Abstract
Comparative genomic hybridization (CGH) is a molecular cytogenetic technique that provides an overview on chromosomal imbalances within the whole tumor cell genome. This method has yet not been applied in effusion cytology. We performed CGH analysis in malignant effusions, fine needle aspirates, and imprint smears from eight ovarian adenocarcinomas, three breast carcinomas, one colon adenocarcinoma, and three malignant mesotheliomas. In part, CGH analysis from fresh frozen tissue and classic karyotyping served as controls. In this series, 14/15 cytologic specimens were suitable for extraction of high molecular weight DNA sufficient for reliable CGH analysis. CGH profiles from cytologic material were equal or even more significant in comparison with corresponding fresh frozen tumor samples. We conclude that CGH analysis from cytologic specimens may support the primary cytologic diagnosis of malignancy, especially in the differential diagnosis of benign proliferating mesothelium, malignant mesothelioma, and metastatic adenocarcinoma. CGH analysis of metastatic lesions may provide information on the site of the primary tumors and detects cytogenetic imbalances affecting oncogenes and tumor suppressor genes involved in tumor progression and metastatic spread.
Similar content being viewed by others
INTRODUCTION
A specific cytologic diagnosis based on morphological criteria alone is limited in various pathologic conditions including, for example, effusion cytology. In such cases, immunocytochemistry is frequently used to provide additional information. However, even the use of a panel of different antibodies does not achieve 100% sensitivity and specificity. Other methods for optimizing sensitivity and specificity in the cytologic diagnosis include DNA cytometry (1) and an increasing number of molecular genetic methods (2, 3, 4).
Comparative genomic hybridization (CGH) is a molecular cytogenetic method first described by Kallioniemi and coworkers (5), which scans the entire genome in a tumor sample and thus provides an overview on DNA sequence copy number changes. Based on fluorescence in situ hybridization, CGH measures the ratio between fluorochrome-labeled tumor DNA and differently fluorochrome-labeled normal DNA hybridized simultaneously to normal target metaphase chromosomes. Computer-based analysis of the fluorescence pattern reveals losses, gains, and amplifications of chromosomal segments (6). CGH analysis has been reported in >2400 solid neoplasms, and the results were recently reviewed (7, 8). However, this method has not yet been widely considered as a diagnostic adjunct in the cytological investigation of human neoplasms.
This study should not provide a detailed cytogenetical evaluation of a specific tumor type, but rather evaluates whether common cytologic materials, especially effusion specimens, are suited for genomic analysis by CGH.
Additionally, conceivable practical applications of CGH analysis in the cytological diagnosis are discussed.
MATERIALS AND METHODS
Cytologic Specimens
The cytologic material investigated consisted of 11 malignant serous effusion specimens, 2 fine needle aspirates (FNAs), and 2 imprint smears from malignant solid tumors. Clinical data and cytologic and histologic diagnoses from all 15 patients are summarized in Table 1. Fine needle aspirates were performed using a 23-gauge needle attached to a 10-mL plastic syringe. From each case, at least two slides were stained according to May-Grünwald-Giemsa (MGG) and Papanicolaou for cytological analysis. In single cases, additional immunocytological stainings were performed to support the morphological diagnosis.
In all cases with the cytological diagnosis of adenocarcinoma or metastatic adenocarcinoma, the histologic slides and/or the clinical charges were reviewed for the primary tumor. The three cases cytologically suspicious for malignant mesothelioma were verified by subsequent histology.
From each case, cytological material was reserved for molecular analysis. From five malignant effusions, 10-mL aliquots were processed by centrifuge at 1000 rpm for 5 minutes, the supernatant was discharged, and pelleted cells were snap-frozen in liquid nitrogen and stored at −70° C until further processing. From the remaining cases, air dried (for ≥24 h at room temperature) and unstained smears were wrapped in aluminum foil and stored at −70° C until DNA extraction.
Fresh Frozen Tumor Tissue
From three carcinomas (Nos. 4, 9, and 11), fresh frozen tumor tissue corresponding to the cytologic specimens were snap-frozen in liquid nitrogen and stored at −70° C.
