Abstract
Helicobacter pylori, strain 26695, has a circular genome of 1,667,867 base pairs and 1,590 predicted coding sequences. Sequence analysis indicates that H. pylori has well-developed systems for motility, for scavenging iron, and for DNA restriction and modification. Many putative adhesins, lipoproteins and other outer membrane proteins were identified, underscoring the potential complexity of host–pathogen interaction. Based on the large number of sequence-related genes encoding outer membrane proteins and the presence of homopolymeric tracts and dinucleotide repeats in coding sequences, H. pylori, like several other mucosal pathogens, probably uses recombination and slipped-strand mispairing within repeats as mechanisms for antigenic variation and adaptive evolution. Consistent with its restricted niche, H. pylori has a few regulatory networks, and a limited metabolic repertoire and biosynthetic capacity. Its survival in acid conditions depends, in part, on its ability to establish a positive inside-membrane potential in low pH.
Similar content being viewed by others
Main
For most of this century the cause of peptic ulcer disease was thought to be stress-related and the disease to be prevalent in hyperacid producers. The discovery1 that Helicobacter pylori was associated with gastric inflammation and peptic ulcer disease was initially met with scepticism. However, this discovery and subsequent studies on H. pylori have revolutionized our view of the gastric environment, the diseases associated with it, and the appropriate treatment regimens2.
Helicobacter pylori is a micro-aerophilic, Gram-negative, slow-growing, spiral-shaped and flagellated organism. Its most characteristic enzyme is a potent multisubunit urease3 that is crucial for its survival at acidic pH and for its successful colonization of the gastric environment, a site that few other microbes can colonize2. H. pylori is probably the most common chronic bacterial infection of humans, present in almost half of the world population2. The presence of the bacterium in the gastric mucosa is associated with chronic active gastritis and is implicated in more severe gastric diseases, including chronic atrophic gastritis (a precursor of gastric carcinomas), peptic ulceration and mucosa-associated lymphoid tissue lymphomas2. Disease outcome depends on many factors, including bacterial genotype, and host physiology, genotype and dietary habits4,5. H. pylori infection has also been associated with persistent diarrhoea and increased susceptibility to other infectious diseases6.
Because of its importance as a human pathogen, our interest in its biology and evolution, and the value of complete genome sequence information for drug discovery and vaccine development, we have sequenced the genome of a representative H.pylori strain by the whole-genome random sequencing method as described for Haemophilus influenzae7, Mycoplasma genitalium8 and Methanococcus jannaschii9.
General features of the genome
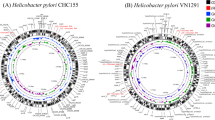
Genome analysis. The genome of H. pylori strain 26695 consists of a circular chromosome with a size of 1,667,867 base pairs (bp) and average G + C content of 39% (Fig 1 (PDF File: 1751k) and 2). Five regions within the genome have a significantly different G + C composition (Table 1 and Fig. 1 (PDF File: 1751k)). Two of them contain one or more copies of the insertion sequence IS605 (see below) and are flanked by a 5S ribosomal RNA sequence at one end and a 521 bp repeat (repeat 7) near the other. These two regions are also notable because they contain genes involved in DNA processing and one contains 2 orthologues of the virB4/ptl gene, the product of which is required for the transfer of oncogenic T-DNA of Agrobacterium and the secretion of the pertussis toxin by Bordetella pertussis10. Another region is the cag pathogenicity island (PAI), which is flanked by 31-bp direct repeats, and appears to be the product of lateral transfer11.
Outer concentric circle: predicted coding regions on the plus strand classified as to role according to the colour code in Fig. 1 (PDF File: 1751k) (except for unknowns and hypotheticals, which are in black). Second concentric circle: predicted coding regions on the minus strand. Third and fourth concentric circles: IS elements (red) and other repeats (green) on the plus and minus strand, respectively. Fifth and sixth concentric circles: tRNAs (blue), rRNAs (red), and sRNAs (green) on the plus and minus strand, respectively.
RNA and repeat elements. Thirty-six tRNA species were identified using tRNAscan-SE12. These are organized into 7 clusters plus 12 single genes. Two separate sets of 23S–5S and 16S ribosomal RNA (rRNA) genes were identified, along with one orphan 5S gene and one structural RNA gene (Table 1). Associated with each of the two 23S–5S gene clusters is a 6-kilobase (kb) repeat containing a possible operon of 5 ORFs that have no database matches.
Eight repeat families (>97% identity) varying in length from 0.47 to 3.8 kb were found in the chromosome (Figs 1 (PDF File: 1751k) and 2). Members of repeat 7 are found in intergenic regions, while the others are associated with coding sequences and may represent gene duplications. Repeats 1, 2, 3 and 6 are associated with genes that encode outer-membrane proteins (OMP) (Fig. 3).
Symbols are as follows: ++, Co2+, Zn2+, Cd2+; ?, unknown; A/G/S, D-alanine/glycine/D-serine; B12, B12/ferric siderophores; E, glutamate; Mo, molybdenum; P, proline; P/G, proline/glycine betaine; Q, glutamine; S, serine;a-k, α-ketoglutarate; a/o, arginine/ornithine; aa, amino acids (specificity unknown); aa2, dipeptides; aaX, oligopeptides; fum, fumarate, succinate; glu, glucose/galactose; h, hemin; lac, L-lactate; mal, malate 2-oxoglutarate; nic, nicotinamide mononucleotides; pyr, pyrimidine nucleosides. Numbers associated with tRNA symbols represent the number of tRNAs at a locus. Numbers associated with GES represent the number of membrane-spanning domains according to the Goldman, Engelman and Steitz scale as calculated by TopPred47.
