Abstract
The chromatin pattern in nuclei from breast ductal proliferative lesions was quantitatively evaluated with the objective of deriving measures of tumor progression. A total of 110 cases were analyzed. There were 38 cases of normal tissue or benign proliferative lesions, 41 cases of ductal carcinoma in situ (DCIS), and 31 cases of microinfiltrating DCIS and of infiltrating cancer. A total of 9424 nuclei were analyzed. High-resolution images were digitally recorded. For each nucleus, 93 karyometric features descriptive of the spatial and statistical distribution of the nuclear chromatin were computed. Data analysis included establishing a profile of relative deviations of each feature from “normal,” called the nuclear signature, and of lesion signatures as well as of trends of lesion progression. Two trends of evolution could be discerned: one from normal to hyperplasia, atypical hyperplasia, and comedo DCIS as representative of high-grade lesions; and the other from normal to hyperplasia to cribriform DCIS, solid DCIS, and infiltrating cancer, representing lower grade lesions. The nuclei in microinfiltrating foci are distinctly different from nuclei in high-grade comedo DCIS. The nuclei in microinfiltrating foci have a statistically significantly lower nuclear abnormality. They may represent outgrowing clones.
Similar content being viewed by others
INTRODUCTION
Breast cancer is known to progress through multiple, genomic changes (1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11). These accumulating changes result in the nuclear alterations on which the pathologic diagnosis and microscopic prognosis of cancer are based. The natural history of ductal breast cancer is, even today, only incompletely known. The suggestion that a clonal selection (12) occurs in the progression from atypical hyperplasia toward ductal carcinoma in situ (DCIS) and invasive cancers seems to be supported by recent cytogenetic and molecular studies (1, 13, 14) and also by the characteristic morphological heterogeneity (15) of various ductal proliferative premalignant lesions. We studied alterations of the nuclear chromatin pattern concomitant with the development of different histologically defined diagnostic categories in ductal proliferative lesion progression.
Recent work led to the development of methods allowing a more extensive use of information extracted from diagnostic imagery. One such method is the characterization of individual nuclei, of sets of nuclei, and of entire lesions by a “signature.” These signatures are based on a substantial number of karyometric features, for each of which a measure of deviation from “normal” is computed. The deviation values are arranged in the form of a profile (16).
In earlier studies, these procedures were applied to images of nuclei from ductal proliferating breast lesions and DCIS (17, 18). It was shown that nuclei from a number of the conventional diagnostic categories of proliferating breast lesions representing breast cancer progression are characterized by a monotonically increasing nuclear abnormality. The histologically defined subjective diagnostic categories, ordered by relative risk for development of infiltrative disease, thus find correspondence and confirmation in an objective, numerically defined measure of progression based on an increasingly abnormal nuclear chromatin pattern.
An unexpected finding was that the lesion signatures, formed from the distribution of nuclear abnormality values, varied widely within a given diagnostic category. Lesion signatures reveal the diversity existent within any given diagnostic category, in a quantifiable manner, as a measure of the extent of nuclear abnormality for each case. This finding may well have prognostic significance because it provides a reproducible, objective measure to assess the relative risk for each individual case.
In this study, a search for “structure” in the recorded data sets is conducted in an effort to document the existence of different pathways for the development of high-grade and low-grade lesions.
MATERIALS AND METHODS
The clinicopathological materials for this study were collected at the Institutes of Pathology at the University of Verona and of Udine, Italy, and at the Department of Pathology, University of Arizona. Only cases for which complete diagnostic agreement between three expert histopathologists was established were included, following the diagnostic criteria recently suggested by the European Pathologist Working Group (18) and the Van Nuys group (19). To represent the main steps in ductal breast cancer progression, eight cases of normal breast glandular tissue (postmenopausal women without hyperplastic lesions or breast cancer) were selected (499 nuclei); there were 11 cases of simple hyperplasia (800 nuclei); 10 cases of florid hyperplasia (1003 nuclei); 12 cases of atypical hyperplasia (896 nuclei); 9 cases of cribriform DCIS, Group 1, low nuclear grade (DIN1; 1080 nuclei); 17 cases of solid, Group 2, intermediate nuclear grade (DIN2) and 4 cases of high grade for a total of 2021 nuclei; and 8 cases of comedo DCIS, high nuclear grade (DIN3; 625 nuclei). There were 5 cases of microinfiltrating cancer (200 nuclei) and 26 cases of ductal, invasive cancer (2300 nuclei). Lesions were selected on the basis of the histologic pattern.