DNA Isolation and CGH Analysis
For CGH, DNA was isolated in 10 cases from either unstained or MGG stained smear preparations containing 105 to 106 cells and in the remaining 5 cases, from frozen cell pellets. MGG-stained smears were stored overnight in xylol to remove cover slides and then subsequently destained in methanol, rehydrated, and washed in PBS-buffer. For efficient yield of tumor DNA from cell smears, lysis buffer (Qiagen, Hilden, Germany) containing proteinase K (1 mg/mL final concentration; Roche, Mannheim, Germany) was dropped on the slides, then covered by a second slide and incubated at 56° C overnight in 50-mL falcon tubes placed in a rotor shaker (30 rpm). The lysate was recovered by centrifugation (5 min, 2500 rpm). In all cases, digestion by proteinase K was followed by spin column purification (Qiagen). CGH analysis was principally performed as described previously (9).
Preparation of Metaphases
Human lymphocytes were cultured in RPMI 1640 medium supplemented with 10% fetal calf serum. Phytohemagglutinin (0.1 mg/40 mL medium) was added to stimulate growth of the lymphocytes. After 3 days of cell culture, mitotic cells were arrested by colchicine incubation followed by hypotonic treatment and subsequent fixation in methanol–acetic glacial acid (3:1) at 4° C. After repeated washing in ice-cold fixation solution, metaphases were dropped on water-rinsed glass slides. They were then subjected to digestion with pepsin in 0.01 N HCl.
Nick Translation and Hybridization
Briefly, labeling of tumor DNA with biotin-16-dUTP (Roche, Mannheim, Germany) and of normal reference DNA with digoxigenin-11-dUTP (Roche) was carried out by standard nick translation. The denatured DNA probe containing 2 μg of tumor DNA, 1.5 μg ov reference DNA, and 80 μg of Cot-1 DNA (Roche) was hybridized for 3 days to a normal metaphase spread (15 × 15 mm cover glass area). The slides were then washed extensively, blocked with bovine serum albumin (BSA) solution, and incubated with fluorescein-conjugated avidin (Vector Laboratories, Burlingame, CA) and rhodamine-conjugated antidigoxigenin (Roche). The slides were washed and mounted in antifade solution (Vector Laboratories) containing 2.5 μg/mL of 4.6-diamino-2-phenylindole (DAPI) counterstain.
Digital Image Analysis
Digital images of metaphase slides were obtained with a charge-coupled device (CCD) camera on a Zeiss Axioskop 2 epifluorescence microscope using filter-set for DAPI, fluorescein isothiocyanate, and rhodamine fluorescence. Images were analyzed with Quips CGH software (Applied Imaging, Santa Barbara, CA). The average green-to-red ratio for each case was calculated for ≥10 metaphases, and only definitively assigned chromosomes were included for analysis. Poorly or inhomogeneously stained metaphases were excluded from analysis, as well as heavily bent or overlapping chromosomes. Threshold values for green-to-red ratio profiles were set on 0.8 and 1.2, respectively, to indicate significant losses and gains of chromosomal material (6). Chromosomal aberrations were scored as borderline when the ratio profile bordered to the threshold value or had broken regions that failed to surpass the threshold value. Imbalances indicated by ratio profiles as a trend were not considered for profile analysis. Gains of chromosomal material surpassing the threshold of 1.5 were defined as high-level amplifications. Centromeric regions, the short arm of the acrocentric chromosomes, and the heterochromatin blocks on 1q12, 9q12, 16q11.2, and Yq12 were excluded from CGH profile analysis.
Cell Culture and Classic Cytogenetics
Sedimented cells from three malignant effusions were seeded in 25-cm2 culture flasks for short-term cell cultures. Cells were cultured in RPMI-1640 medium supplemented with L-glutamine and 10% fetal calf serum (FCS) at 37° C in a humidified 5% CO2–95% air atmosphere. In one case, the tumor cells did not grow. From both remaining cases, chromosome preparations were performed as described previously (10). More than 20 DAPI-banded metaphases were karyotyped according to the International System for Human Cytogenetic Nomenclature.
