Abstract
There will be an increasing need of methods for assessing the suitability of specimens for genetic-based assays as DNA markers become an integral part of molecular diagnosis. The targeting of specimens for specific analyses will require the ability to rapidly screen for DNA quality. Conventional methods such as Southern analysis and gene specific-polymerase chain reaction (PCR) often require quantities of material that represent a significant portion of the specimen, especially in microdissected samples. Here we describe a novel application of a commonly used PCR-based DNA-fingerprinting technology that requires minimal quantities of DNA to simultaneously assess multiple regions throughout the genome for DNA quality. Randomly amplified polymorphic DNA (RAPD) PCR generates DNA fragments of a broad size range with the product size reflecting the degree of sample fragmentation. Fourteen DNA samples extracted from cells microdissected from seven formalin-fixed, paraffin-embedded oral cancer biopsies were assessed for DNA quality using gene-specific PCR and RAPD-PCR. Although the more conventional assay required 2-ng DNA (or 300-cell equivalents) to examine DNA quality at a single locus, RAPD-PCR provided a more informative profile of DNA quality from the same microdissected archival specimens.
Similar content being viewed by others
INTRODUCTION
Archival specimens represent a vast resource for discovery and evaluation of prognostic DNA markers. Knowledge of DNA quality is important to determine the types of techniques that the material can support. The quality of these specimens is dependent on fixation and storage conditions. Some DNA samples are highly fragmented or damaged by preservation procedures and require special handling in reactions, for example, the use of closer-spaced primers or larger quantities of DNA in a PCR-based assay (1, 2, 3, 4). Other samples are better preserved, generating amplified products of larger size. Because specimen yield is often a limiting factor in studying nonrenewable, archival material, the ability to assess DNA quality, with a minimal amount of material, would allow the sorting of such specimens for specific analytical use.
Several methods are commonly used to assess DNA quality (Table 1). Although these methods provide information on the degree of DNA fragmentation, not all can predict the sample's ability to support PCR. For example, procedures used to preserve hospital specimens may introduce a wide array of DNA alterations, including fragmentation as well as cross-linking and base loss (1, 2, 3, 4). In most cases, researchers do not prescreen samples for DNA quality but rather directly measure the status of a gene by single or multiplex PCR, accepting a possible high failure rate (5, 6, 7). Consequently, this requires the repetition of the PCR reaction with an increased amount of DNA and/or with multiple primers that generate the gene in smaller pieces consuming a significant portion of the DNA sample. RAPD-PCR (8, 9, 10, 11), a robust technique, enables consistent genetic fingerprints from archival, microdissected specimens by the random amplification of multiple loci throughout the genome, producing an unbiased representation of DNA quality. Here we report the use of this technique as an effective method for assessing DNA quality in archival specimens.
MATERIALS AND METHODS
Microdissection of Tissue and Extraction of DNA
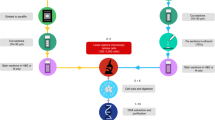
Formalin-fixed, paraffin-embedded archival biopsies from seven oral squamous cell carcinomas were obtained from the Oral Biopsy Service of British Columbia. Tissue sections of these samples were deparaffinized using xylene and alcohol, stained with hematoxylin and eosin, and manually microdissected by LZ, an oral pathologist. Two samples were collected from the same section for each specimen: tumor and connective tissue. Dissected cells were digested with proteinase K, and DNA was extracted and quantified fluorometrically (Picogreen kit, Molecular Probes, Eugene, OR).