In selecting cases for the diagnostic categories, cases were chosen in which no lesion of higher progression was present. For example, in the selection of DCIS cases, those with microinvasive foci were not included. For the category of ductal comedo microinvasive cancer, cases for which the foci of microinvasion were > 1 mm in diameter were excluded.
Sections were cut to 5-μm thickness and stained with a routine hematoxylin and eosin (H&E) procedure under carefully controlled conditions.
Digital imagery was recorded first on a videophotometer equipped with a 20:1, N.A. 0.75 apochromatic flat field objective by NIKON (Melville, NY) and by a three-chip color CCD camera from Sony (Park Ridge, NJ). From this imagery, several videoframes of each lesion were seamlessly merged to cover a diagnostically representative area (19). The merged images were then laser printed and subregions outlined in which individual nuclei were randomly selected for measurement. These fields and nuclei were relocated on a high-resolution videophotometer for measurement. This instrument is equipped with a 63:1, 1.40 N.A. oil immersion planapochromatic objective by Zeiss (Oberkochen, Germany) and a COHU (San Diego, CA) black and white video camera. The relay optics adjusted image sampling to six pixels per linear micron. An interference filter with a maximum band-pass of 610 nm was used to enhance contrast of the H&E sections. The nuclear images were edited using an interactive procedure and filed for feature extraction.
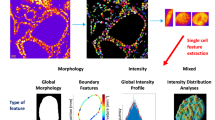
For each nucleus, a set of 93 karyometric features was computed. These features include global features: total optical density (O.D.), nuclear area, variance of pixel O.D. values: 3 features, histogram of pixel O.D. values, 0.10 O.D. intervals: 18 features; co-occurrence features, six O.D. intervals 0.30 O.D. units wide, upper diagonal of the co-occurrence matrix only 21 features: run length features, six O.D. intervals 0.30 O.D. units wide, six run length intervals from 1 to 2 pixels to 11 to 12 pixels: 36 features; and a number of summarizing features, such as run length emphasis, pixel O.D. histogram shape: 15 features.
To form the nuclear signature, data from a reference set of normal nuclei were used. For each feature, the mean and standard deviation was computed. Then, for any nucleus, the deviation from that mean value was computed as an absolute value and divided by the standard deviation from the reference set value for that feature. The deviation from normal thus is expressed as a standardized distance, or z value. The nuclear signature is formed by arranging the z values for the 93 features in an arbitrary but consistent order.
The nuclear signatures offer very detailed information. It is useful to employ a number of summarizing procedures. One may form an average over the 93 z values. The result is a single number that is also referred to as averaged standardized distance but more simply as nuclear abnormality.
The distribution of such nuclear abnormality values for all nuclei measured in a given lesion is called a “lesion signature.” One may also form an average nuclear signature for a set of nuclei, either from a given case or averaged over all cases from a given diagnostic category. In the same way, one may average the lesion signature over all cases from a given diagnostic category.
RESULTS
To demonstrate the relationship between the nuclear chromatin pattern and the nuclear abnormality, Figure 1 shows a representative image of a nucleus from normal breast epithelium, from cribriform DCIS, from solid DCIS, and from comedo DCIS, with the nuclear abnormality value for the nuclei shown, and their nuclear signatures.
Plotting the nuclear abnormality values averaged within each of the diagnostic categories (i.e., over all nuclei and all cases in the category) as a function of diagnostic category, a monotonically rising trend is seen in Figure 2, from normal glandular epithelium, simple hyperplasia, cribriform DCIS, and solid-pattern DCIS, to infiltrating cancer. Within these diagnostic categories, there is some variance in the case-averaged nuclear abnormality. Generally, though, the nuclear abnormality values fall into the range from 0.5 standard deviations from normal to 1.5 standard deviations.
