Abstract
The polymerase chain reaction (PCR) analysis of DNA extracted from tissue sections can be applied to a variety of research and diagnostic protocols. To analyze selectively the specific areas of tissue, a direct microdissection of histochemically or immunohistochemically stained sections, if satisfactory for PCR, is helpful. However, the influence of various staining methods on PCR has been poorly investigated. In this study, paraffin sections of formalin-fixed lymph node samples were histochemically stained with Mayer’s hematoxylin, eosin Y, methyl green, or May-Grunwald solution and immunostained for CD45 using 3,3′-diaminobenzidine (DAB), DAB with cobalt ion (DAB-Co), or new fuchsin as the chromogen. In addition, unstained sections were treated with trypsin, microwave, or pressure cooker, the techniques frequently used in immunostains for antigen unmasking. DNA was extracted from each section, and the PCR efficiency in amplifying a 110 bp portion of the beta-globin gene was evaluated by two parameters: the cycle count in which the first visible band was obtained (CYCLEmin) and the maximum amount of PCR products (CONCmax). The hematoxylin stain showed a significantly prolonged CYCLEmin (P <.01) and lower CONCmax (P <.05) in comparison with unstained and untreated control sections. The May-Grunwald stain showed a prolonged CYCLEmin (P <.01), although the CONCmax was not significantly different from that of the control (P =.051). The eosin and methyl green stains showed no effects against PCR. In immunostains, the DAB-Co method showed a lower CONCmax (P <.05), whereas the CYCLEmin was not prolonged. The DAB and new fuchsin methods had no untoward effects. Antigen-unmasking treatments showed deteriorating effects on PCR. The trypsin treatment significantly prolonged the CYCLEmin (P <.01), and the PCR amplification did not reach the “plateau” level with a maximum of 60 cycles. The PCR efficiency was worse in microwave or pressure cooker treatment, with neither CYCLEmin nor CONCmax being obtained. When target areas from sections for subsequent PCR amplification are microdissected, methyl green is most suitable as a dye for nuclear staining. The immunohistochemical visualization with DAB or new fuchsin yields no unfavorable effects. A successful PCR amplification may not be expected in sections that are pretreated in a microwave oven or pressure cooker.
Similar content being viewed by others
INTRODUCTION
Polymerase chain reaction (PCR) used for tissue sections as a source of DNA plays an important role in pathologic research and diagnosis (1, 2, 3, 4, 5, 6, 7). To analyze selectively the specific areas of interest, microdissection of the sections on glass slides usually has to be performed. Now the combination of microdissection and PCR can be applied to even a single cell (e.g., Hodgkin or Reed-Sternberg cells in malignant lymphoma sections) (4, 5, 6, 7).
In microdissecting such small targets, an accurate morphologic identification is necessary, where the histochemical or immunohistochemical stain is of help. Tissue samples from hematoxylin and eosin–stained sections, however, yield an unsatisfactory PCR amplification (8, 9). Although various dyes, including Wright’s solution instead of hematoxylin, have been applied for this purpose (8), correlation between each dye and efficiency of PCR remains largely unknown. The conventional phenol/chloroform purification of the DNA template may decrease the influence of the various inhibitors (10, 11, 12); however, a considerable amount of DNA may be lost during the purification (10).
In the present study, we evaluated in more detail the effects against PCR amplification of histochemical dyes commonly used in microdissection (Mayer’s hematoxylin, eosin Y, methyl green, and May-Grunwald solution). We also performed a similar analysis on the signal visualizing methods by immunohistochemistry (peroxidase-diaminobenzidine [DAB], peroxidase-DAB-cobalt [DAB-Co], and alkaline phosphatase–new fuchsin). To our knowledge, no such influence of immunostains on PCR has been studied. Furthermore, we examined the influence of signal-enhancing or antigen-unmasking methods that frequently are used in modern immunohistochemistry (trypsin treatment, microwave heating, and high-temperature treatment by pressure cooker).