Gene-Specific PCR
Two sizes of fragments were amplified from the human glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene (GenBank accession no. J04038) in 25-μL reaction mixtures containing 50 mm of KCl; 10 mm of Tris-HCl, pH 9; 0.1% Triton X-100; 2 mm of MgCl2; 0.2 mm of deoxyribose trinucleotides (dNTPs); 0.8 μm of each GAPDH primer; 2.5 U of recombinant Taq DNA polymerase; and 2 ng of DNA. The PCR primers were as follows: GAP1 (5′-AACCTGCCAAATATGATGACATCA-3′), GAP2 (5′-GTCGTTGAGGGCAATGCCA-3′), and GAP3 (5′-GTCTTACTCCTTGGAGGCCATGA-3′). GAP1 and 2 produced a 163–base pair (bp) fragment, whereas GAP1 and 3 produced a 363-bp fragment (Fig. 1A). PCR conditions were as follows: denaturation at 94° C for 2 minutes, followed by 35 cycles of amplification (94° C for 45 s, 60° C for 45 s, and 72° C for 2 min) and a final extension (72° C for 10 min) in a PTC-100 Thermal Cycler (MJ Research, Waltham, MA).
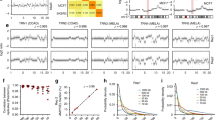
DNA quality assessment of 7 microdissected archival oral tumor and connective tissue pairs. A, glyceraldehyde 3-phosphate dehydrogenase (GAPDH) gene with primer annealing positions. Numbers indicate nucleotide position on GAPDH sequence (GenBank Accession No. J04038). B, PCR amplification (35 cycles) of a 163-bp and a 363-bp GAPDH gene fragment. PCR products were size separated on 1% agarose gels. C, connective tissue; T, tumor. C, RAPD-PCR fingerprints of samples shown in B. +, positive control, high–molecular weight DNA extracted from a frozen lung biopsy; −, negative control, a mock reaction with no template DNA.
Randomly Amplified Polymorphic DNA (RAPD)-PCR
RAPD-PCR primers were selected on the basis of the number of fragments generated and their distribution on a 6% nondenaturing polyacrylamide gel. A number of primers purchased from Operon Technologies (http://www.operon.com) were assessed, and PCR primers (5′-aatcgggctg-3′ and 5′-gaaacgggtg-3′) were chosen for this study. Twenty pmol of each primer were labeled with 1 μCi of [γ-32P] adenosine triphosphate (Amersham Pharmacia Biotech, Piscataway, NJ). Reactions were performed in a 10-μL volume containing 2 ng DNA, 2.5 U of recombinant Taq DNA polymerase, 200 μm of each dNTP, 10 mm of Tris-HCl (pH 8.3), 50 mm of KCl, 2 mm of MgCl2, and 0.001% gelatin. All reactions were performed for 45 cycles consisting of 94° C for 1 minute, 35° C for 1 minute, and 72° C for 2 minute. PCR products were resolved on 6% non-denaturing polyacrylamide gels (37:1 acrylamide: N′, N′-methylene-bis-acrylamide) and imaged by autoradiography.
RESULTS AND DISCUSSION
Assessment of DNA Using Gene-Specific PCR
Seven arbitrarily chosen sample pairs (14 samples in total) were assessed by gene-specific PCR. Figure 1B illustrates the results. Although all samples generated 163-bp fragments, only eight yielded a 363-bp PCR product. The latter included samples from Cases 402, 446, 448, and 451. For each specimen, DNA extracted from connective tissue (C) and tumor (T) showed the same PCR response.
Assessment of DNA Quality Using RAPD
RAPD profiles from these samples were generated using the same quantity of DNA (Fig. 1C). The size range of RAPD products varied among sample pairs, again with both members of a sample pair giving a similar profile. Three of the sample pairs (446, 448, and 451) had fingerprint patterns that resembled the profile of cryopreserved tissue that contains high-quality DNA. A fourth (402) gave reproducible PCR products up to 350 bp. All four of these sample pairs gave a 363-bp GAPDH fragment (Fig. 1B). In contrast, Sample 405 gave a reliable profile up to 200 bp, with less amplification of larger sized fragments (lighter bands) and variation between C and T for fragments of >200 bp. Specimens 414 and 416 had the worst profiles with fragments restricted to <150 bp. Increasing DNA concentration did not improve the quality of these profiles. These results are consistent with the absence of a GAPDH 363-bp fragment in Figure 1B.