RESULTS
Cytomorphologic Analysis
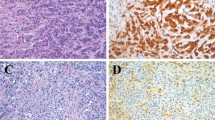
In 12 cases, the cytologic smears were predominately composed of atypical cells, and the cytologic diagnosis of adenocarcinoma or metastatic adenocarcinoma was made (Fig. 1A). In the three remaining cases, the cytologic findings were suspected of malignant mesothelioma, and all of these were confirmed by subsequent histology, as well as differentiated malignant mesothelioma of epithelial type in one case and biphasic malignant mesothelioma in two cases (Fig. 1B). The cytological findings and histological diagnoses from corresponding primary tumors are summarized in Table 1.
A, the smear preparation from malignant ascites (Case 6) contains approximately 2 × 105 metastatic carcinoma cells from serous ovarian carcinoma (May-Grünwald-Giemsa; MGG). B, pleural effusion specimen from malignant mesothelioma (Case 15) demonstrates numerous malignant cells intermingled with some histiocytes (MGG). C, fine needle aspiration smear from mucinous breast carcinoma (Case 9) composed almost exclusively of carcinoma cells embedded in epithelial mucin (MGG). D, in the corresponding tissue section, the carcinoma cells are surrounded by nonneoplastic stroma cells.
DNA Extraction
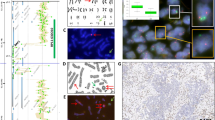
DNA extraction from air-dried cytologic smears and from cell pellets yielded genomic DNA of high molecular weight as shown for three samples in Figure 2. The DNA extraction from the fresh frozen tissues also yielded high molecular weight DNA comparable to the cytologic material.
CGH Analysis
CGH analysis was carried out successfully in all cases except Case 15, which lacked a green fluorescence signal because of insufficient incorporation of biotin-16-dUTP during nick translation.
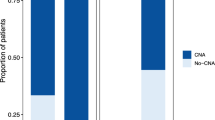
CGH analysis demonstrated in all 14 cytologic specimens composed of various malignant cells, mostly complex genomic aberrations with gains and losses of chromosomal material (Table 2). Imbalances seen only as a trend were not considered for analysis. Chromosomal gains were a more frequent finding than chromosomal losses (mean 8.8 gain per tumor versus 5 loss per tumor). The genetic imbalances are shown graphically in Figure 3 for the eight ovarian carcinomas and for the remaining six malignancies.
Summary of chromosomal imbalances detected by comparative genomic hybridization (CGH) in cytologic specimens. Vertical lines on the left of each chromosome ideogram represent a loss of genetic material, whereas vertical lines on the right correspond to gains. Ratio profiles that bordered to the threshold value are indicated in gray lines. Bold lines on the right side represent high amplification sites surpassing the threshold value of 1.5. Chromosomal imbalances found in ovarian carcinomas are shown separately (A) from CGH results detected in breast carcinomas, colon carcinoma, and malignant mesotheliomas (B).
Ovarian Carcinomas
On average, 19.6 genetic changes were found in the ovarian carcinomas (Nos. 1–8), with a range of 8 to 35. The most frequent gains involved parts or the entire long arm of chromosomes 8 (100%), 20 (88%), and 3 (63%). The most frequent losses occurred at chromosome 4q (75%), 18q (63%), and 9q (50%). High-amplification sites were found in three cases on 8q, in two cases on 20q, and in one case on 17q and 20p. Ratio profiles of −9p and −Xp in Case 2 reached the value of 0.6.
Breast Carcinomas
DNA copy number changes in three breast carcinomas (Nos. 9, 10, and 11) ranged from 7 to 11 and demonstrated, among others, gains at 8q23qter, 19q, and 20q in two cases each and losses at 11q in two cases. A high-level amplification was detected at 11q11q14.
Colonic Carcinoma
The metastatic adenocarcinoma of the colon (No. 12) demonstrated seven DNA copy number changes, including gain of Xq11q21, 8q23q24.1, and 17q11q12.
Malignant Mesotheliomas
This study included three cases (Nos. 13, 14, and 15) of histologically verified malignant mesothelioma. CGH analysis from a MGG-stained smear preparation (No. 13) and one cell pellet (No. 14) demonstrated significant genomic imbalances. CGH from malignant mesothelioma of epithelial type (No. 13) revealed the loss of 6q. In this case, a loss of 22q was found tendentiously, which is not included in Figure 3B. The biphasic malignant mesothelioma No. 14 showed gains at chromosomes 1p36, 16p, 17, and 19. In Case 15, the CGH analysis failed.
