Abstract
Epstein-Barr virus has been linked to an increasing number of nonhematolymphoid conditions. Epstein-Barr virus was recently described in association with fibroadenomas of the breast occurring in immunosuppressed patients. To further investigate the potential association of Epstein-Barr virus with fibroadenoma in the context of immune dysfunction, 11 cases of fibroadenoma of the breast in immunosuppressed organ transplant recipients were examined. Cases were evaluated for the presence of Epstein-Barr virus by polymerase chain reaction, in situ hybridization, and immunohistochemical methods. The presence of Epstein-Barr virus genomic DNA was studied by polymerase chain reaction amplification using primers flanking the BamHI-W fragment of the Epstein-Barr virus genome, as well as the Epstein-Barr virus nuclear antigen-4 and latent membrane protein-1 genes. Cases were also evaluated for the presence of defective heterogeneous Epstein-Barr virus DNA. In addition, morphologic analysis by in situ hybridization for Epstein-Barr virus–encoded RNA-1 and immunohistochemistry for latent membrane protein-1 were performed. Epstein-Barr virus DNA was detected in 4 of 11 (36%) cases with BamHI-W polymerase chain reaction. Polymerase chain reaction studies for Epstein-Barr virus nuclear antigen-4 and latent membrane protein-1 genes were positive in two and four cases, respectively. No defective Epstein-Barr virus genomes were identified in any of the cases. Quantitative polymerase chain reaction demonstrated low levels of Epstein-Barr virus in the fibroadenomas studied. Despite the detection of Epstein-Barr virus genomes in a subset of the cases examined, the constituent epithelial and stromal components of all fibroadenomas demonstrated no evidence of Epstein-Barr virus–encoded RNA-1 by in situ hybridization or latent membrane protein-1 expression by immunohistochemistry. Rare Epstein-Barr virus–encoded RNA-1–positive lymphocytes were observed in some cases, which may account for the positive polymerase chain reaction results. The findings of the present study argue against a significant relationship between Epstein-Barr virus and fibroadenomas of the breast in the setting of transplant-associated immunosuppression.
Similar content being viewed by others
Main
Epstein-Barr virus (EBV) is a ubiquitous human herpesvirus, with serologic evidence of infection present in the majority of the population. EBV has been associated with a variety of hematolymphoid malignancies and also has been observed in a number of epithelial tumors including nasopharyngeal carcinoma (1), lymphoepithelioma-like carcinoma of various sites (2), and gastric adenocarcinoma (3, 4, 5, 6). In addition, several neoplasms arising in patients with primary and secondary immunodeficiencies are linked to EBV and include posttransplant lymphoproliferative disorders (7), as well as AIDS-associated lymphomas (8, 9) and smooth muscle tumors (10, 11).
Fibroadenoma is the most common benign tumor of the breast, occurring predominantly in premenopausal women. The etiology of fibroadenomas remains unclear; however, similar to breast carcinomas, the development of fibroadenomas is believed to be in part hormonally related, possibly secondary to estrogenic stimulus (12, 13). Several investigators have suggested an association between breast carcinoma and EBV (14, 15, 16, 17, 18); however, the results of other studies have argued against a significant role for EBV in the pathogenesis of breast carcinoma (19, 20, 21, 22, 23). EBV in fibroadenomas of the breast has not been well studied, but a recent report has documented the presence of EBV in a subset of these tumors. In that study, Kleer et al. (24) demonstrated molecular evidence of the EBV genome in 72% of rapidly growing fibroadenomas in immunosuppressed patients. EBV latent membrane protein (LMP-1) was also observed immunohistochemically in the cytoplasm of epithelial cells in 60% of the tumors studied. In contrast, EBV was not detected in a group of fibroadenomas from nonimmunosuppressed patients, suggesting a causative role for the virus in the pathogenesis of fibroadenomas in the context of immunocompromised hosts.
To further investigate the potential role of EBV in fibroadenomas of the breast in the setting of immunosuppression, we analyzed a series of fibroadenomas occurring in transplant allograft recipients for evidence of EBV. EBV expression was evaluated by determining the presence of EBV DNA (BamHI-W, EBV nuclear antigen-4 [EBNA-4], LMP-1) and defective heterogeneous (het) EBV DNA by polymerase chain reaction (PCR), of EBV-encoded RNA-1 (EBER-1) by in situ hybridization, and of EBV protein (LMP-1) by immunohistochemistry.