Targeting Analyses to Maximize Use of Material
We have extracted DNA from 137 oral premalignant lesions and have obtained on average 100 ng per sample. Although DNA yield varied between samples, on average, we retrieved 100 ng of DNA from three to four 12-μm sections for tumors and from 10–12 sections for oral premalignant lesions. As shown in Figure 2, 43% of cases produced profiles with >500-bp fragments in an RAPD assay, 29% produced intermediate-size fragments of 200 to 500 bp, and 28% yielded only small fragments of <200 bp. It should be noted that there were no obvious histological characteristics of these specimens that would have predicted this variation in DNA quality. This information could have several applications. Although all of the samples can be used for microsatellite analysis (12, 13, 14) and the amplification of small exons, only a portion of samples will support procedures that require the amplification of larger fragments; that is, degenerate oligonucleotide primed PCR for comparative genomic hybridization analysis (15, 16, 17, 18, 19, 20) or Southern blot analysis, cycle sequencing, and single strand conformational polymorphism assessment for mutation analysis (21). Through prescreening by RAPD-PCR, it is possible to prevent the depletion of low-quality DNA in experiments that require more intact material, thus maximizing the use of precious specimens.
Accession codes
References
Giannella C, Zito FA, Colonna F, Paradiso A, Marzullo F, Alaibac M, et al. Comparison of formalin, ethanol, and Histochoice fixation on the PCR amplification from paraffin-embedded breast cancer tissue. Eur J Clin Chem Clin Biochem 1997; 35(8): 633–635.
Koshiba M, Ogawa K, Hamazaki S, Sugiyama T, Ogawa O, Kitajima T . The effect of formalin fixation on DNA and the extraction of high-molecular-weight DNA from fixed and embedded tissues. Pathol Res Pract 1993; 189(1): 66–72.
Murase T, Inagaki H, Eimoto T . Influence of histochemical and immunohistochemical stains on polymerase chain reaction. Mod Pathol 2000; 13(2): 147–151.
Tbakhi A, Totos G, Pettay J, Myles J, Tubbs R . The effect of fixation on detection of B-cell clonality by polymerase chain reaction. Mod Pathol 1999; 12(3): 272–278.
Cawkwell L, Quirke P . Direct multiplex amplification of DNA from a formalin fixed, paraffin wax embedded tissue section. Mol Pathol 2000; 53(1): 51–52.
Chakrabarti S, Price BD, Tetradis S, Fox EA, Zhang Y, Maulik G, et al. Highly selective isolation of unknown mutations in diverse DNA fragments: toward new multiplex screening in cancer. Cancer Res 2000; 60(14): 3732–3737.
Lin Z, Cui X, Li H . Multiplex genotype determination at a large number of gene loci. Proc Natl Acad Sci U S A 1996; 93(6): 2582–2587.
Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV . DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 1990; 18(22): 6531–6535.
Maeda T, Jikko A, Hiranuma H, Fuchihata H . Analysis of genomic instability in squamous cell carcinoma of the head and neck using the random amplified polymorphic DNA method. Cancer Lett 1999; 138(1–2): 183–188.
Ong TM, Song B, Qian HW, Wu ZL, Whong WZ . Detection of genomic instability in lung cancer tissues by random amplified polymorphic DNA analysis. Carcinogenesis 1998; 19(1): 233–235.
Anami Y, Takeuchi T, Mase K, Yasuda J, Hiroshashi S, Perucho M, et al. Amplotyping of microdissected, methanol-fixed lung carcinoma by arbitrarily primed polymerase chain reaction. Int J Cancer 2000; 89(1): 19–25.
Wistuba II, Behrens C, Virmani AK, Mele G, Milchgrub S, Girard L, et al. High resolution chromosome 3p allelotyping of human lung cancer and preneoplastic/preinvasive bronchial epithelium reveals multiple, discontinuous sites of 3p allele loss and three regions of frequent breakpoints. Cancer Res 2000; 60(7): 1949–1960.