These proteins were identified as OMPs based on the characteristic alternating hydrophobic residues at their carboxy termini. All members of this family have one domain of similarity at the amino-terminal end and seven domains of similarity at their carboxy-terminal end. Note that the first 11 of these OMPs share extensive similarity over their entire length. Four of the OMPs were identified as porins (Hops) based on identity to published amino-terminal sequences, represented at the top of the alignment50. The most likely candidate for HopD is HP0913, which has 15 matches to the first 20-residue N-terminal peptide sequence50. These differences may be due to strain variability. The program Signal-P48 was used to identify cleavage sites and signal peptides (underlined). Four of the OMPs have TTG start codons (HP1156, HP0252, HP1113, HP0796). Numbers embedded in the sequences represent amino acids omitted from the alignment. The star symbols indicate that HP722, HP725 and HP9 proteins contain a frameshift in their signal-peptide-coding region. These frameshifts are associated with the presence of dinucleotide repeats (Table 3).
Two distinct insertion sequence (IS) elements are present. There are five full-length copies of the previously described IS60511,13 and two of a newly discovered element designated IS606. In addition, there are eight partial copies of IS605 and two partial copies of IS606. Both elements encode two divergently transcribed transposases (TnpA and TnpB). IS606 has less than 50% nucleotide identity with IS605 and the IS606 transposases have 29% amino-acid identity with their IS605 counterpart. Both copies of the IS606 TnpB may be non-functional owing to frameshifts.
Origin of replication.As a typical eubacterial origin of replication was not identified14, we arbitrarily designated basepair one at the start of a 7-mer repeat, (AGTGATT)26, that produces translational stops in all reading frames, as this repeated DNA is unlikely to contain any coding sequence.
Open reading frames.One thousand five hundred and ninety predicted coding sequences were identified. They were searched against a non-redundant protein database resulting in 1,091 putative identifications that were assigned biological roles using a classification system adapted from Riley15 (Table 2 (PDF File: 169k)). The 1,590 predicted genes had an average size of 945 bp, similar to that observed in other prokaryotes7,8,9, and no genome-wide strand bias was observed (Fig. 2). More than 70% of the predicted proteins in H. pylori have a calculated isoelectric point (pI) greater than 7.0, compared to ∼40% in H. influenzae and E. coli. The basic amino acids, arginine and lysine, occur twice as frequently in H. pylori proteins as in those of H. influenzae and E. coli, perhaps reflecting an adaptation of H. pylori to gastric acidity.
Paralagous families.Ninety-five paralogous gene families comprising 266 gene products (16% of the total) were identified (>www.tigr.org/tdb/mdb/hpdb/hpdb.html). Of these, 67 (173 proteins) have an assigned role. Sixty-four have only 2 members, while the porin/adhesin-like outer membrane protein family (Fig.2) is the largest with 32 members. The largest number of paralogues with assigned roles fall into the functional categories of cell envelope, transport and binding proteins, and proteins involved in replication. The large number of cell envelope proteins might reflect either a reduced biosynthetic capacity or a need to adapt to the challenging gastric environment.
Cell division and protein secretion
The gene content of H. pylori suggests that the basic mechanisms of replication, cell division and secretion are similar to those of E. coli and H. influenzae. However, important differences are noted. For example, apparently missing from the H. pylori genome are orthologues of DnaC, MinC, and the secretory chaperonin, SecB. In oriC-type primosome formation, the DnaB and DnaC proteins form a B–C complex that delivers the DnaB helicase to the developing primosome complex16. The apparent absence of DnaC in H. pylori suggests that either a novel mechanism for recruiting DnaB exists or a DnaC orthologue with no detectable sequence similarity is present. Similar arguments can be made for other seemingly missing important functions.
H. pylori has a classical set of bacterial chaperones (DnaK, DnaJ, CbpA, GrpE, GroEL, GroES, and HtpG). The transcriptional regulation of H. pylori chaperone genes is likely to be different from that in E. coli, as it seems not to have the sigma factors that upregulate chaperone synthesis in E. coli (heat-shock sigma 32 and stationary-phase sigma S).
In addition to the SecA-dependent secretory pathway, H. pylori has two specialized export systems. One is associated with the cag pathogenicity island11 and the other is the flagellar export pathway which is assembled from orthologues of FliH, FliI, FliP, FlhA, FlhB, FliQ, FliR and FliP17. Apparently absent from H. pylori is a type IV signal peptidase and orthologues of the dsbABC system, which in other species are required for the maturation of pili and pilin-like structures18 and assembly of surface structures involved in virulence and DNA transformation19.
Recombination, repair and restriction systems
Systems for homologous recombination and post-replication, mismatch, excision and transcription-coupled repair appear to be present in H. pylori. Also present are genes with similarity to DNA glycosylases which have associated AP endonuclease activity. The RecBCD pathway, which mediates homologous recombination and double-strand break repair, and RecT and RecE orthologues, proteins involved in strand exchange during recombination20, seem to be absent. The ability of H. pylori to perform mismatch repair is suggested by the presence of methyl transferases, mutS and uvrD. However, orthologues of MutH and MutL were not identified. Components of an SOS system also appear to be absent.
Bacteria commonly use restriction and modification systems to degrade foreign DNA. In H. pylori, this defence system is well developed with eleven restriction-modification systems identified on the basis of gene order and similarity to endonucleases, methyltransferases, and specificity subunits. Three type I, one type II, and three type IIS systems were identified, as well as four type III systems, including the recently identified epithelial responsive endonuclease, iceA1, and its associated DNA adenine methyltransferase (M. HypI) genes21,22. In addition to the complete systems, seven adenine-specific, and four cytosine-specific methyltransferases, and one of unknown specificity were found. Each of these has an adjacent gene with no database match, suggesting that they may function as part of restriction-modification systems.