Comparison between CGH and Classic Cytogenetics
In one case of ovarian serous papillary adenocarcinoma (No. 6), short-term cell culturing of tumor cells from ascites for conventional karyotyping was successful and revealed the following complex aneuploid karyotype: 55–64, X, −X, −X, del (1) (p36), add (1) (p32), +add (1) (p36), −2, ins (3) (p25), del (3) (p21), + add (3) (p11), −4, add (4) (p15), del (5) (q11q13)×2, add (6) (p21), +8, −9, −11, −12, −13, −15, add (15) (q25), −16, −17, −19, +mar 1, +mar2(cp29). Although direct comparison remains problematic because of the complex karyotype with several aberrant chromosomes, CGH analysis from the cytologic specimens shows in part corresponding genetic imbalances including gain of 3q and loss of X, 1p36, 3p21, 5q13, and 17.
Conventional karyotyping of one case of biphasic malignant mesothelioma (No. 15) presented loss of chromosomes 14 and 22. However, in this case CGH analysis from corresponding cytologic slides for comparison was not successful.
Comparison between CGH from Cytologic Specimens and from Fresh Frozen Tissue
In three carcinomas (Nos. 4, 9, and 11), CGH analysis was performed simultaneously from the cytologic material (Fig. 1C) and from corresponding fresh frozen tumor tissue (Fig. 1D). In comparison to the fresh frozen tumor samples, CGH from cytologic specimens demonstrated equal or more significant ratio profiles of chromosomal gains and losses as shown in Figure 4 for the imbalances on chromosomes 11 and 20 in Case 9.
DISCUSSION
In the present work, CGH results were successfully obtained in 14 of 15 cases under investigation, demonstrating strikingly the efficacy of cytologic specimens for CGH. Cell pellets from serous effusions, air-dried unstained, and MGG-stained smear preparations from effusion specimens and from fine needle aspirations as well as imprint smears are all suitable for isolation of well-preserved high molecular weight DNA. Generally, one smear preparation containing 105 to 106 malignant cells was sufficient in quality and quantity for CGH analysis without prior PCR amplification of genomic DNA. Degradation of DNA caused by formalin fixation and paraffin embedding, which frequently decreases sensitivity of molecular analysis in routine histologic material (11, 12), did not occur in the cytologic specimens.
Moreover, for CGH analysis, there is not only a considerable advantage of cytologic specimens in comparison to formalin-fixed tissue but also to fresh frozen tissue. The chromosomal imbalances detected by CGH analysis were as significant or even more significant in fine needle aspirates and imprint smears than in corresponding fresh frozen tumor tissue (Fig. 4). This difference may be caused by the more selective aspiration of tumor cells in fine needle aspirates, which prevents dilution effects from admixture with tumor-accompanying nonneoplastic stromal cells and adjacent nonneoplastic tissue (6). For instance, the fine needle aspiration smear (Fig. 1C) from mucinous breast carcinoma (Case 9) contains almost exclusively carcinoma cells embedded in a mucinous substance, whereas in the corresponding histologic section (Fig. 1D), areas of carcinoma cells are surrounded by a benign stroma tissue.
The resolution of CGH in detection of chromosomal imbalances is reported to be ≥3–10 megabases (6, 13). However, the sensitivity of detecting single copy number changes is decreased in polyploid tumors (14). Balanced structural aberrations as inversions, reciprocal translocations, or imbalances affecting only smaller regions or single mutations are not detectable by means of CGH. There are in fact only few reports on the comparison of CGH and karyotyping in solid tumors, and these deal with concordant results in most but not all cases, and usually more imbalances are detected by CGH analysis (11). Discrepancies of both methods are likely to be explained by the selection of specific tumor cell clones in cell culturing but not in material for CGH analysis. Therefore, the limited part of human solid tumors that present evaluable karyotypes by cell culturing cannot provide a representative overview of cytogenetic aberrations for the broad spectrum of tumors (7).
