cell. 87: 651–660
Miller L . 1999 An exegesis of IAPs: salvation and surprises from BIR motifs. Trends Cell. Biol. 9: 323–328
Deveraux QL and Reed JL . 1999 IAP family proteins-suppressors of apoptosis. Genes Dev. 13: 239–252
Jones G, Jones D, Zhou L, Steller H and Chu Y . 2000 Deterin, a new inhibitor of apoptosis from Drosophila melanogaster. J. Biol. Chem. 275: 22157–22165
Hay BA, Wassarman DA and Rubin GM . 1995 Drosophila homologs of baculovirus inhibitor of apoptosis proteins function to block cell death. Cell 83: 1253–1262
Hawkins C, Wang S and Hay B . 1999 A cloning method to identify caspases and their regulators in yeast: Identification of Drosophila IAP1 as an inhibitor of the Drosophila caspase DCP-1. Proc. Natl. Acad. Sci. USA 96: 2885–2890
Kaiser W, Vucic D and Miller L . 1998 The Drosophila Inhibitor of Apoptosis D-IAP1 suppresses cell death induced by the caspase drICE. FEBS Lett. 440: 243–248
Meier P, Silke J, Leevers SJ and Evan GI . 2000 The Drosophila caspase DRONC is regulated by DIAP1. EMBO J. 19: 598–611
Fraser AG, James C, Evan GI and Hengartner MO . 1999 Caenorhabditis elegans inhibitor of apoptosis protein (IAP) homologue BIR-1 plays a conserved role in cytokinesis. Curr. Biol. 9: 292–301
Uren AG, Beilharz T, O'Connell MJ, Bugg SJ, van Driel R, Vaux DL and Lithgow T . 1999 Role for yeast inhibitor of apoptosis (IAP)-like proteins in cell division. Proc. Natl. Acad. Sci. USA 96: 10170–10175
Vucic D, Kaiser WJ and Miller LK . 1998 Inhibitor of apoptosis proteins physically interact with and block apoptosis induced by Drosophila proteins HID and GRIM. Mol. Cell. Biol. 18: 3300–3309
Vucic D, Kaiser WJ, Harvey AJ and Miller LK . 1997 Inhibition of Reaper-induced apoptosis by interaction with inhibitor of apoptosis proteins (IAPs). PNAS, USA 94: 10183–10188
Wing J, Zhou L, Schwartz L and Nambu J . 1998 Distinct killing properties of the Drosophila reaper, head involution defective, and grim genes. Cell Death Differ. 5: 930–939
Chen P, Lee P, Otto L and Abrams JM . 1996 Apoptotic activity of REAPER is distinct from signalling by the Tumor Necrosis Factor Receptor 1 death domain. J. Biol. Chem. 271: 25735–25737
Vucic D, Seshagiri S and Miller LK . 1997 Characterization of reaper and FADD-induced apoptosis in a lepidopteran cell line. Mol. Cell. Biol. 17: 667–676
Du C, Fang M, Li Y, Li L and Wang X . 2000 Smac, a mitochondrial protein that promotes cytochrome c-dependent caspase activation by eliminating IAP inhibition. Cell 102: 33–42
Verhagen A, Ekert P, Pakusch M, Silke J, Connolly L, Reid G, Moritz R, Simpson R and Vaux D . 2000 Indentification of DIABLO, a mammalian protein that promotes apoptosis by binding to and antagonizing IAP proteins. Cell 102: 43–53
Inohara N, Koseki T, Hu Y, Chen S and Nunez G . 1997 CLARP, a death effector domain-containing protein interacts with caspase-8 and regulates apoptosis. Proc. Natl. Acad. Sci. USA 94: 10717–10722
Chen P, Rodriguez A, Erskine R, Thach T and Abrams J . 1998 Dredd, a novel effector of the apoptosis activators reaper, grim, and hid in Drosophila. Dev. Biol. 201: 202–216
Dorstyn L, Colussi P, Quinn L, Richardson H and Kumar S . 1999 DRONC, an ecdysone-inducible Drosophila caspase. PNAS 96: 4307–4312
Song Z, McCall K and Steller H . 1997 DCP-1, a Drosophila cell death protease essential for development. Science 275: 536–540
Fraser A and Evan G . 1997 Identification of a Drosophila melanogaster ICE/CED-3-related protease, drICE. EMBO J. 16: 2805–2813
Dorstyn L, Read S, Quinn L, Richardson H and Kumar S . 1999 DECAY, a novel Drosophila caspase related to mammalian caspase-3 and caspase-7. J. Biol. Chem. 274: 30778–30783