Transcription and translation
Although analysis of gene content suggests that H. pylori has a basic transcriptional and translational machinery similar to that of E. coli, interesting differences are observed. For example, no genes for a catalytic activity in tRNA maturation (rnd, rph, or rnpB) were identified and of the three known ribonucleases involved in mRNA degradation, only polyribonucleotide phosphorylase was found. Twenty-one genes coding for 18 of the 20 tRNA synthetases normally required for protein biosynthesis were found.
As in most other completely sequenced bacterial genomes, the gene for glutaminyl-tRNA synthetase, glnS, is missing, and the existence of a transamidation process is assumed. It is also possible that the product of the second glutamyl-tRNA synthetase gene, gltX, present in H. pylori, may have acquired the glutaminyl-tRNA synthetase function. H. pylori provides the first example of a bacterial genome apparently lacking an asparaginyl-tRNA synthetase gene, asnS. A transamidation process to form Asn-tRNAAsn from Asp-tRNAAsn has been reported for the archaeon Haloferax volcanii22 and may also operate in H. pylori. Most intriguing, however, is the finding that in H. pylori the genes encoding the β and β′ subunits of RNA polymerase are fused. In all studied prokaryotes the two genes are contiguous, but separate, and are part of the same transcriptional unit. Whether this gene fusion in H. pylori results in a fused protein, or whether the transcriptional or translational product of the fusion is subject to splicing, is currently not known. It is worth noting that an artificial fusion of the E. coli rpoB and rpoC genes is viable and results in a transcriptional complex, which has the same stoichiometry as the native complex (K. Severinov, personal communication).
Adhesion and adaptive antigenic variation
Most pathogens show tropism to specific tissues or cell types and often use several adherence mechanisms for successful attachment. H. pylori may use at least five different adhesins to attach to gastric epithelial cells5. One of them, HpaA (HP0797), was previously identified as a lipoprotein in the flagellar sheath and outer membrane5,23. In addition to the HpaA orthologue, we have identified 19 other lipoproteins. Few have an identifiable function, but some are likely to contribute to the adherence capacity of the organism.
Two adhesins24,25,26, one of which mediates attachment to the Lewisbhisto-blood group antigens, belong to the large family of outer membrane proteins (OMP) (Fig. 3) (T. Boren and R. Haas, personal communication). It is conceivable that other members of these closely related proteins also act as adhesins. Given the large number of sequence-related genes encoding putative surface-exposed proteins, the potential exists for recombinational events leading to mosaic organization. This could be the basis for antigenic variation in H. pylori and an effective mechanism for host defence evasion, as seen in M. genitalium27.
At least one other mechanism for antigenic variation could operate in H. pylori. The DNA sequence at the beginning of eight genes, including five members of the OMP family, contain stretches of CT or AG dinucleotide repeats (Table 3a). In addition, poly(C) or poly(G) tracts occur within the coding sequence of nine other genes (Table 3b>). Slipped-strand mispairing within such repeats are documented features of one mechanism of genotypic variation28,29. These mechanisms may have evolved in bacterial pathogens to increase the frequency of phenotypic variation in genes involved in critical interactions with their hosts28. Such ‘contingency’ genes encode surface structures like pilins, lipoproteins or enzymes that produce lipopolysaccharide molecules28. Our analysis suggests that the seventeen genes reported in Table 3a, b belong to this category and thus may provide an example of adaptive evolution in H. pylori.
Phenotypic variation at the transcriptional level may also operate in H. pylori. Examples of repetitive DNA mediating transcriptional control have been documented by the presence of oligonucleotide repeats in promoter regions29. Homopolymeric tracts of A or T in potential promoter regions of eighteen genes were found, including eight members of the OMP family (Table 3c).
Virulence
The virulence of individual H. pylori isolates has been measured by their ability to produce a cytotoxin-associated protein (CagA) and an active vacuolating cytotoxin (VacA)5. The cagA gene, though not a virulence determinant, is positioned at one end of a pathogenecity island containing genes that elicit the production of interleukin (IL)-8 by gastric epithelial cells11,30. Consistent with its more virulent character, H. pylori strain 26695 contains a single contiguous PAI region11 (Fig. 4).
The twenty nine ORFs of the contiguous PAI in strain 26695 are represented together with the corresponding ORFs from the PAI present in NCTC11638 (AC000108 and U60176). The PAI in NCTC11638 is divided by the IS 605 elements into two regions, cagI and cagII. The PAI in NCTC11638 is flanked by a 31-bp (TTACAATTTGAGCCCATTCTTTAGCTTGTTTT) direct repeat (vertical arrows) as described11. Some of the genes encode proteins with similarity to proteins involved either in DNA transfer (Vir and Tra proteins) or in export of a toxin (Ptl protein)10. However, these genes do not have the conserved contiguous arrangement found in the VirB, Tra and Ptl operons, suggesting that this PAI is not derived from these systems. Most genes of the PAI have no database match, contrary to a previous suggestion11. Thirteen of the proteins have a signal peptide (squiggle line), three of them with a weaker probability (squiggled line+?). The average length of the signal peptides is 25 amino acids, suggesting that this PAI is of Gram-negative origin. Eight proteins are predicted to have at least two membrane-spanning domains and to be integral membrane proteins (IM)47. Although the two PAI are ∼97% identical at the nucleotide level, there are several notable and perhaps biologically relevant differences between the two sequences. Four of the genes differ in size. In the PAI of strain 26695, HP 520 and 521 are shorter, whereas HP523 is longer, and HP 527 actually spans both ORF13 and 14. In addition, the N-terminal part of HP527 is 129 amino acids longer than the corresponding region in ORF14. HP548/549 contains a frameshift and is therefore probably inactive in strain 26695. The stippled box preceding ORF13 represents an N-terminal extension not annotated in the Genbank entry for the PAI of NCTC11638. The ‘x’ indicates ORFs that are neither GeneMark-positive nor GeneSmith-positive, so were not included in our gene list. However, these ORFs may be biologically significant. We do not represent cagR as an ORF, because it is completely contained within ORFQ, and is GeneMark-negative.