MATERIALS AND METHODS
Tissue Samples
Six paragastric lymph nodes of similar size (approximately 1 cm in diameter) were collected from six patients at surgery for gastric cancer. These nodes were fixed in 10% buffered formalin for 12 to 72 hours and embedded in paraffin. The tissue sections (3 μm thick) were deparaffinized and rehydrated in distilled water. The hematoxylin and eosin stain showed a normal nodal architecture with no primary or metastatic tumors in each lymph node. All of the procedures used in this study were performed with ample precautions for avoiding cross-contaminations.
Histochemical Stains, Immunohistochemical Stains, and Antigen-Unmasking Treatments
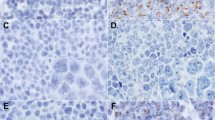
For histochemical stains, the dewaxed sections were stained with each of the following dyes of usual concentration: 1.0% w/v Mayer’s hematoxylin, 0.5% w/v eosin Y, 0.5% w/v methyl green, and May-Grunwald solution (containing 0.25% w/v nonoxidized methylene blue and eosin Y). When appropriate contrast was obtained after incubation for 10 to 30 min, the sections were rinsed in distilled water and then air dried. For immunohistochemistry, the dewaxed sections were stained for CD45 (leukocyte common antigen), because most of the cells in the lymph node express this antigen. The signals were visualized using one of the DAB, DAB-Co, and new fuchsin methods. Briefly, the sections were treated with 3% hydrogen peroxide for 15 min and reacted with anti-CD45 antibody (2B11 + PD7/26; DAKO, Kyoto, Japan) for 1 hour in a humid chamber at room temperature, followed by incubation with biotinylated antimouse immunoglobulins (Nichirei, Tokyo, Japan). Then the reaction products were visualized by each of the following methods. In the DAB method, the slides were reacted with peroxidase-conjugated streptavidin (Nichirei, Tokyo, Japan), and the brown signals were developed by immersing slides in the DAB solution containing 0.2 mg/mL DAB and 0.003% H2O2 (13). In the DAB-Co method, using cobalt ion to intensify the DAB signals, 0.02% w/v CoCl2 was added to the above DAB solution; the reaction with peroxidase resulted in black color (14). In the new fuchsin method, the slides were incubated with streptavidin-alkaline phosphatase, and the red signals were developed with new fuchsin solution (0.5 mL of a 5% solution of new fuchsin in 2N HCl added to 1.25 mL of a freshly prepared 4% solution of sodium nitrite) (15). After developing, each slide was rinsed in distilled water and air dried without counterstaining. To evaluate the effect of the antigen-unmasking pretreatments, dewaxed sections were treated with three different methods as follows: incubation in 0.1% w/v trypsin in phosphate buffered saline for 10 min at 37° C, microwave treatment for 10 min with the sections in citrate buffer (0.01 m, pH 6.0), and high-temperature heating in a pressure cooker by dipping slides in pure water for 4 min after the water boiled.