Rosin MP, Cheng X, Poh C, Lam WL, Huang Y, Lovas J, et al. Use of allelic loss to predict malignant risk for low-grade oral epithelial dysplasia. Clin Cancer Res 2000; 6(2): 321–322.
Zhang L, Michelsen C, Cheng X, Zeng T, Priddy R, Rosin MP . Molecular analysis of oral lichen planus. A premalignant lesion? Am J Pathol 1997; 151(2): 323–327.
Aubele M, Mattis A, Zitzelsberger H, Walch A, Kremer M, Hutzler P, et al. Intratumoral heterogeneity in breast carcinoma revealed by laser—microdissection and comparative genomic hybridization. Cancer Genet Cytogenet 1999; 110(2): 94–102.
Aubele M, Mattis A, Zitzelsberger H, Walch A, Kremer M, Welzl G, et al. Extensive ductal carcinoma in situ with small foci of invasive ductal carcinoma: evidence of genetic resemblance by CGH. Int J Cancer 2000; 85(1): 82–86.
Daigo Y, Chin SF, Gorringe KL, Bobrow LG, Ponder BA, Pharoah PD, et al. Degenerate oligonucleotide primed-polymerase chain reaction-based array comparative genomic hybridization for extensive amplicon profiling of breast cancers: a new approach for the molecular analysis of paraffin-embedded cancer tissue. Am J Pathol 2001; 158(5): 1623–1631.
Jones C, Foschini MP, Chaggar R, Lu YJ, Wells D, Shipley JM, et al. Comparative genomic hybridization analysis of myoepithelial carcinoma of the breast. Lab Invest 2000; 80(6): 831–836.
Looijenga LH, Rosenberg C, van Gurp RJ, Geelen E, van Echten-Arends J, de Jong B, et al. Comparative genomic hybridization of microdissected samples from different stages in the development of a seminoma and a non-seminoma. J Pathol 2000; 191(2): 187–192.
Weber RG, Scheer M, Born IA, Joos S, Cobbers JM, Hofele C, et al. Recurrent chromosomal imbalances detected in biopsy material from oral premalignant and malignant lesions by combined tissue microdissection, universal DNA amplification, and comparative genomic hybridization. Am J Pathol 1998; 153(1): 295–303.
Suzuki Y, Orita M, Shiraishi M, Hayashi K, Sekiya T . Detection of ras gene mutations in human lung cancers by single-strand conformation polymorphism analysis of polymerase chain reaction products. Oncogene 1990; 5(7): 1037–1043.
Acknowledgements
The authors wish to thank Jenny Siwoski, Chad Malloff, Kim Lonergan, Sandra Henderson, and Laura Jane Henderson for careful reading of this manuscript and expert assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article was supported by a grant from the Canadian Institute of Health Research (CIHR) using samples collected through Grant 1 R01 DE13124–01 from NIDCR.
Rights and permissions
About this article
Cite this article
Siwoski, A., Ishkanian, A., Garnis, C. et al. An Efficient Method for the Assessment of DNA Quality of Archival Microdissected Specimens. Mod Pathol 15, 889–892 (2002). https://doi.org/10.1097/01.MP.0000024288.63070.4F
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/01.MP.0000024288.63070.4F
Keywords
This article is cited by
-
Comparison of Different Drying Methods for Recovery of Mushroom DNA
Scientific Reports (2017)
-
High-efficiency genotype analysis from formalin-fixed, paraffin-embedded tumor tissues
The Pharmacogenomics Journal (2011)
-
Assessing the digestion of a genetically modified tomato (Solanum lycopersicum) R8 DNA in simulated gastric fluid using event-specific real-time PCR
European Food Research and Technology (2011)
-
Large fragment Bst DNA polymerase for whole genome amplification of DNA from formalin-fixed paraffin-embedded tissues
BMC Genomics (2006)
-
Chromosome 5p aberrations are early events in lung cancer: implication of glial cell line-derived neurotrophic factor in disease progression
Oncogene (2005)