MATERIALS AND METHODS
Patient Samples
Eleven cases of fibroadenoma of the breast occurring in transplant allograft recipients were obtained from the files of the Department of Anatomic Pathology at the University of Pittsburgh Medical Center and the Department of Pathology at Stanford University Medical Center. Formalin-fixed, paraffin-embedded tissue from each case were used for PCR, in situ hybridization, and immunohistochemical studies as described below. Pertinent clinical information for each case was obtained.
Polymerase Chain Reaction
Viral genomic DNA was extracted from formalin-fixed, paraffin-embedded tissue blocks, using 0.2 mg/mL of proteinase K digestion buffer overnight, followed by denaturation by boiling. The PCR studies were performed with 2 μL of extracted DNA in a 30-μL mixture containing 50 mmol/L KCL, 10 mmol/L Tris buffer (pH 8.3), 50 μm of each deoxynucleotide triphosphate, 2.5 mmol/L MgCl2, 1 U of HotStarTaq DNA Polymerase (QIAGEN, Valencia, CA), and 20 pmol of each primer. Oligonucleotide primers corresponding to the BamHI-W fragment of the EBV genome were used. The primers used nucleotide positions 45451 to 45470 and nucleotide positions 45556 to 45575, respectively: 5′-CGGTCGCCCAGTCCTACCAG-3′ and 5′-CCTGGAGAGGTCAGGTTACT-3′. The expected amplification product size was 125 bp. For the EBNA-4 gene, primers were used that flank the DNA region encoding epitopes 399 to 408 and 416 to 424 of the prototype B95.8 EBV virus, using the nucleotide positions 96541 to 96540 and nucleotide positions 96770 to 96751, respectively: 5′-GAGGAGGAAGACAAGAGTGG-3′ and 5′-GATTCAGGCGTGGCTCTTGG-3′. The expected PCR product size was 230 bp. Primers for the EBV LMP-1 gene that flank the site of the characteristic 30-bp deletion of the LMP-1 gene were employed, using the nucleotide positions 168350 to 168331 and nucleotide positions 168190 to 168209, respectively: 5′-CGGAAGAGGTGGAAAACAAA-3′ and 5′-GTGGGGGTCGTCATCATCTC-3′. The expected LMP-1 gene product size was 161 bp, whereas a product containing the characteristic deletion was 131 bp. PCR for defective het EBV DNA was performed using primers that framed the junction of rearranged genomic DNA (5′-GCACATTAGCAATGCCTGTG-3′ and 5′-GTCCAGCGCGTTTACGTAAG-3′), as described elsewhere (25). Primers flanking the β-globin gene were used as a positive control for DNA preservation. After initial denaturation for 15 minutes at 95° C, 45 amplification cycles were performed as follows: denaturing at 94° C for 30 seconds, annealing for 30 seconds at 56° C for BamHI-W and het DNA, 57° C for EBNA-4, and 61° C for LMP-1, and extension at 72° C for 40 seconds. A final extension at 72° C for 7 minutes completed the PCR amplification. The PCR setup and post-PCR work were performed in separate laboratories to minimize the possibility of contamination. The amplified products obtained were separated by electrophoresis on gels and visualized with ethidium bromide staining under ultraviolet light. PCR products were subsequently subjected to Southern blot analysis and hybridized with appropriate probes specific for each particular region of the EBV genome of interest, as previously described (25, 26, 27, 28).
DNA extracted from paraffin embedded tissues was also used in a quantitative fluorogenic PCR assay to measure EBV DNA in the fibroadenomas. Primers EcoF (5′-TGGAGTTTCCCCCGATTCAA-3′) and EcoR (5′-TCCATGCTCTCGTCCACATCT-3′) were used to amplify EBV. Amplification of GAPDH (GAPDH Control Reagents; Applied Biosystems, Foster City, CA) was also performed as a normalization control. The cycling conditions were 50° C for 2 minutes, 95° C for 10 minutes, and 45 cycles at 95° C for 20 seconds and 60° C for 1.5 minutes. Serial dilutions of DNA extracted from the EBV-positive cell line Namalwa (American Type Culture Collection, CRL-1432), which contains two copies of EBV per cell, was used as a standard. A conversion factor of 6.6 pg of DNA per cell was used for expression of results as copy numbers. Data were collected and analyzed with the ABI Prism 7700 Sequence Detection System (Applied Biosystems). For EBV quantification, EBV copy number per sample was normalized to the amount of GAPDH present, and results were expressed as copies of EBV genomes per 100 cells.