For cases of florid hyperplasia, atypical hyperplasia, and comedo DCIS, a much greater nuclear abnormality is observed, along with greater case-to-case variance (Fig. 3), compared with the range of abnormality values found in cribriform DCIS, solid DCIS, and infiltrating carcinoma. There are some cases of solid DCIS diagnosed as high-grade solid DCIS that exceed that range.
Nuclear abnormality values for lesions falling into a range of higher deviation from normal. The region of nuclear abnormality values for lesions in a lower range, from Figure 2, is outlined in the lower portion of the plot.
In Figures 2 and 3, the diagnostic categories were arbitrarily arranged along the abscissa at an equal spacing, in the rank order of relative risk for the development of infiltrating disease. Such spacing may distort the true structure of the data sets. A plot based on two numerically defined measures is preferable. For the abscissa, the nuclear area was chosen for two reasons. The difference in nuclear area between nuclei from normal epithelium and nuclei, such as from comedo DCIS, is substantial. The feature provides a good spread. Figure 4 shows a direct comparison of two representative nuclei. Also, all sections for this study had been cut to 5 μm. Total optical density would have been a preferred choice, as it relates to ploidy—but it might be compromised in a nonlinear fashion by the sectioning of large-size nuclei.
The nuclear area undergoes a steady increase from normal epithelium to comedo DCIS. This is shown in Table 1.
Figure 5 shows a plot of the diagnostic category–averaged nuclear abnormality values versus the nuclear area. Again, there is a clear indication of a dual trend in the development of the proliferating lesions.
Plot of nuclear abnormality versus nuclear area, averaged over all cells and all cases in each diagnostic category. Two trends of lesion progression are in evidence. Also note that the nuclei from microinfiltrating comedo ductal carcinoma in situ (DCIS)are distinctly different from the nuclei found in the comedo DCIS lesion.
There seems to exist a bimodal distribution of the comedo DCIS nuclear abnormality: almost half of all cases have an average nuclear abnormality of more than three standard deviations; the other half exhibits values only slightly above the values seen in noncomedo DCIS and in infiltrating cancer. This is shown in Figure 3.
The distinction between lesions of low grade and those of high deviations from normal is expressed already at the level of hyperplastic lesions.
The nuclear signatures for nuclei from normal breast epithelium, simple hyperplasia, and atypical hyperplasia are shown in Figure 6. Also shown for comparison is the nuclear signature for comedo DCIS. The standard deviations for all karyometric features in normal epithelium are less than one standard deviation for all features. In simple hyperplasia, there are modest increases in the deviations from normal for a few features. The total optical density is slightly increased, and for example, the pixel optical density histogram is changing.
In atypical hyperplasia, drastic changes are seen. The nuclear area increases, and the pixel O.D. features, the co-occurrence features, and the run length features all begin to show high deviations.
The nuclear signatures of comedo DCIS expressed the very same changes, in a more pronounced fashion. The changes seen in atypical hyperplasia thus are similar to those seen in comedo DCIS.
Figure 7A shows the lesion signatures, averaged over all cases of hyperplasia, falling into the range of nuclear abnormality demarcating a lower grade pathway in Figure 2. Figure 7B shows the lesion signatures averaged over all cases from simple hyperplasia, florid hyperplasia, and atypical hyperplasia, with a nuclear abnormality in excess of 1.2 standard deviations for the case average. The distribution of nuclear abnormality values for the comedo DCIS is seen in Figure 7C, for comparison.
There is an indication of a bimodality in the distributions for the high-grade hyperplasias as well as for the comedo DCIS cases.
The high degree of nuclear abnormality in the higher grade hyperplastic lesions is not associated with increased aneuploidy. Table 2 shows the mean values for the total optical densities. The total optical density values seen in atypical hyperplasia fall into the same range as those from simple hyperplasia. The values found in comedo DCIS lesions are about twice as large as those in a normal diploid cell population. In comedo DCIS, the total optical density distribution shows a broadly spread pattern filling the range between the total optical densities the corresponds with 2 N to 5 N ploidy values and extending up into the 7 N and higher range. Thus, there is a definite indication of aneuploidy.