VacA induces the formation of acidic vacuoles in host epithelial cells, and its presence is associated epidemiologically with tissue damage and disease31. VacA may not be the only ulcer-causing factor as 40% of H. pylori strains do not produce detectable amounts of the cytotoxin in vitro5. Sequence differences at the amino terminus and central sections are noted among VacA proteins derived from Tox+ and Tox− strains31. This Tox+H. pylori strain contains the more toxigenic S1a/m1 type cytotoxin and three additional large proteins with moderate similarities to the carboxy-terminal end of the active cytotoxin (∼26–31%) (Fig. 5). However, they lack the paired-cysteine residues and the cleavage site required for release of the VacA toxin from the bacterial membrane31 (Fig. 5). We propose that these proteins may be retained on the outside surface of the cell membrane and contribute to the interaction between H. pylori and host cells.
HP887 is the vacuolating cytotoxin (vacA) gene from H. pylori 26695 strain. HP610, HP922 and HP289 are related proteins. Blocks of aligned sequence and the length of each protein are shown. Arrows designate the extents of each VacA domain. The hydrophilic domain (blue boxes) contains the site in VacA at which the N-terminal domain is cleaved into 37K and 58K fragments. The putative cleavage site (ANNNQQNS) differs from that of three cytotoxic strains (CCUG 1784, 60190, G39; AKNDKXES) and is not conserved in the other three VacA-related proteins. The cleavage domain (black boxes) of VacA contains a pair of Cys residues 60 residues upstream from the site at which the C terminus is cleaved. These residues are not conserved in the other three proteins. The 33K C-terminal hydrophobic domain (red boxes) in VacA is thought to form a pore through which the toxin is secreted. The other three proteins show 26–31% sequence similarity to VacA in this region. The other coloured boxes represent regions of similarity.
The surface-exposed lipopolysaccharide (LPS) molecule plays an important role in H. pylori pathogenesis32. The LPS of H. pylori is several orders of magnitude less immunogenic than that of enteric bacteria33 and the O antigen of many H. pylori isolates is known to mimic the human Lewisx and Lewisy blood group antigen32. Genes for synthesis of the lipid A molecule, the core region, and the O antigen were identified. Two genes with low similarity to fucosyltransferases (HP379, HP651) were found and may play a role in the LPS-Lewis antigen molecular mimicry. Our analysis also suggests that three genes, two glycosyltransferases (HP208 and HP619) and one fucosyltransferase (HP379), may be subject to phase variation (Table 3a, b).
As with other pathogens, H. pylori probably requires an iron-scavenging system for survival in the host5. Genome analysis suggests that H. pylori has several systems for iron uptake. One is analogous to the siderophore-mediated iron-uptake fec system of E. coli34, except that it lacks the two regulatory proteins (FecR and FecI) and is not organized in a single operon. Unlike other studied systems, H. pylori has three copies of each of fecA, exbB and exbD. A second system, consisting of a feoB-like gene without feoA, suggests that H. pylori can assimilate ferrous iron in a fashion similar to the anaerobic feo system of E. coli. Other systems for iron uptake present in H. pylori consist of the three frpB genes which encode proteins similar to either haem- or lactoferrin-binding proteins. Finally, H. pylori contains NapA, a bacterioferritin34, and Pfr, a non-haem cytoplasmic iron-containing ferritin used for storage of iron35. The global ferric uptake regulator (Fur) characterized in other bacteria is also present in H. pylori. Consensus sequences for Fur-binding boxes were found upstream of two fecA genes, the three frnB genes and fur.
H. pylori motility is essential for colonization36. It enables the bacterium to spread into the viscous mucous layer covering the gastric epithelium. At least forty proteins in the H. pylori genome appear to be involved in the regulation, secretion and assembly of the flagellar architecture. As has bene reported for the flaA and flaB genes, we identified sigma 28 and sigma 54-like promoter elements upstream of many flagellar genes, underscoring the complexity of the transcriptional regulation of the flagellar regulon5.
Acidity, pH and acid tolerance
H. pylori is unusual among pathogenic bacteria in its ability to colonize host cells in an environment of high acidity. As it enters the gastric environment by oral ingestion, the organism is transiently subjected to the extreme pH of the lumen side of the gastric mucous layer (pH ∼2). The survival of H. pylori in acidic environments is probably due to its ability to establish a positive inside-membrane potential37 and subsequently to modify its microenvironment through the action of urease and the release of factors that inhibit acid production by parietal cells5. A switch in membrane polarity provides an electrical barrier that prevents the entry of protons (H+). A positive cell interior can be created by the active extrusion of anions or by a proton diffusion potential. The latter model appears more likely as no clear mechanism for electrogenic anion efflux is apparent in the genome. A proton diffusion potential would require the anion permeability of the cytoplasmic membrane to be low and, thus far, only three anion transporters have been identified. However, it remains to be determined whether anion conductances are associated with other proteins: the MDR-like transporters (HP600, HP1082 and HP1206) or hypotheticals. Although it has been suggested that proton-translocating P-type ATPases could mediate survival in acid conditions by the extrusion of protons from the cytoplasm38, this idea is not supported by the identified transporter genes. The P-type ATPase sequences in H. pylori (copAP, HP791, and HP1503) are more closely related to divalent cation transporters than to ATPases with specificity for protons or monovalent cations. One of them, HP0791, is involved in Ni2+ supply, an essential component of urease activity39. The others may be involved in the elimination of toxic metals from the cytoplasm and not in pH regulation.
Additional mechanisms of pH homeostasis may well contribute to H. pylori survival. A change in protein content observed in response to a shift of extracellular pH from 7.5 to 3.0 suggests the presence of an acid-inducible response40. Although H. pylori lacks most orthologues of the genes that are acid-induced in E. coli and Salmonella typhimurium, including the amino-acid decarboxylases and formate hydrogen lyase, certain virulence factors, outer membrane proteins, sensor-regulator pairs and other proteins may be acid-induced.