In this series, a comparison of CGH results with findings of classic cytogenetic techniques was expedient in one case (Case 6) of a metastatic serous papillary ovarian adenocarcinoma. Tumor cells obtained from ascites fluid were successfully obtained in short-term cell culture. The numerical and structural aberrations of DAPI-banded karyotype were in part also identified by CGH analysis in corresponding cytologic material either significantly or as borderline imbalances. This partial correspondence and the detection of additional aberrations by CGH that were not identified by karyotyping are in accordance with data from the literature (11) and confirm that CGH is a reliable and sensitive method for detecting DNA copy number changes in malignant effusions. Our experience with CGH in cytogenetic analysis of malignant effusions is that CGH is much more efficient than short-term culturing of metastatic carcinoma cells obtained from effusion specimens because of the contamination of cell cultures with proliferating mesothelial cells and fibroblasts.
From a diagnostic point of view, molecular genetic methods should provide information to support the morphologic diagnosis. In cytodiagnosis, a reliable distinction between benign reactive mesothelial proliferation, malignant mesothelioma, and metastatic adenocarcinoma requires additional techniques in many cases (1, 3).
Our series includes three cases of histologically verified malignant mesothelioma. In one case (Case 15), conventional karyotyping demonstrated loss of chromosomes 14 and 22 (data not shown), and in the others, CGH analysis revealed loss of 6q and 22q tendentiously (No. 13) as well as gain of 1p36, 16p, 17, and 19 (No. 14). Such obvious chromosomal imbalances favor the diagnosis of a benign or malignant neoplasm and argue against a reactive condition (15, 16). In addition, loss of 6q and 22q, as well chromosomes 14 and 22 represent some of the most frequent and tumor-specific genetic imbalances in malignant mesothelioma (17, 18, 19, 20, 21), thus supporting cytologic diagnosis of malignancy (22). In contrast, loss of 22q is a very uncommon CGH finding in non–small-cell lung carcinomas (8, 23), a tumor type, which must be distinguished from malignant mesothelioma.
Recently, it was demonstrated that genetic aberrations of primary tumors also frequently occur in their metastases, often combined with additional imbalances (24). This consistency in the pattern of chromosomal imbalances during tumor progression may be useful in cytology of metastatic lesions. On the one hand, detection of characteristic imbalances may result in conclusions on the site of the primary tumor (7), and on the other hand, comparison of the cytogenetic pattern in the metastatic lesion with a known primary tumor supports the relationship between these two lesions and rules out a second independent malignant tumor.
In metastatic ovarian carcinomas, which made up the majority of tumors in this series, the most frequent chromosomal gains included the long arms of chromosomes 8, 20, and 3, the most frequent losses included the long arms of chromosomes 4, 18, and 9. This pattern of chromosomal imbalances is in accordance with results of previous investigations of CGH in large numbers of primary ovarian carcinomas (14, 25, 26, 27, 28). Although these imbalances are not exclusively found in ovarian carcinomas, our findings support the hypothesis that chromosomal aberrations of the primary tumor can also be identified in its metastatic lesions. Moreover, in all metastatic ovarian carcinomas of our series, gain or high amplification of 8q24.1 was detected, as has previously been reported to indicate advanced-stage ovarian carcinomas (26). Our results confirm these data from the literature and highlight the perspectives of CGH analysis of cytologic specimens. Cytologic material from metastatic lesions as malignant effusions represent advanced-stage, generally genetically unstable tumors. Cytogenetic analysis of such tumors by means of CGH may detect genetic imbalances that will give important information on prognosis in particular types of cancer (16). Furthermore, detailed molecular genetic analysis of such unstable chromosomal sites might identify the involvement of oncogenes and tumor suppressor genes that contribute to progression and metastasis of cancer (20, 27). This hypothesis may be applicable for the myc oncogene that is located at 8q24.1 and that is amplified in ≤30% of ovarian carcinomas (26) or for the cyclin D1 gene in breast carcinomas, which is located on 11q13 (29). Cytologic specimens present a methodical advantage for molecular investigations (30), because the yield of well-preserved messenger RNA is more efficient in contrast to formalin-fixed tissue. Therefore, cytologic specimens appear to be suited for subsequent RNA expression analysis in tumors that present gene amplification sites in CGH.