In Situ Hybridization
The EBV RNA in situ hybridization study methods have been described previously (29). Briefly, in situ hybridization was performed using a 30-base oligonucleotide probe complementary to a portion of the EBER-1 gene, a region of the EBV genome that is actively transcribed in latently infected cells. The sequence used was 5′-AGACACCGTCCTCACCACCCGGGACTTGTA-3′ (Operon Technologies, San Pablo, CA). The probe was labeled with biotin at its 3′ end using methods previously described (30). Sections cut from formalin-fixed, paraffin-embedded tissue were deparaffinized, dehydrated, digested with pronase, incubated with prehybridization solution, and then hybridized overnight at a concentration of 0.25 ng/μL of probe. Sections were then incubated in a solution of avidin-alkaline phosphatase conjugate, washed for 3 minutes, incubated in McGadey’s substrate, briefly washed in distilled water, air dried, and coverslipped. No counterstain was used. A case was considered positive if the nucleus, or nucleus and cytoplasm, of a tumor cell stained dark brown or black over background levels. A poly d(T) was used as a control for total RNA preservation as described elsewhere (31). A known EBV-positive tumor and EBV-negative lymphoid tissue were used as controls.
Immunohistochemistry
Immunohistochemical staining was performed on formalin-fixed, paraffin-embedded sections using the mouse monoclonal antibody clone CS1–4 to LMP-1 protein (DAKO, Carpinteria, CA) at 1:600 dilution. Sections were deparaffinized in xylene and rehydrated in a graded ethanol series. Antigen retrieval was performed by steam heating slides in a 1 mmol/L solution of EDTA buffer (pH 8.0) in a steamer (Black and Decker, Shelton, CT) for 20 minutes. Staining was performed using an automated immunostainer (DAKO), followed by antibody detection using the DAKO EnVision+ System and 3,3′-diaminobenzidine as a chromogen. The slides were counterstained with hematoxylin and coverslipped. Sections of known EBV-positive classical Hodgkin lymphoma were used as positive controls. Membrane and cytoplasmic staining was considered positive.
RESULTS
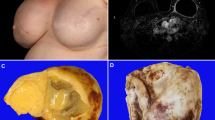
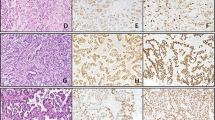
The clinical information and results of EBV studies are summarized in Table 1. All patients were therapeutically immunosuppressed female organ transplant recipients. The age of the patients ranged from 18 to 73 years, with a median age of 45 years. Types of transplant allografts included liver (n = 3), lung (n = 3), heart (n = 2), kidney (n = 2), and kidney/pancreas (n = 1). The size of the fibroadenomas ranged from 0.6 cm to 5.0 cm, with a mean size of 2.3 cm. Histologically, the fibroadenomas exhibited typical morphologic features that were characterized by a proliferation of stromal and epithelial elements in pericanalicular and intracanalicular patterns. Stromal cellularity was not increased, and the epithelial components did not exhibit hyperplastic proliferative changes. None of the fibroadenomas contained an unusual number of infiltrating lymphocytes, although rare scattered small lymphoid cells could be identified within the stroma in all cases.