Regulation of gene expression
Bacteria regulate the transcription of their genes in response to many environmental stimuli, such as nutrient availability, cell density, pH, contact with target tissue, DNA-damaging agents, temperature and osmolarity. In the case of pathogens, the regulated expression of certain key genes is essential for successful evasion of host responses and colonization, adaptation to different body sites, and survival as the pathogen passes to new hosts. In H. pylori, global regulatory proteins are less abundant than in E. coli. For example, orthologues of many DNA-binding proteins that regulate the expression of certain operons such as OxyR (oxidative stress), Crp (carbon utilization), RpoH (heat shock), and Fnr (fumarate and nitrate regulation) are absent. Only four H. pylori proteins have a perfect match to helix–turn–helix (HTH) motifs, a signature of transcription factors; a putative heat-shock protein (HspR), two proteins with no database match (HP1124 and HP1349) and SecA, a component of the general secretory machinery. In contrast, 34 proteins containing an HTH motif were found in H. influenzae and 148 in E. coli. We identified several other putative regulatory functions, including SpoT and CstA for ‘stringent response’ to amino-acid starvation and to carbon starvation, respectively.
Environmental response requires sensing changes and transmission of this information to cellular regulatory networks. Two-component regulator systems, consisting of a membrane histidine kinase sensor protein and a cytoplasmic DNA-binding response regulator, provide a well studied mechanism for such signal transduction. Four sensor proteins and seven response regulators were found in H. pylori, similar to the number found in H. influenzae7. This is approximately one third the number found in E. coli which, in contrast to H. pylori and H. influenzae, may be exposed to more environments.
Metabolism
Metabolic pathway analysis of the H. pylori genome suggests the following features. H. pylori uses glucose as the only source of carbohydrate and the main source for substrate-level phosphorylation. It also derives energy from the degradation of serine, alanine, aspartate and proline. The glycolysis–gluconeogenesis metabolic axis constitutes the backbone of energy production and the start point of many biosynthetic pathways. The biosynthesis of peptidoglycan, phospholipids, aromatic amino acids, fatty acids and cofactors is derived from acetyl-CoA or from intermediates in the glycolytic pathway (Fig. 6). The metabolism of pyruvate reflects the microaerophilic character of this organism. Neither the aerobic pyruvate dehydrogenase (aceEF) nor the strictly anaerobic pyruvate formate lyase (pfl) associated with mixed-acid fermentation are present. The conversion of pyruvate to acetyl CoA is performed by the pyruvate ferrodoxin oxidoreductase (POR), a four-subunit enzyme thus far only described in hyperthermophilic organisms41. The tricarboxylic acid cycle (TCA) is incomplete and the glyoxylate shunt is absent. The analysis of degradative pathways, uptake systems and biosynthetic pathways for pyrimidine, purine and haem suggests that H. pylori uses several substrates as nitrogen source, including urea, ammonia, alanine, serine and glutamine. The assimilation of ammonia, an abundant product of urease activity, is achieved by the glutamine synthase enzyme and α-ketoglutarate is transformed into glutamate by glutamate dehydrogenase rather than by the glutamate synthase enzyme.
Transporters identified by sequence comparisons are characteristc of Gram-negative bacteria. Colours correspond to transport role categories defined by Riley15: blue, amino acids, peptides and amines; red, anions; yellow, carbohydrates, organic alcohols and acids; green, cations; and purple, nucleosides, purines and pyrimidines. Numerous permeases (ovals) with specificity for amino acids (recE, proP, dagA, gltS, putP and sdaC) or carbohydrates (SODiTl, gluP, lactP, cduA, kgtP) import organic nutrients. Structurally related permease proteins maintain ionic homeostasis by transporting HPO42− (HI1604), NO32− (narK), and Na+ (nhA, napA). Primary active-transport systems, independent of the proton cycle, are also apparent. Included in this group are ATP-binding protein-cassette (ABC) transporters (composite figures of 2 diamonds, 2 circles, 1 oval)forthe uptake of oligopeptides (oppACD), dipeptides (dppABCDF), proline (proVWX), glutamine (glnHMPQ), molybdenum (modABD), and iron III (fecED), P-type ATPases that extrude toxic metals from the cell (copAP and cadA), and the glutathione-regulated potassium-efflux protein (kefB). Transporters for the accumulation of ionic cofactors are encoded by nixA (Ni2+ for urease activation), corA (Mg2+ for phosphohydrolases, phosphotransferases, ATPases) and feoB (Fe2+ import under anaerobic conditions for cytochromes, catalase). An integrated view of the main components of the central metabolism of H. pylori strain 26695 is presented. The use of glucose as the sole carbohydrate source is emphasized. Urease, a multisubunit Ni2+-binding enzyme, is crucial for colonization and for survival of H. pylori at acid pH, and is indicated as a complex (purple circle) with Hpn, a Ni2+-binding cofactor, and a newly identified Hpn-like protein (HP1432). A question mark is attached to pathways that could not be completely elucidated. Pathways or steps for which no enzymes were identified are represented by a redarrow. Pathways for macromolecular biosynthesis (RNA, DNA and fatty acids)have been omitted. ackA, acetate kinase; acnB, aconitase B; aspC, aspartate aminotransferase; dld, D-lactate dehydrogenase; gdhA, glutamate dehydrogenase; glnA, glutamine synthetase; gltA citrate synthase; HydABC, hydrogenase complex; icd, isocitrate dehydrogenase; pfl, pyruvate formate lyase; por, pyruvate ferredoxin oxidoreductase; ppc, phosphoenolpyruvate carboxylase; pps, phosphoenolpyruvate synthase; pta, phosphate acetyltransferase; gldD, glycerol-3-phosphate dehydrogenase; NDH-1, NADH–ubiquinone oxidoreductase complex.