DNA Extraction and PCR
The above sections as well as unstained and untreated (control) sections from six lymph nodes were totally scraped off the slides and subjected to DNA extraction. Each section was digested in 0.05 m Tris buffer (200 μl) containing 200 μg/mL proteinase K (Boehringer Mannheim, Stuttgart, Germany) at 55° C overnight, then heated at 95° C for 10 min to inactivate proteinase K (16). The influence of each staining method on PCR was evaluated by amplifying a 110 bp portion of beta-globin gene (17). Subjected to PCR were eight tubes per section, containing 25 μl solution with a mixture of 10 mm Tris-HCl (pH 8.4), 50 mm KCl, 0.01% gelatin, 1.5 mm MgCl2, 0.2 mm dNTPmix, 1.0 μm each primer, 0.05 unit per microliter of Taq GOLD DNA polymerase (Perkin-Elmer, New York, NY), and 5 μl extracted DNA. The sequences of the primers for the complementary portions were 5′-ACACAACTGTGTTCACT-AGC-3′ and 5′-CAACTTCATCCACGTTCACC-3′, respectively (17). The samples were incubated at 95° C for 10 min to activate Taq GOLD DNA polymerase, then PCR was performed for each tube from 25 to 60 cycles at 5-cycle intervals; the cycle condition was for 1 min at 95° C, 1 min at 55° C, and 1 min at 72° C. One μl of PCR product in each tube was electrophoresed through 2% agarose gel containing ethidium bromide with a size and concentration marker. The PCR efficiency was evaluated by two parameters: the cycle count at which the 110 bp PCR product was first visible (CYCLEmin) and the maximum concentration of the PCR band (CONCmax). The CONCmax was obtained when the amount of PCR product reached its “plateau” state. The gel images were digitized (Fluor-S MultiImager & Multi-Analyst, Bio-Rad Laboratories, Melville, NY), and the concentrations of the bands were quantified using NIH image software (version 1.57; Macintosh, Cupertino, CA). Correlation between the intensity of the bands and the concentration of the PCR product in the range from 0.01 to 1.50 μg/μl was estimated, and the intensity was proportionate to the concentration. The PCR amplification at each five-cycle interval in each specimen was performed in duplicate, resulting in occasional minor inconsistency. The examination was repeated in such cases, and the random artifacts did not affect the values of CYCLEmin or CONCmax. In statistical analysis, paired t test was used, and P <.05 was considered to be significant.
RESULTS
The PCR products in each method in a representative sample are illustrated in Figure 1, and the average CYCLEmin and CONCmax values of six samples in each method are shown in Table 1. In the control samples from the unstained and untreated sections, the visible 110 bp beta-globin band first appeared at 30 to 45 cycles of PCR amplification, and the mean ± standard error (SE) of the CYCLEmin was 38 ± 2.5 cycles. The concentration of the PCR product reached the maximum within the next 10 cycles and showed a plateau thereafter. This CONCmax ranged from 0.33 to 1.05 μg/μl (0.61 ± 0.11 μg/μl).
The influence of the stains and treatments on the 110 bp band of beta-globin gene in polymerase chain reaction from 25 to 60 cycles at 5-cycle intervals (a representative sample). The DNA was extracted from the sections of unstained and untreated control (A), Mayer’s hematoxylin stain (B), eosin Y stain (C), methyl green stain (D), May-Grunwald stain (E), 3,3′-diaminobenzidine stain (F), 3,3′-diaminobenzidine with cobalt ion stain (G), new fuchsin stain (H), trypsin treatment (I), microwave treatment (J), and pressure cooker treatment (K). Note the delayed appearance of visible bands in B, E, I, and J; lower maximum concentration in B and G; and unobtainable bands in K. M, size and concentration marker.
Influence of Histochemical Stains
The CYCLEmin in the samples from the hematoxylin- and May-Grunwald–stained sections ranged from 35 to 50 cycles (43 ± 2.1 cycles) and from 35 to 50 cycles (43 ± 2.8 cycles), respectively, showing that more PCR cycles were required to obtain the first visible band in the samples with these stains than in the control (P <.01). In contrast, the CYCLEmin of the eosin- and methyl green–stained samples was similar to that of the control. The CONCmax of the hematoxylin-stained samples ranged from 0.20 to 0.45 μg/μl with an average of 0.27 μg/μl, a significantly lower value than that of the control (P <.05). The CONCmax in May-Grunwald–stained samples ranged from 0.20 to 1.07 μg/μl with an average of 0.50 μg/μl, showing lower values of marginal significance (P =.051). In eosin- and methyl green–stained samples, the CONCmax for each was only slightly lower than the control value, with no significance.
Influence of Immunostain Chromogens
The CYCLEmin was not extended in the immunostained samples developed with DAB or new fuchsin, whereas that in the DAB-Co–stained samples ranged from 30 to 50 cycles, a slight extension with no statistical significance. The CONCmax in DAB-Co samples ranged from 0.09 to 0.79 μg/μl, with the mean value significantly lower than that of the control (P <.05). The CONCmax in DAB or new fuchsin–stained samples was similar to that in the control.