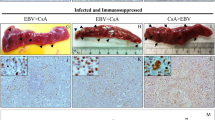
Strong β-globin amplified bands were identified by PCR from all 11 cases, indicating adequate DNA present. Of the 11 samples studied, 4 (36%) were found to be positive when amplified and probed for the BamHI-W sequence (Fig. 1). EBNA-4 genomic DNA was identified in two cases, and amplified EBV LMP-1 bands were observed in four cases. Two of the four EBV LMP-1–positive cases showed a 30-bp LMP-1 gene deletion. All positive fibroadenomas expressed more than one EBV gene. None of the 11 cases yielded a PCR product that was indicative of defective EBV genomes. In the EBV-positive fibroadenomas, an average of 4.1 genome copies per 100 cells (range, 0.2 to 10.1) were present, as determined by quantitative PCR. No evidence of EBV RNA was identified in the epithelial or stromal components of any of the fibroadenomas that were studied by in situ hybridization using the EBER-1 probe. Rare EBER-1–positive stromal lymphocytes were observed in three of the cases (Fig. 2). All cases studied exhibited strong positivity for poly d(T), indicating adequate RNA preservation. All of the fibroadenomas were negative for the LMP-1 protein by immunohistochemistry, with no evidence of immunoreactivity identified in either the stromal or epithelial cells.
DISCUSSION
In addition to a subset of non-Hodgkin and Hodgkin lymphomas (32), EBV has been linked to an increasing number of epithelial tumors (33, 34). Several studies have examined EBV in the context of breast neoplasia with disparate results. Although EBV DNA has been detected in a number of cases of breast carcinoma by PCR (14, 15, 16, 17, 18), there has been inconsistent demonstration of EBV RNA by in situ hybridization and EBV gene products by immunohistochemistry in the constituent tumor cells, leading some investigators to suggest that breast carcinomas are not truly associated with EBV (19, 20, 21, 22, 23).
In contrast to the data available regarding EBV and breast carcinoma, there is relatively little information in the literature pertaining to EBV and fibroadenomas. Grinstein et al. (18), in a study investigating EBV and epithelial tumors of various primary sites, observed no evidence of EBNA-1 expression in nine fibroadenomas that were examined by immunohistochemistry. In another recent report, Kleer et al. (24) examined the association of EBV with rapidly growing fibroadenomas of the breast occurring in a group of immunocompromised patients. The majority of patients were organ transplant recipients, whereas two patients were on immunosuppressive therapy for systemic lupus erythematosus. In this particular study, EBER-2 DNA was detected in 13 of 18 (72%) fibroadenomas by PCR, whereas cytoplasmic LMP-1 immunoreactivity was observed in the epithelial cells by immunohistochemistry in 12 of 19 cases. Nine cases were positive for EBV by both PCR and immunohistochemical methods. In contrast, no immunohistochemical evidence of LMP-1 expression was identified in a control group of 11 fibroadenomas from non-immunocompromised patients. Based on these findings, those investigators suggested that EBV infection is associated with fibroadenomas in immunosuppressed hosts.
In the current study, 4 of 11 (36%) fibroadenomas occurring in immunosuppressed organ transplant recipients had evidence of EBV by PCR. However, in contrast to the findings of Kleer et al. (24), no evidence of LMP-1 immunoreactivity was identified in either the epithelial or stromal components of the fibroadenomas studied. The reasons for this discrepancy in observed positivity for EBV by immunohistochemistry are unclear, but may be the result of methodological differences.
Although EBV DNA was detected by PCR in a number of cases of fibroadenoma in the present study, morphological analysis by EBER-1 in situ hybridization failed to confirm the presence of EBV in the constituent cells, demonstrating only rare scattered stromal EBER-1–positive lymphocytes. The low copy numbers of EBV, as determined by quantitative PCR, is also consistent with the observed in situ hybridization results. PCR is highly sensitive and can detect EBV in contaminating infected lymphocytes, which may account for the PCR positive cases in the current study. In immunosuppressed individuals in particular, loss of cytotoxic T-lymphocyte activity allows for accumulation and proliferation of EBV-infected cells, resulting in increased numbers of circulating infected lymphocytes. Although all of the PCR-positive cases in this study did not reveal EBER-1–positive lymphoid cells, infected lymphocytes may not have been present on these particular in situ hybridization preparations because of sampling error.
Although in situ hybridization demonstration of EBERs is the most effective means of establishing a significant association of EBV with a given neoplasm, in rare instances, an EBER-negative form of EBV infection has been observed (25, 35, 36). In some cases, disrupted EBER expression has been attributed to the presence of a defective EBV genome, termed het DNA, as suggested in a recent study demonstrating het EBV DNA in 8 of 24 cases of EBER-negative Hodgkin lymphoma (25). Defective EBV was not detected in any of the fibroadenomas in the present study and is an unlikely cause for the absence of EBER expression in these particular cases.