In H. pylori, proton translocation is mediated by the NDH-1 dehydrogenase and the different cytochromes, including the primitive-type cytochrome cbb3 (Table 2 (PDF File: 169k)). Four respiratory electron-generating deydrogenases have been identified, glycerol-3-phosphate dehydrogenase (GlpD), D-lactate dehydrogenase, NADH–ubiquinone oxidoreductase complex (NDH-1), and a hydrogenase complex (HydABC). Our analysis also suggests that H. pylori is not able to use nitrate, nitrite, dimethylsulphoxide, trimethylamine N-oxide or thiosulphate as electron acceptors. Much of our metabolic analysis is supported by experimental evidence41,42.
Evolutionary relationships of H. pylori
H. pylori is currently classified in the Proteobacteria, a large, diverse division of Gram-negative bacteria which includes two other completely sequenced species, H. influenzae and E. coli. Given this taxonomic placement, based primarily on 16S rRNA sequence comparisons, one might expect the proteins of H. pylori more closely to resemble their H. influenzae and E. coli homologues rather than those in other genomes such as Synechocystis sp., M. genitalium, M. pneumoniae, M. jannaschii, and Saccharomyces cerevisae. This is indeed the case for many proteins. There are, however, many examples of H. pylori proteins in amino-acid biosynthesis, energy metabolism, translation and cellular processes that have greater sequence similarity to those found in non-Proteobacteria. For example, Dhs1, the initial enzyme in the chorismate biosynthesis pathway is 75.5% similar to Arabidopsis thaliana chloroplast Dhs1 gene product, and has minimal sequence similarity to the equivalent E. coli AroH, AroF or AroG gene products. The remaining enzymes in this pathway have strong sequence similarity to their E. coli counterpart. Similarly, the H. pylori prephenate dehydrogenase (TyrA), which converts chorismate to tyrosine, and six out of 15 enzymes in the aspartate amino acid biosynthetic pathways, resemble those from B. subtilis. A similar pattern can be seen in a different functional category. Nearly all H. pylori tRNA synthetases have eubacterial homologues, mostly with best matches to Proteobacteria species. However, histidyl-tRNA synthetase shows several amino-acid sequence signatures in common with eukaryotic and archaeal (M. jannaschii) homologues.
Such observations of discordant sequence similarity are often interpreted as evidence of lateral gene transfer in the evolutionary history of an organism. It is also possible that H. pylori diverged early from the lineage that led to the gamma Proteobacteria, and retained more ancient forms of enzymes that have been subsequently replaced or have diverged extensively in H. influenzae and E.coli.
Conclusion
Our whole-genome analysis of H. pylori gives new insight into its pathogenesis, acid tolerance, antigenic variation and microaerophilic character. The availability of the complete genome sequence will allow further assessment of H. pylori genetic diversity. This is an important aspect of H. pylori epidemiology as allelic polymorphism within several loci has already been associated with disease outcome5,21,31. The extent of molecular mimicry between H. pylori and its human host, an underappreciated topic, can now be fully explored43. The identification of many new putative virulence determinants should allow critical tests of their roles and thus new insight into mechanisms of initial colonization, persistence of this bacterium during long-term carriage, and the mechanisms by which it promotes various gastroduodenal diseases.
Methods
H. pylori strain 26695 (ref. 44) was originally isolated from a patient in the United Kingdom with gastritis (K. Eaton, personal communication) and was chosen because it colonizes piglets and elicits immune and inflammatory responses. It is also toxigenic, and transformable, and thus amenable to mutational tests of gene function.
The H. pylori genome sequence was obtained by a whole-genome random sequencing method previously applied to genomes of Haemophilus influenzae7, Mycoplasma genitalium8, and Methanococcus jannaschii9. Ninety-two per cent of the genome was covered by at least one λ clone and only 0.56% of the genome had single-fold coverage.
Open reading frames (ORFs) and predicted coding regions were identified using three methods. The predicted protein-coding regions were initially defined by searching for ORFs longer than 80 codons. Coding potential analysis of the entire genome was performed with a version of GeneMark45 trained with a set of H. pylori ORFs longer than 600 nucleotides. Coding sequences and potential starts of translation were also determined using GeneSmith (H.S., unpublished), a program that evaluates ORF length, separation of ORFs and overlap and quality of ribosome binding site. ORFs with low GeneMark coding potential, no database match, and not retained by GeneSmith were eliminated. GeneSmith identified 25 ORFs that are smaller than 100 codons, had no database match and were GeneMark negative. Frameshifts were detected by inspecting pairwise alignments, families of orthologues (similar proteins derived from different species) and paralogues (similar proteins from within the same organism), and regions containing homopolymer stretches and dinucleotide repeats. Ambiguities were resolved by an alternative sequencing chemistry (terminator reactions), and by sequencing PCR products obtained using the genomic DNA as template. Frameshifts that remain in the genome are considered authentic and not sequencing artefacts.
To determine their identity, ORFs were searched against a non-redundant amino-acid database as previously described9. ORFs were also analysed using 175 hidden Markov models constructed for a number of conserved protein families (pfam v1.0) using hmmer43. In addition, all ORFs were searched against the prosite motif database using MacPattern46. Families of paralogues were constructed by pairwise searches of proteins using FASTA. Matches that spanned at least 60% of the smaller of the protein pair were retained and visually inspected.
A unix version of the program TopPred47 was used to identify membrane-spanning domains (MSD) in proteins. Six hundred and sixty three proteins containing at least one MSD were found; of these, 300 had 2 potential MSDs or more. The presence of signal peptides and the probable position of the cleavage site in secreted proteins were detected using Signal-P, a neural net program that had been trained on a curated set of secreted proteins from Gram-negative bacteria48. 367 proteins were predicted to have a signal peptide. Lipoproteins were identified by scanning for the presence of a lipobox in the first 30 amino acids of every protein; 20 lipoproteins were identified, eighteen of which were Signal-P positive. Outer-membrane proteins were found by searching for aromatic amino acids at the end of the proteins.