Rubin GM, Yandell MD, Wortman JR, Gabor Miklos GL, Nelson CR, Hariharan IK, Fortini ME, Li PW, Apweiler R, Fleischmann W, Cherry JM, Henikoff S, Skupski MP, Misra S, Ashburner M, Birney E, Boguski MS, Brody T, Brokstein P, Celniker SE, Chervitz SA, Coates D, Cravchik A, Gabrielian A, Galle RF, Gelbart WM, George RA, Goldstein LS, Gong F, Guan P, Harris NL, Hay BA, Hoskins RA, Li J, Li Z, Hynes RO, Jones SJ, Kuehl PM, Lemaitre B, Littleton JT, Morrison DK, Mungall C, O'Farrell PH, Pickeral OK, Shue C, Vosshal LB, Zhang J, Zhao Q, Zheng XH, Zhong F, Zhong W, Gibbs R, Venter JC, Adams MD and Lewis S . 2000 Comparative genomics of the eukaryotes. Science 287: 2204–2215
Hawkins CJ, Yoo SJ, Peterson EP, Wang SL, Vernooy SY and Hay BA . 2000 The Drosophila caspase DRONC is a glutamate/aspartate protease whose activity is regulated by DIAP1, HID and GRIM. J. Biol. Chem. 275: 27084–27093
Rodriguez A, Oliver H, Zou H, Chen P, Wang X and Abrams J . 1999 DARK, a Drosophila homolog of Apaf-1/ced-4, functions in an evolutionarily conserved death pathway. Nature Cell Biol. 1: 272–279
Song ZW, Guan B, Bergman A, Nicholson DW, Thornberry NA, Peterson EP and Steller H . 2000 Biochemical and genetic interactions between Drosophila caspases and the proapoptotic genes rpr, hid, and grim. Mol. Cell. Biol. 20: 2907–2914
McCall K and Steller H . 1998 Requirement for DCP-1 caspase during Drosophila oogenesis. Science 279: 230–234
Kanuka H, Sawamoto K, Inohara N, Matsuno K, Okano H and Miura M . 1999 Control of the cell death pathway by Dapaf-1, a Drosophila Apaf-1/CED-4-related caspase activator. Mol. Cell. 4: 757–769
Zhou L, Song Z, Tittel J and Steller H . 1999 HAC-1, a Drosophila homolog of APAF-1 and CED-4, functions in developmental and radiation-induced apoptosis. Mol. Cell. 4: 745–755
Brachmann CB, Jassim OW, Wachsmuth BD and Cagan RL . 2000 The Drosophila bcl-2 family member dBorg-1 functions in the apoptotic response to UV-irradiation. Curr. Biol. 10: 547–550
Igaki T, Kanuka H, Inohara N, Sawamoto K, Nunez G, Okano H and Miura M . 2000 Drob-1, a Drosophila member of the Bcl-2/CED-9 family that promotes cell death. Proc. Natl. Acad. Sci. USA 97: 662–667
Colussi PA, Quinn LM, Huang DC, Coombe M, Read SH, Richardson H and Kumar S . 2000 Debcl, a proapoptotic Bcl-2 homologue, is a component of the Drosophila melanogaster cell death machinery. J. Cell. Biol. 148: 703–714
Flybase . 1994 The Drosophila genetic data base. Nucleic Acids. Res. 22: 3456–3458
Vaux D . 1997 ced-4- The third horseman of apoptosis. Cell 90: 389–390
Thress K, Henzel W, Shillinglaw W and Kornbluth S . 1998 Scythe: a novel reaper-binding apoptotic regulator. EMBO J. 17: 6135–6143
Thress K, Evans EK and Kornbluth S . 1999 Reaper-induced dissociation of a Scythe-sequestered cytochrome C- releasing activity. EMBO J. 18: 5486–5493
Gross A, McDonnell JM and Korsmeyer SJ . 1999 BCL-2 family members and the mitochondria in apoptosis. Genes Dev. 13: 1899–1911
Varkey J, Chen P, Jemmerson R and Abrams JM . 1999 Altered cytochrome C display precedes apoptotic cell death in Drosophila. J. Cell. Biol. 144: 701–710
Wyllie AH, Kerr JFR and Currie AR . 1980 Cell death: the significance of apoptosis. Int. Rev. Cytol. 68: 251–306
Raff M . 1992 Social controls on cell survival and cell death. Nature 356: 397–400
Brown EJ . 1995 Phagocytosis. Bioessays 17: 109–117
Ellis R, Jacobson D and Horvitz H . 1991 Genes required for the engulfment of cell corpses during programmed cell death in Caenorhabditis elegans. Genetics 129: 79–94
Sonnenfeld MJ and Jacobs JR . 1995 Macrophages and glia participate in the removal of apoptotic neurons from the Drosophila embryonic nervous system. J. Comp. Neurol. 359: 644–652
Tepass U, Fessler LI, Aziz A and Hartenstein V . 1994 Embryonic origin of hemocytes and their relationship to cell death in Drosophila. Development 120: 1829–1837
Lebestky T, Chang T, Hartenstein V and Banerjee U . 2000 Specification of Drosophila hematopoietic lineage by conserved transcription factors. Science 288: 146–149
Rizki T and Rizki R . 1984 In: Insect Ultrastructure. King R, Akai H (eds) pp. 579–604 (Plenum Press: New York)