Fibroadenomas originate in the terminal duct-lobular units of the breast and are thought to develop from proliferation and expansion of the specialized connective tissue stroma (37). Etiologic factors influencing the development of fibroadenomas remain unclear. Fibroadenomas of the breast have been previously reported in the immunosuppressed organ transplant population and are, in this setting, frequently multiple and bilateral (38, 39, 40, 41, 42). Development of fibroadenomas in this context has been attributed to the effects of cyclosporin A therapy, though the exact mechanism by which this occurs has not been elucidated (38, 39, 40, 41, 42). Laboratory studies have shown that breast fibroadenomas can be induced in rats by inoculation with adenovirus type 9 (43, 44, 45, 46), raising the possibility of a viral etiology for human fibroadenomas. Using in situ hybridization with adenovirus type 9–specific probes, viral mRNA was identified only in the stromal cells and not the epithelial component of the fibroadenomas, suggesting virus-transformed stromal cells as the origin of these lesions in this particular animal model system (45). Although this rat model of viral-mediated fibroadenoma development may not be applicable to humans, one might expect that should a subset of human breast fibroadenomas be associated with EBV, the virus would be similarly localized to constituent stromal cells. No morphologic evidence of EBV was identified in the fibroadenomas that were analyzed by immunohistochemistry and in situ hybridization in the current study, and interestingly, although Kleer et al. (24) reported localization of the virus to epithelial cells in their EBV-associated fibroadenomas, the stromal component of the tumors was negative.
In summary, the results of the present study demonstrate no strong evidence for an association between EBV and fibroadenomas of the breast occurring in immunosuppressed organ transplant recipients. The lack of localization of EBV by morphological analyses in these cases argues against an etiologic role for the virus in the development of fibroadenomas in this particular patient population.
References
Nonoyama M, Huang CH, Pagano JS, Klein G, Singh S . DNA of Epstein-Barr virus detected in tissue of Burkitt’s lymphoma and nasopharyngeal carcinoma. Proc Natl Acad Sci U S A 1973; 70: 3265–3268.
Iezzoni JC, Gaffey MJ, Weiss LM . The role of Epstein-Barr virus in lymphoepithelioma-like carcinomas. Am J Clin Pathol 1995; 103: 308–315.
Shibata D, Weiss LM . Epstein-Barr virus-associated gastric adenocarcinoma. Am J Pathol 1992; 140: 769–774.
Tokunaga M, Land CE, Uemura Y, Tokudome T, Tanaka S, Sato E . Epstein-Barr virus in gastric carcinoma. Am J Pathol 1993; 143: 1250–1254.
Imai S, Koizumi S, Sugiura M, Tokunaga M, Uemura Y, Yamamoto N, et al. Gastric carcinoma: monoclonal epithelial malignant cells expressing Epstein-Barr virus latent infection protein. Proc Natl Acad Sci U S A 1994; 91: 9131–9135.
Hayashi K, Chen WG, Chen YY, Murakami I, Chen HL, Ohara N, et al. Deletion of Epstein-Barr virus latent membrane protein 1 gene in Japanese and Brazilian gastric carcinoma, metastatic lesions, and reactive lymphocytes. Am J Pathol 1998; 152: 191–198.
Hopwood P, Crawford DH . The role of EBV in post-transplant malignancies: a review. J Clin Pathol 2000; 53: 248–254.
Hamilton-Dutoit SJ, Pallesen G, Franzmann MB, Karkov J, Black F, Skinhoj P, et al. AIDS-related lymphoma. Histopathology, immunophenotype, and association with Epstein-Barr virus as demonstrated by in situ nucleic acid hybridization. Am J Pathol 1991; 138: 149–163.
Neri A, Barriga F, Inghirami G, Knowles DM, Neequaye J, Magrath IT, et al. Epstein-Barr virus infection precedes clonal expansion in Burkitt’s and acquired immunodeficiency syndrome-associated lymphoma. Blood 1991; 77: 1092–1095.
McClain KL, Leach CT, Jenson HB, Joshi VV, Pollock BH, Parmley RT, et al. Association of Epstein-Barr virus with leiomyosarcomas in young people with AIDS. N Engl J Med 1995; 332: 12–18.