In conclusion, CGH is a powerful tool for identifying chromosomal imbalances in cytologic specimens, in fine needle aspirates as well as in fluid samples. CGH analysis in cytology may help to confirm the diagnosis of malignancy, especially in the differential diagnosis of benign mesothelial proliferation, malignant mesothelioma, and metastatic adenocarcinoma in serous effusions. In cytology of metastatic lesions, the relation to primary tumors may be elucidated by the consistency of chromosomal imbalances in primary and metastatic lesions. Cytology combined with CGH analysis represents a noninvasive method for genetic studies of advanced stage tumors and involvement of specific genes in progression of cancer.
References
Motherby H, Marcy T, Hecker M, Roß B, Nadjari B, Auer H, et al. Static DNA cytometry as a diagnostic aid in effusion cytology. Anal Quant Cytol Histol 1998; 20: 153–161.
Magda JL, Minger BA, Rimm DL . Polymerase chain reaction-based detection of clonality as a non-morphologic diagnostic tool for fine-needle aspiration of the breast. Cancer Cytopathol 1998; 84: 262–267.
Dejmek A, Yahata N, Ohyashiki K, Ebihara Y, Kakihana M, Hirano T, et al. In situ telomerase activity in pleural effusions. Diagn Cytopathol 2001; 24: 11–15.
Nagel H, Schlott T, Schulz GM, Droese M . Gene expression analysis of the catalytic subunit of human telomerase (hEST2) in the differential diagnosis of serous effusions. Diagn Mol Pathol 2001; 10: 60–65.
Kallioniemi A, Kallioniemi OP, Sudar D, Rurowitz D, Gray JW, Waldman F, et al. Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 1992; 258: 818–821.
Kallioniemi O-P, Kallioniemi A, Piper J, Isola J, Waldman FM, Gray JW, et al. Optimizing comparative genomic hybridization for analysis of DNA sequence copy number changes in solid tumors. Genes Chromosomes Cancer 1994; 10: 231–234.
Gebhart E, Liehr T . Patterns of genomic imbalances in human solid tumors. Int J Oncol 2000; 16: 383–389.
Rooney PH, Murray GI, Stevenson DAJ, Haites NE, Cassidy J, McLeod HL . Comparative genomic hybridization and chromosomal instability in solid tumors. Br J Cancer 1999; 80: 862–873.
Brinck U, Gunawan B, Schulten H-J, Pinzon W, Fischer U, Füzesi L . Clear-cell odontogenic carcinoma with pulmonary metastases resembling pulmonary meningothelial-like nodules. Virchows Arch 2001; 438: 412–417.
Gunawan B, Braun S, Cortés MJ, Bergmann F, Karl C, Füzesi L . Characterization of a newly established endometrial stromal sarcoma cell line. Int J Cancer 1998; 77: 424–428.
Persson K, Pandis N, Mertens F, Borg Å, Baldetorp B, Killander D, et al. Chromosomal aberrations in breast cancer: a comparison between cytogenetics and comparative genomic hybridization. Genes Chromosomes Cancer 1999; 25: 115–122.
Bürki NG, Caduff R, Walt H, Moll C, Pejovic T, Haller U, et al. Comparative genomic hybridization of fine needle aspirates from breast carcinomas. Int J Cancer 2000; 88: 607–613.
Bentz M, Plesch A, Stilgenbauer S, Döhner H, Lichter P . Minimal sizes of deletions detected by comparative genomic hybridization. Genes Chromosomes Cancer 1998; 21: 172–175.
Jacobsen A, Arnold N, Weimar J, Kiechle M . Comparison of comparative genomic hybridization and interphase fluorescence in situ hybridization in ovarian carcinomas: possibilities and limitations of both techniques. Cancer Genet Cytogenet 2000; 122: 7–12.
Ried T, Knutzen R, Steinbeck R, Blegen H, Schrock E, Heseleyer K, et al. Comparative genomic hybridization reveals a specific pattern of chromosomal gains and losses during the genesis of colorectal tumors. Genes Chromosomes Cancer 1996; 15: 234–245.
Friedrich K, Scheithauer J, Dimmer V, Meyer W, Theissig F, Haroske G, et al. DNA ploidy and chromosomal imbalances in invasive ductal breast cancer. A comparative study of DNA image cytometry and comparative genomic hybridization (CGH). Anal Cell Pathol 2000; 20: 69–82.