Homopolymer and dinucleotide repeats were found by using RepScan (H.O.S., unpublished) which finds direct repeats of any length. All features identified using these programs were validated by visual inspection to remove false positives. Metabolic pathways were curated by hand and by reference to EcoCyc49.
References
Warren, J. R. & Marshall, B. Unidentified curved bacilli on gastric epithelium in active chronic gastritis. Lancet 1, 1273–1275 (1983).
Cover, T. L. & Blaser, M. J. Helicobacter pylori infection, a paradigm for chronic mucosal inflammation: pathogenesis and implications for eradication and prevention. Adv. Int. Med. 41, 85–117 (1996).
Mobley, H. L. T., Island, M. D. & Hausinger, R. P. Molecular Biology of Microbial Ureases. Microbiol. Rev. 59, 451–480 (1995).
Go, M. F. & Graham, D. Y. How dos Helicobacter pylori cause duodenal ulcer disease: The bug, the host, or both? J. Gastroentrol. Hepatol. (suppl.) 9, 8–12 (1994).
Labigne, A. & de Reuse, H. Determinants of Helicobacter pylori pathogenicity. Infect. Agents Disease 5, 191–202 (1996).
Clemens, J.et al. Impact of infection by Helicobacter pylori on the risk and severity of endemic cholera. J. Inf. Dis. 171, 1653–1656 (1995).
Fleischmann, R. D.et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496–512 (1995).
Fraser, C. M.et al. The Mycoplasma genitalium genome sequence reveals a minimal gene complement. Science 270, 397–403 (1995).
Bult, C. J.et al. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273, 1058–1073 (1996).
Winans, S. C., Burns, D. L. & Christie, P. J. Adaptation of a conjugal transfer system for the export of pathogenic macromolecules. Trends Microbiol. 4, 64–68 (1996).
Censini, S.et al. Cag, a pathogenicity island of Helicobacter pylori, encodes typeI-specific and disease-associated virulence factors. Proc. Natl Acad. Sci. USA 93, 14648–14653 (1996).
http://genome.wustl.edu/eddy/low/tRNAscan-SE-Manual/Manual.html
Akopyants, N. S., Kersulyte, D. & Berg, D. E. DNA rearrangement in the 40 kb cag (virulence) region in the Helicobacter pylori genome. Gut 39 (suppl. 2), A67 (1996).
Marczynski, G. T. & Shapiro, L. Bacterial chromosome origins of replication. Curr. Opin. Gen. Dev. 3, 775–782 (1993).
Riley, M. Functions of gene products of Escherichia coli. Microbiol. Rev. 57, 862–952 (1993).
Kornberg, A. & Baker, T. A. Replication mechanisms and operations in DNA replication.(ed. Kornberg, A. & Baker, T.) 471–510 (Freeman, New York, (1992)).
Macnab, R. M. in Escherichia coli and Salmonella Cellular and Molecular Biology(eds Neidhardt, F. C. et al.) 123–145 (ASM, Washington DC, (1996)).
Strom, M. S., Nunn, D. N. & Lory, S. Posttranslational processing of type IV prepilin and homologs by PilD of Pseudomonas aeruginosa. Meth. Enzymol. 235, 527–540 (1994).
Bardwell, J. C. Building bridges: disulphide bond formation in the cell. Mol. Microbiol. 14, 199–205 (1994).
Linn, S. in Escherichia coli and Salmonella Cellular and Molecular Biology(eds Neidhardt, F. C. et al.) 764–772 (ASM, Washington D.C., (1996)).
Peek, R. M., Thompson, S. A., Atherton, J. C., Blaser, M. J. & Miller, G. G. Expression of iceA, a novel ulcer-associated Helicobacter pylori gene, is induced by contact with gastric epithelial cells and is associated with enhanced mucosal IL-8. Gut 39 (suppl. 2), A71 (1996).
Curnow, A. W., Ibba, M. & Soll, D. tRNA-dependent asparagine formation. Nature 382, 589–590 (1996).
Jones, A. C., Foynes, S., Cockayne, A. & Penn, C. W. Gene cloning of a flagellar sheath protein of Helicobacter pylori shows its identity with the putative adhesin, HpaA. Gut 39 (suppl. 2), A62 (1996).
Boren, T., Falk, P., Roth, K. A., Larson, G. & Normark, S. Attachment of Helicobacter pylori to human gastric epithelium mediated by blood group antigens. Science 262, 1892–1895 (1993).
Ilver, D.et al. The Helicobacter pylori blood group antigen binding adhesin. Gut 39 (suppl. 2), A55 (1996).
Odenbreit, S., Till, M. & Haas, R. Optimized blaM-transposon shuttle mutagenesis of Helicobacter pylori allows identification of novel genetic loci involved in bacterial virulence. Mol. Microbiol. 20, 361–373 (1996).
Peterson, S. N.et al. Characterization of repetitive DNA in the Mycoplasma genitalium genome: possible role in the generation of antigenic variation. Proc. Natl Acad. Sci. USA 92, 11829–11833 (1995).
Moxon, E. R., Rainey, P. B., Nowak, M. A. & Lenski, R. E. Adaptive evolution of highly mutable loci in pathogenic bacteria. Curr. Biol. 4, 24–33 (1994).
Jonsson, A. B., Nyberg, G. & Normark, S. Phase variation of gonococcal pili by frameshift mutation in pilC, a novel gene for pilus assembly. EMBO J. 10, 477–488 (1991).