Lee ES, Locker J, Nalesnik M, Reyes J, Jaffe R, Alashari M, et al. The association of Epstein-Barr virus with smooth-muscle tumors occurring after organ transplantation. N Engl J Med 1995; 332: 19–25.
Block GE, Zlatnik PA . Giant fibroadenomata of the breast in a prepubertal girl. A case report with observations of hormonal influences. Arch Surg 1960; 80: 155–159.
Martin PM, Kuttenn F, Serment H, Mauvais-Jarvis P . Studies on clinical, hormonal, and pathological correlations in breast fibroadenomas. J Steroid Biochem 1978; 9: 1251–1255.
Labrecque LG, Barnes DM, Fentiman IS, Griffin BE . Epstein-Barr virus in epithelial cell tumors: a breast cancer study. Cancer Res 1995; 55: 39–45.
Luqmani YA, Shousha A . Presence of Epstein-Barr virus in breast carcinoma. Int J Oncol 1996; 6: 899–903.
Bonnet M, Guinebretiere JM, Kremmer E, Grunewald V, Benhamou E, Contesso G, et al. Detection of Epstein-Barr virus in invasive breast cancers. J Natl Cancer Inst 1999; 91: 1376–1381.
Fina F, Romain S, Ouafik LH, Palmari J, Ben Ayed F, Benharkat S, et al. Frequency and genome load of Epstein-Barr virus in 509 breast cancers from different geographical areas. Br J Cancer 2001; 84: 783–790.
Grinstein S, Preciado MV, Gattuso P, Chabay PA, Warren WH, De Matteo E, et al. Demonstration of Epstein-Barr virus in carcinomas of various sites. Cancer Res 2002; 62: 4876–4878.
Chu JS, Chen CC, Chang KJ . In situ detection of Epstein-Barr virus in breast cancer. Cancer Lett 1998; 124: 53–57.
Glaser SL, Ambinder RF, DiGiuseppe JA, Horn-Ross PL, Hsu JL . Absence of Epstein-Barr virus EBER-1 transcripts in an epidemiologically diverse group of breast cancers. Int J Cancer 1998; 75: 555–558.
Brink AATP, van den Brule AJC, van Diest P, Meijer CJLM . Re. Detection of Epstein-Barr virus in invasive breast cancers. J Natl Cancer Inst 2000; 92: 655–656.
Chu PG, Chang KL, Chen YY, Chen WG, Weiss LM . No significant association of Epstein-Barr virus infection with invasive breast carcinoma. Am J Pathol 2001; 159: 571–578.
Deshpande CG, Badve S, Kidwai N, Longnecker R . Lack of expression of the Epstein-Barr virus (EBV) gene products, EBERs, EBNA1, LMP1, and LMP2A, in breast cancer cells. Lab Invest 2002; 82: 1193–1199.
Kleer CG, Tseng MD, Gutsch DE, Rochford RA, Wu Z, Joynt LK, et al. Detection of Epstein-Barr virus in rapidly growing fibroadenomas of the breast in immunosuppressed hosts. Mod Pathol 2002; 15: 759–764.
Gan YJ, Razzouk BI, Su T, Sixbey JW . A defective, rearranged Epstein-Barr virus genome in EBER-negative and EBER-positive Hodgkin’s disease. Am J Pathol 2002; 160: 781–786.
Imai S, Usui N, Sugiura M, Osato T, Sato T, Tsutsumi H, et al. Epstein-Barr virus genomic sequences and specific antibodies in cerebrospinal fluid in children with neurologic complications of acute and reactivated EBV infections. J Med Virol 1993; 40: 278–284.
Vasef MA, Kamel OW, Chen YY, Medeiros LJ, Weiss LM . Detection of Epstein-Barr virus in multiple sites involved by Hodgkin’s disease. Am J Pathol 1995; 147: 1408–1415.
Chu PG, Chang KL, Chen WG, Chen YY, Shibata D, Hayashi K, et al. Epstein-Barr virus (EBV) nuclear antigen (EBNA)-4 mutation in EBV-associated malignancies in three different populations. Am J Pathol 1999; 155: 941–947.