The substantial differences in nuclear size suggested that for the chosen section thickness of five μm, the total optical density values might, for the diagnostic categories with very large nuclei, underestimate “ploidy.” This is unquestionably true. However, if the total optical density distribution is found to extend well into the high optical density range, as was the case here, one may safely deduce that an aneuploid cell population is present.
The nuclei measured in foci of microinvasion are distinctly different from nuclei in comedo DCIS. The average nuclear abnormality is lower in all five measured cases and falls into the range of the trend for low-grade DCIS lesions, as seen in Table 1. Figure 8 shows the respective nuclear signatures.
To document these differences, an analysis of variance (ANOVA) was carried out for a randomly selected subset of five cases each of comedo DCIS and five cases of microinfiltrating comedo DCIS, with 40 nuclei per case, to maintain a balanced design. Table 3 shows the ANOVA results. A two-level nested design was chosen, with diagnostic category as the first, fixed level and with cases as the second, random level.
The difference in nuclear abnormality between diagnostic categories is highly significant and allows for 92.7% of the total mean square, with 7.1% and 0.2% accounting for case to case and nucleus to nucleus variability.
DISCUSSION
The characterization of nuclei by a detailed evaluation of the chromatin pattern has allowed definition of a numeric measure of progression, for nuclei and for lesions, in terms of deviation from normal. A preceding study (17) has shown that the histologically established ordering of proliferating breast lesions and DCIS can be confirmed by a monotonically rising numeric measure of deviation from normal for nuclei from different diagnostic categories. In this current study, the analysis leads in a different direction, providing possible clues to the precursor relationship between these lesions and to the changes that characterize the start of microinvasion.
Although the deviations in the nuclear chromatin pattern are monotonically increasing and continuous, they seem to follow two different trends of progression. From these results, it appears that categories of cribriform/solid DCIS and comedo DCIS represent different subtypes of breast in situ cancers, expressing different evolutionary changes. They may correspond to the dual trend for breast cancer progression suggested by Ellis et al. (20).
The recent publications of many different subclassifications of DCIS (15, 21, 22, 23, 24, 25, 26) reflect an effort to have a reproducible system of grading. These subclassifications of DCIS are overall based on nuclear criteria that are subjectively evaluated within the limits of visual subjective assessment.
Karyometric data, such as the nuclear signatures, provide objective measures reflecting the continuity of change in both trends of progression. They allow an objective determination of the degree of progression for each individual case. The objective evaluation offers an option to define a system of continuous grading suitable for individual patient-targeted prognosis.
The data structure revealed by the quantitative evaluation of karyometric features from nuclei in proliferative breast lesions suggests the existence of at least two distinctly different pathways of progression, representing lower and high-grade lesions. There is some evidence that the nuclei from microinvasive foci represent a different clone from the nuclei forming the majority of nuclei in comedo DCIS lesions, a clone capable of starting stromal invasion. These nuclei—either newly arising from highly heterogeneous, polyclonal aneuploid, and atypical cells of in situ cancers, or finally gaining a growth advantage and becoming noticeable—show phenotypical homogeneity. It seems likely that the more atypical cells of the highly heterogeneous cell population, with their numerous chromosomal and genomic abnormalities, might not be able to give rise to a new and more aggressive cell population.
Previous experience (18, 27, 28) with cervical and breast cancer has shown that in tissue sections, the homogeneous populations of microinfiltrative foci usually have cells with near-diploid DNA contents and without prominent evidence of somatic abnormalities.
This is also strongly reminiscent of the changes in the nuclear chromatin pattern seen in nuclei from proliferating lesions in prostatic intraepithelial neoplasia (PIN). The chromatin pattern in nuclei from high-grade PIN undergoes a drastic change in nuclei measured in the region of microinvasion (16). Also, in these cases, it was possible to show that the nuclei in the region of microinvasion were of lower ploidy than the highly abnormal nuclei of the high-grade PIN lesions.