Rehorn KP, Thelen H, Michelson AM and Reuter R . 1996 A molecular aspect of hematopoiesis and endoderm development common to vertebrates and Drosophila. Development 122: 4023–4031
Bernardoni R, Vivancos B and Giangrande A . 1997 glide/gcm is expressed and required in the scavenger cell lineage. Dev. Biol. 191: 118–130
Nelson R, Fessler L, Takagi Y, Blumberg B, Keene D, Olson P, Parker C and Fessler J . 1994 Peroxidasin: a novel enzyme-matrix protein of Drosophila development. EMBO J. 13: 3438–3447
Diez-Roux G and Lang RA . 1997 Macrophages induce apoptosis in normal cells in vivo. Development 124: 3633–3638
Savill J . 1997 Recognition and phagocytosis of cells undergoing apoptosis. Br. Med. Bull. 53: 491–508
Franc NC, Dimarcq JL, Lagueux M, Hoffmann J and Ezekowitz RA . 1996 Croquemort, a novel Drosophila hemocyte/macrophage receptor that recognizes apoptotic cells. Immunity 4: 431–443
Franc N, Heitzler P, Ezekowitz R and White K . 1999 Requirement for croquemort in phagocytosis of apoptotic cells in Drosophila. Science 284: 1994–1998
Ren Y, Silverstein R, Allen J and Savill J . 1995 CD36 gene transfer confers capacity for phagocytosis of cells undergoing apoptosis. J. Exp. Med. 181: 1857–1862
Savill J, Hogg N, Ren Y and Haslett C . 1992 Thrombospondin cooperates with CD36 and the vitronectin receptor in macrophage recognition of neutrophils undergoing apoptosis. J. Clin. Invest. 90: 1513–1522
Fadok VA, Bratton DL, Rose DM, Pearson A, Ezekewitz RA and Henson PM . 2000 A receptor for phosphatidylserine-specific clearance of apoptotic cells. Nature 405: 85–90
Pearson A, Lux A and Krieger M . 1995 Expression cloning of dSR-C1, a class C macrophage-specific scavenger receptor from Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 92: 4056–4060
van den Eijnde S, Boshart L, Baehrecke E, De Zeeuw C, Reutelingsperger C and Vermeij-Keers C . 1998 Cell surface exposure of phosphatidylserine during apoptosis is phylogenetically conserved. Apoptosis 3: 9–16
Suzuki H, Kurihara Y, Takeya M, Kamada N, Kataoka M, Jishage K, Ueda O, Sakaguchi H, Higashi T, Suzuki T, Takashima Y, Kawabe Y, Cynshi O, Wada Y, Honda M, Kurihara H, Aburatani H, Doi T, Matsumoto A, Azuma S, Noda T, Toyoda Y, Itakura H, Yazaki Y, Kodama T et al.1997 A role for macrophage scavenger receptors in atherosclerosis and susceptibility to infection. Nature 386: 292–296
Platt N, Suzuki H, Kurihara Y, Kodama T and Gordon S . 1996 Role for the class A macrophage scavenger receptor in the phagocytosis of apoptotic thymocytes in vitro. Proc. Natl. Acad. Sci. USA 93: 12456–12460
Fukasawa M, Adachi H, Hirota K, Tsujimoto M, Arai H and Inoue K . 1996 SRB1, a class B scavenger receptor, recognizes negatively charged liposomes and apoptotic cells. Exp. Cell. Res. 222: 246–250
Asano M, Nevins JR and Wharton RP . 1996 Ectopic E2F expression induces S phase and apoptosis in Drosophila imaginal discs. Genes Dev. 10: 1422–1432
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by S Kumar
Rights and permissions
About this article
Cite this article
Bangs, P., Franc, N. & White, K. Molecular mechanisms of cell death and phagocytosis in Drosophila. Cell Death Differ 7, 1027–1034 (2000). https://doi.org/10.1038/sj.cdd.4400754
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4400754
Keywords
This article is cited by
-
Active JNK-dependent secretion of Drosophila Tyrosyl-tRNA synthetase by loser cells recruits haemocytes during cell competition
Nature Communications (2015)
-
The mechanism of peptide-binding specificity of IAP BIR domains
Cell Death & Differentiation (2008)
-
Relationship among follicular apoptosis, integrin β1 and collagen type IV during early ovarian regression in the teleost Prochilodus argenteus after induced spawning
Cell and Tissue Research (2008)
-
Illuminating the role of caspases during Drosophila oogenesis
Cell Death & Differentiation (2006)
-
IAP proteins: blocking the road to death's door
Nature Reviews Molecular Cell Biology (2002)