Chang KL, Chen YY, Shibata D, Weiss LM . Description of an in situ hybridization methodology for detection of Epstein-Barr virus RNA in paraffin-embedded tissues, with a survey of normal and neoplastic tissues. Diagn Mol Pathol 1993; 1: 246–255.
Weiss LM, Movahed LA, Chen YY, Shin SS, Stroup RM, Bui N, et al. Detection of immunoglobulin light-chain mRNA in lymphoid tissues using a practical in situ hybridization method. Am J Pathol 1990; 137: 979–988.
Weiss LM, Chen YY . Effects of different fixatives on detection of nucleic acids from paraffin-embedded tissues by in situ hybridization using oligonucleotide probes. J Histochem Cytochem 1991; 39: 1237–1242.
Chang KL, Weiss LM . The association of the Epstein-Barr virus with malignant lymphoma. Biomed Pharmacother 1996; 50: 459–467.
Gaffey MJ, Weiss LM . Association of Epstein-Barr virus with human neoplasia. Pathol Annu 1992; 27: 55–74.
Anagnostopoulos I, Hummel M . Epstein-Barr virus in tumours. Histopathology 1996; 29: 297–315.
Gilligan K, Rajadurai P, Resnick L, Raab-Traub N . Epstein-Barr virus small nuclear RNAs are not expressed in permissively infected cells in AIDS-associated leukoplakia. Proc Natl Acad Sci U S A 1990; 87: 8790–8794.
Sugawara Y, Mizugaki Y, Uchida T, Torri T, Imai S, Makuuchi M, et al. Detection of Epstein-Barr virus (EBV) in hepatocellular carcinoma tissue: a novel EBV latency characterized by the absence of EBV-encoded small RNA expression. Virology 1999; 256: 196–202.
Koerner FC, O’Connell JX . Fibroadenoma: morphologic observations and a theory of pathogenesis. Pathol Annu 1994; 29: 1–20.
Rolles K, Calne RY . Two cases of benign lumps after treatment with cyclosporine A [letter]. Lancet 1980; 2: 795.
Baildam AD, Higgins RM, Hurley E, Furlong A, Walls J, Venning MC, et al. Cyclosporin A and multiple fibroadenomas of the breast. Br J Surg 1996; 83: 1755–1757.
Campbell A, Moazami N, Ditkoff BA, Kurtz E, Eastbrook A, Schnabel F . Short-term outcome of chronic immunosuppression on the development of breast lesions in premenopausal heart and lung transplant patients. J Surg Res 1998; 78: 27–30.
Weinstein SP, Orel SG, Collazzo L, Conant EF, Lawton TJ, Czerniecki B . Cyclosporin A-induced fibroadenomas of the breast: report of five cases. Radiology 2001; 220: 465–468.
Sangthawan P, Fox J, Atkins RC, Kerr PG . Increased incidence of benign breast disease in female renal transplant patients receiving cyclosporin. Aust N Z J Surg 2002; 72: 222–225.
Ankerst J, Jonsson N, Kjellen L, Norrby E, Sjogren HO . Induction of mammary fibroadenomas in rats by adenovirus type 9. Int J Cancer 1974; 13: 286–290.
Jonsson N, Ankerst J . Studies on adenovirus type 9-induced mammary fibroadenomas in rats and their malignant transformation. Cancer 1977; 39: 2513–2519.
Javier R, Raska K, Macdonald GJ, Shenk T . Human adenovirus type 9-induced rat mammary tumors. J Virol 1991; 65: 3192–3202.
Javier R, Shenk T . Mammary tumors induced by human adenovirus type 9: a role for the viral early region 4 gene. Breast Cancer Res Treat 1996; 39: 57–67.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lau, S., Chen, YY., Berry, G. et al. Epstein-Barr Virus Infection Is Not Associated with Fibroadenomas of the Breast in Immunosuppressed Patients after Organ Transplantation. Mod Pathol 16, 1242–1247 (2003). https://doi.org/10.1097/01.MP.0000097363.72401.00
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/01.MP.0000097363.72401.00
Keywords
This article is cited by
-
The possible involvement of virus in breast cancer
Journal of Cancer Research and Clinical Oncology (2009)
-
Analysis of Epstein-Barr virus reservoirs in paired blood and breast cancer primary biopsy specimens by real time PCR
Breast Cancer Research (2006)