One must expect that the nuclei measured in the preinfiltrating lesions do not constitute homogeneous cell populations. Rather, there might be nuclei representing two or more different phenotypes, as has been found in endometrial and in prostatic lesions. The proportional composition of the phenotypes in a given case would greatly affect the case-averaged nuclear abnormality. This, in turn has the effect of increasing the variance of that measure and consequently might mask the existence of even more pathways than the low-grade and high-grade lesions demonstrated above. The slight bimodality in the distribution of nuclear abnormality values in high-grade hyperplasia and in comedo DCIS may point to such additional pathways.
The interpretation of data given above is compatible with a model of lesion progression in which a succession of clones evolve, some of which gain a growth advantage and persist, whereas others may not continue.
The trend or “progression” shown in Figures 2 and 3 may suggest biologic behavior but should not conclusively be interpreted in this manner. The data merely demonstrate the monotonic and continuous rise in nuclear abnormality. This is to be seen as independent of the histologic pattern, on which histopathologic diagnosis is based and correlated with risk for infiltrating disease. It is, in this model, quite possible for any premalignant lesion to progress directly to infiltrating disease, and it is quite possible for any lesion not to progress at all.
References
Stratton MR, Collins N, Lakhani SR, Sloane JP . Loss of heterozygosity in ductal carcinoma in situ of the breast. J Pathol 1995; 175: 195–201.
Paulson TG, Wright FA, Parker BA, Russack V, Wahl GM . Microsatellite instability correlates with reduced survival and poor disease prognosis in breast cancer. Cancer Res 1996; 56(17): 4021–4026.
Porter PL, Malone KE, Heagerty PJ, Alexander GM, Gatti LA, Firpo EJ, et al. Expansion of cell-cycle regulators p27/Kip1 and cyclin E, alone and in combination, correlate with survival in young breast cancer patients. Nat Med 1997; 3(2): 222–225.
Sourvinos G, Kianis H, Tsikkinis A, Vassilanos S, Spardidos DA . Microsatellite instability and loss of hetrozygosity in primary breast tumor. Tumor Biol 1997; 18(3): 155–166.
Valgarsdottir R, Tryggvadottir L, Steinarsdottir M, Olafsdottir K, Jonasdottir S, Jonasson JG, et al. Genomic instability and poor prognosis associated with abnormal TP53 in breast carcinomas. Molecular and immunohistochemical analysis. APMIS 1997; 105(2): 121–130.
Wolman SR, Heppner GH, Wolman E . New directions in breast cancer research. FASEB J 1997; 11: 535–543.
Wang BL, Springer GF, Carlstedt SC . Quantitative computerized image analysis of Tn and T (Thomsen-Friedenreich) epitopes in prognostication of human breast carcinoma. J Histochem Cytochem 1997; 45: 1393–1400.
Albertazzi E, Cajone F, Leone BE, Nagrib FN, Lakshani MS, Sherbet GV . Expression of metastasis-associated genes h-mts 1 (S100A4) and nm23 in carcinoma of breast is related to disease progression. DNA Cell Biol 1998; 17(47): 335–342.
Burger AM, Zhang X, Li H, Ostrowski JL, Beatty B, Venanzoni M, et al. Down-regulation of T1A12/mac25, a novel insulin-like growth factor binding protein related gene, is associated with disease progression in breast carcinomas. Oncogene 1998; 16: 2459–2467.
Tsukamoto Y, Kuwabara K, Hirota S, Kawano K, Yoshikawa K, Ozawa K, et al. Expression of the 150-kd oxygen-regulated protein in human breast cancer. Lab Invest 1998; 78(6): 699–706.
Watson BL, Marcus JN, Lynch HT . Prognosis of BRCA1 hereditary breast cancer. Lancet 1998; 351: 304–305.
Nowell PC . The clonal evolution of tumor cell population. Science 1976; 194: 23–38.
Poller DN, Roberts EC, Bell JA, Elston CW, Blamey RW, Ellis IO . p53 protein expression in mammary ductal carcinoma in situ: relationship to immunohistochemical expression of estrogen receptor and c-erbB-2 protein. Hum Pathol 1993; 24: 463–468.