Tummuru, M. K. R., Sharma, S. A. & Blaser, M. J. Helicobacter pylori picB, a homologue of the Bordetella pertussis toxin secretion protein, is required for induction of IL-8 in gastric epithelial cells. Mol. Microbiol. 18, 867–876 (1995).
Atherton, J. C.et al. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. Association of specific vacA types with cytotoxin production and peptic ulceration. J. Biol. Chem. 270, 17771–17777 (1995).
Moran, A. P. The role of lipopolysaccharide in Helicobacter pylori pathogenesis. Aliment. Pharmacol. Ther. 10 (suppl. 1), 39–50 (1996).
Baker, P. J.et al. Molecular structures that influence the immunomodulatory properties of the lipid A and inner core region oligosaccharides of bacterial lipopolysaccharides. Infect. Immun. 62, 2257–2269 (1994).
Earhart, C. F. in Escherichia coli and Salmonella Cellular and Molecular Biology(eds Neidhardt, F. C. et al.) 1075–1090 (ASM, Washington DC, (1996)).
Evans, D. J. J, Evans, D. G., Lampert, H. C. & Nakano, H. Identification of four new prokaryotic bacterioferritins, from Helicobacter pylori, Anabaena variabilism, Bacillus subtilis and Treponema pallidum, by analysis of gene sequences. Gene 153, 123–127 (1995); Frazier, B. A.et al. Paracrystalline inclusions of a novel ferritin containing nonheme iron, produced by the human gastric pathogen Helicobacter pylori: evidence for a third class of ferritins. J. Bacteriol. 175, 966–972 (1993).
Suerbaum, S. The complex flagella of gastric Helicobacter species. Trends Microbiol. 3, 168–170 (1995).
Matin, A., Zychlinsky, E., Keyhan, M. & Sachs, G. Capacity of Helicobacter pylori to generate ionic gradients at low pH is similar to that of bacteria which grow under strongly acidic conditions. Infect. Immun. 64, 1434–1436 (1996).
Melchers, K.et al. Cloning and membrane topology of a P type ATPase from Helicobacter pyroli. J. Biol. Chem. 271, 446–457 (1996).
Melchers, K.et al. Cloning and analysis of two P type ion pumps of Helicobacter pylori, a cation resistance ATPase and a membrane pump necessary for urease activity. Gut 39 (suppl. 2), A67 (1996).
McGowan, C. C., Cover, T. L. & Blaser, M. J. Helicobacter pylori and gastric acid: biological and therapeutic implications. Gastroenterology 110, 926–938 (1996).
Hughes, N. J., Chalk, T. L., Clayton, C. L. & Kelly, D. J. Identification of carboxylation enzymes and characterization of a novel four-subunit pyruvate:flavodoxin oxidoreductase from Helicobacter pylori. J. Bacteriol. 177, 3953–3959 (1995).
Mendz, G. L. & Hazell, S. L. Aminoacid utilizaiton by Helicobacter pylori. Int. J. Biochem. Cell. Biol. 27, 1085–1093 (1995).
Sonnhammer, E. L. L., Eddy, S. R. & Durbin, R. Pfam: A comprehensive database of protein families based on seed alignments. Proteins(in the press).
Akopyants, N. S., Eaton, K. A. & Berg, D. E. Adaptive mutation and co-colonization during Helicobacter pylori infection of gnotobiotic piglets. Infect. Immun. 63, 116–121 (1995).
Borodovsky, M., Rudd, K. E. & Koonin, E. V. Intrinsic and extrinsic approaches for detecting genes in a bacterial genome. Nucleic Acids Res. 22, 4756–4767 (1994).
Fuchs, R. MacPattern: protein pattern searching on the Apple MacIntosh. Comput. Appl. Biosci. 7, 105–106 (1991).
Claros, M. G. & von Heijne, G. TopPred II: an improved software for membrane protein structure predictions. Comput. Appl. Biosci. 10, 685–686 (1994).
Nielsen, H., Engelbrecht, J., Brunak, S. & von Heijne, G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 10, 1–6 (1997).
Karp, P. D., Riley, M., Paley, S. M., Pellegrini-Toole, A. & Krummenacker, M. EcoCyc: Encyclopedia of Escherichia coli genes and metabolism. Nucleic Acids Res. 25, 43–51 (1997).
Doig, P., Exner, M. M., Hancock, R. E. & Trust, T. J. Isolation and characterization of a conserved porin protein from Helicobacter pylori. J. Bacteriol. 177, 5447–5452 (1995).
Acknowledgements
D.E.B., M.B. and W.H. are supported by grants from the NIH; P.K. is supported by a grant from the National Center for Research Resources. We thank N. S. Akopyants for preparing high quality chromosomal DNA from H. pylori strain 26695; M. Heaney, J. Scott, A. Saeed and R. Shirley for software and database support; and V. Sapiro, B. Vincent, J. Meehan and D. Mass for computer system support.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
Supplementary material for the article (PDF 525 kb)
Rights and permissions
About this article
Cite this article
Tomb, JF., White, O., Kerlavage, A. et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388, 539–547 (1997). https://doi.org/10.1038/41483
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/41483
This article is cited by
-
Integrative proteomic characterization of adenocarcinoma of esophagogastric junction
Nature Communications (2023)
-
Comparative genomics of two Vietnamese Helicobacter pylori strains, CHC155 from a non-cardia gastric cancer patient and VN1291 from a duodenal ulcer patient
Scientific Reports (2023)
-
Trimer stability of Helicobacter pylori HtrA is regulated by a natural mutation in the protease domain
Medical Microbiology and Immunology (2023)
-
Lactate causes downregulation of Helicobacter pylori adhesin genes sabA and labA while dampening the production of proinflammatory cytokines
Scientific Reports (2022)
-
Helicobacter pylori: an up-to-date overview on the virulence and pathogenesis mechanisms
Brazilian Journal of Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.