Björquist A-M, Tammilehto L, Antilla S, Mattson K, Knuutila S . Recurrent DNA copy number changes in 1q, 4q, 6q, 9p, 13q, 14q and 22q detected by comparative genomic hybridization in malignant mesothelioma. Br J Cancer 1997; 75: 523–527.
Björkquist A-M, Wolf M, Nordling S, Tammilehto L, Knuuttila A, Kere J, et al. Deletions at 14q in malignant mesothelioma detected by microsatellite marker analysis. Br J Cancer 1999; 81: 1111–1115.
Balsara BR, Bell DW, Sonoda G, De Rienzo A, du Manoir S, Jhanwar SC, et al. Comparative genomic hybridization and loss of heterozygosity analyses identify a common region of deletion at 15q11. 1–15 in human malignant mesothelioma. Cancer Res 1999; 59: 450–454.
De Rienzo A, Testa JR . Recent advances in the molecular analysis of human malignant mesothelioma. Clin Ter 2000; 151: 433–438.
Krismann M, Adams H, Jaworska M, Müller KM, Johnen G . Patterns of chromosomal imbalances in benign solitary fibrous tumours of the pleura. Virchows Arch 2000; 437: 248–255.
Björkquist A-M, Tammilehto L, Nordling S, Nurminen M, Anttila S, Mattson K, et al. Comparison of DNA copy number changes in malignant mesothelioma, adenocarcinoma and large-cell anaplastic carcinoma of the lung. Br J Cancer 1998; 77: 260–269.
Michelland S, Gazzeri S, Brambilla E, Robert-Nicoud M . Comparison of chromosomal imbalances in neuroendocrine and non-small-cell lung carcinomas. Cancer Genet Cytogenet 1999; 114: 22–30.
Al-Mulla F, Keith WN, Pickford IR, Going JJ, Birnie GD . Comparative genomic hybridization analysis of primary colorectal carcinomas and their synchronous metastases. Genes Chromosomes Cancer 1999; 24: 306–314.
Izutzu T, Kudo T, Shoji T, Nishiya I . Comparative cytogenetic studies of benign, borderline and malignant epithelial ovarian tumors. J Obstet Gynaecol Res 1996; 22: 541–549.
Sonoda G, Palazzo J, du Manoir S, Godwin AK, Feder M, Yakushiji M, et al. Comparative genomic hybridization detects frequent overrepresentation of chromosomal material from 3q26, 8q24 and 20q13 in human ovarian carcinomas. Genes Chromosomes Cancer 1997; 20: 320–328.
Watson JEV, Gabra H, Taylor KJ, Rabiasz GJ, Morrison H, Perry P, et al. Identification and characterization of a homozygous deletion found in ovarian ascites by representational difference analysis. Genome Res 1999; 9: 226–233.
Suehiro Y, Sakamoto M, Umayahara K, Iwabuchi H, Sakamoto H, Tanaka N, et al. Genetic aberrations detected by comparative genomic hybridization in ovarian clear cell adenocarcinomas. Oncology 2000; 59: 50–56.
Tanner MM, Karhu RA, Nupponen NN, Borg Å, Baldetorp B, Pejovic T, et al. Genetic aberrations in hypodiploid breast cancer. Am J Pathol 1998; 153: 191–199.
Forozan F, Mahlamäki EH, Monni O, Chen Y, Vedman R, Jiang Y, et al. Comparative genomic hybridization analysis of 38 breast cancer cell lines: a basis for interpreting complementary DNA microarray data. Cancer Res 2000; 60: 4519–4525.
Acknowledgements
The authors thank Mrs. Cynthia Buncker for reviewing the manuscript and Mr. Klaus Müller for assistance in preparing graphics.
Author information
Authors and Affiliations
Corresponding author
Additional information
These data were presented in part at the 11th meeting of the Swiss, Austrian, and German Societies of Cytology, April 2–4, 2001, in Flims, Switzerland.
Rights and permissions
About this article
Cite this article
Nagel, H., Schulten, H., Gunawan, B. et al. The Potential Value of Comparative Genomic Hybridization Analysis in Effusion—and Fine Needle Aspiration Cytology. Mod Pathol 15, 818–825 (2002). https://doi.org/10.1097/01.MP.0000024521.67720.0F
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/01.MP.0000024521.67720.0F