Bobrow LG, Happerfield LC, Gregory WM, Springall MD, Millis RR . The classification of ductal carcinoma in situ and its association with biological markers. Semin Diagn Pathol 1994; 11: 199–207.
Lennington WJ, Jensen RA, Dalton LW, Page DL . Ductal carcinoma in situ of the breast: Heterogeneity of individual lesions. Cancer 1994; 73: 118–124.
da Silva V, Montironi R, Thompson D, Bartels HG, Vaught L, Hamilton PW, et al. Chromatin texture in high grade prostatic intraepithelial neoplasia and early invasive carcinoma. Anal Quant Cytol Histol 1999; 21(2): 113–120.
Mariuzzi L, Mombello A, Rucco V, Morelli L, Zamo A, Thompson D, et al. Quantitative study of ductal breast cancer progression: signatures of nuclei in proliferating breast lesions and in situ cancers. Adv Clin Pathol 2000; 4: 87–97.
Frank DH, Davis JR, Alberts DS, Thompson D, Bartels PH . Nuclear chromatin characteristics in breast solid pattern ductal carcinoma in situ (DCIS). Anal Quant Cytol Histol (in press).
Thompson D, Richards D, Montironi RM, Scarpelli M, Ham-ilton PW, Bartels PH . Multimegapixel images in histopathology. Anal Quant Cytol Histol 2001; 23: 169–177.
Ellis IO, Pinder SE, Lee AH, Elston CW . A critical appraisal of existing classification systems of epithelial hyperplasia and in situ neoplasia of the breast with proposals for future methods of categorization: where are we going? Semin Diagn Pathol 1999; 16: 202–208.
Ottensen GL, Graversen HP, Blichert TM, Zedeler K, Andersen JA . Ductal carcinoma in situ of the female breast. Short-term results of a prospective nationwide study. The Danish Breast Cancer Cooperative Group Am J Surg Pathol 1992; 16: 1183–1196.
Bellamy CO, McDonald C, Salter DM, Chetty U, Anderson DJ . Noninvasive ductal carcinoma of the breast: the relevance of histologic categorization. Hum Pathol 1993; 24: 16–23.
Lagios MD . Heterogeneity of ductal carcinoma in situ of the breast. J Cell Biochem 1993; 17G(Suppl): 49–52.
Holland R, Peterse JL, Millis RR . Ductal carcinoma in situ: a proposal for a new classification. Semin Diagn Pathol 1994; 11: 167–180.
Poller DM, Silverstein MJ, Galea M . Ideas in pathology. Ductal carcinoma in situ of the breast: a proposal for a new simplified histological classification association between cellular proliferation and c-erbB-2 protein expression. Mod Pathol 1994; 7: 257–262.
Silverstein MJ, Poller DN, Waisman J, Colburn WJ, Barth A, Gierson ED, et al. Prognostic classification of breast ductal carcinoma in situ. Lancet 1995; 345: 1154–1157.
Maraldi NM, Marinell F, Ranoldi R, Papa S, Mariuzzi GM, Manzoli FA . Morphometric and topologic analysis of freeze-fractured interphase nuclei. Anal Quant Cytol Histol 1986; 8: 343–348.
Marinelli F, Squarzoni S, Maraldi NM, DiLoreto C, Rubini C, Mariuzzi GM, et al. Morphometric study of chromatin pattern in freeze-fractured rat liver nuclei during malignancy evolution. Pathol Res Pract 1989; 185: 769–773.
Acknowledgements
This research was supported in part by a grant from the National Institutes of Health, National Cancer Institute, CA 53877 (to PHB) and in part by grants from AIRC (Milan, Italy) and MURST (Verona, Italy) to GMM.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mariuzzi, L., Mombello, A., Granchelli, G. et al. Quantitative Study of Breast Cancer Progression: Different Pathways for Various In Situ Cancers. Mod Pathol 15, 18–25 (2002). https://doi.org/10.1038/modpathol.3880485
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3880485
Keywords
This article is cited by
-
Molecular profile of atypical hyperplasia of the breast
Breast Cancer Research and Treatment (2018)
-
Quantitative Image Analysis in Mammary Gland Biology
Journal of Mammary Gland Biology and Neoplasia (2004)