Influence of Antigen-Unmasking Treatments
The trypsin, microwave, and pressure cooker methods had a considerably harmful effect on PCR. The CYCLEmin in the trypsin-treated samples ranged from 45 to 60 cycles, significantly greater than that in the control samples (P <.01). The intensity of the band in these samples was increased according to the cycles of PCR in two cases but remained weak in others at 60 cycles in the present setting; therefore, the mean CONCmax was not evaluated. No visible bands were obtained by 60 cycles of PCR in two and five of six samples by the microwave and pressure cooker treatments, respectively, leaving the CYCLEmin as well as the CONCmax undetermined.
DISCUSSION
The present study showed that both histochemical and immunohistochemical stains and antigen-unmasking treatments of sections had more or less adverse effects on PCR DNA amplification. We evaluated the PCR efficiency by CYCLEmin and CONCmax. Using these two parameters, we could assess the influence of dyes or treatments more precisely. In previous studies, PCR efficiency was evaluated by measuring only one point—the concentration of the product after 30 cycles of PCR amplification (8, 9). The results obtained by this approach should be interpreted carefully because there is no information on whether the measured point represents the maximum concentration of PCR product at the plateau level or a concentration at the exponentially increasing curve.
Of the four histochemical stains examined, those by eosin Y and methyl green showed the least inhibiting effect with the CYCLEmin and CONCmax comparable to the control values. Conversely, the hematoxylin and May-Grunwald stains considerably inhibited the PCR amplification. The untoward effect on PCR by the hematoxylin stain has been described, and two possible explanations are proposed: the dye may bind to DNA and subsequently interfere with proteinase digestion (8, 18, 19), or it might influence divalent cation (Mg++) concentration that is important in maintaining the Taq polymerase activity (18, 20). Because the present CYCLEmin may depend on either the state of template DNA or the process of PCR amplification and the CONCmax may depend predominantly on the latter, the lower CYCLEmin and CONCmax values in the hematoxylin-stained samples in this study suggest that both mechanisms may be responsible. In the May-Grunwald solution, the nonoxidized methylene blue may have a harmful effect on template DNA because only the CYCLEmin but not the CONCmax was impaired in the samples by this stain. However, the methyl green dye also reported to bind to DNA directly (21, 22, 23, 24, 25) did not affect the PCR amplification.
PCR was successfully performed with the samples where the cell membrane was immunostained and the signals were developed using DAB or new fuchsin as a chromogen. In these methods, the CYCLEmin and CONCmax were almost identical with those of the control. When DAB signals were intensified with CoCl2, the subsequent PCR gave a significantly lower CONCmax (75% of the control), although CYCLEmin was not prolonged. The reason may be that excess cobalt ion (Co++) in the PCR solution might interfere with optimization of Mg++ concentration. The influence of immunostains of various antigens in the cytoplasm and nucleus remains to be elucidated.
The antigen-unmasking treatments—trypsin, microwave, and pressure cooker—resulted in an unsatisfactory PCR. After each treatment, the CONCmax was frequently not obtained in the setting of 60 cycles of PCR. Especially, the microwave and pressure cooker treatments gave deteriorating effects. In the process of these two treatments, most of the DNA in the tissue sections might be highly degraded on exposure to the high temperature, resulting in the unobtainable CYCLEmin. The trypsin pretreatment still can be used, although the CYCLEmin was markedly delayed and the amount of the final PCR product was not satisfactory at 60 cycles. A considerable amount of the DNA may drop out of the section during trypsin digestion.
In conclusion, we suggest that methyl green may be the most suitable dye for the nuclear staining in a direct microdissection of the targeted areas. Although the eosin stain also did not impair PCR amplification, the morphologic details are hard to recognize by this stain alone. DAB or new fuchsin in immunostains of the membrane antigen gave almost no inhibiting effects on PCR. The pretreatments for antigen retrieval were unsatisfactory for PCR. When such a treatment is mandatory for immunohistochemistry, only the trypsin treatment may be applicable. In the latter case, additional PCR or nested PCR may be helpful to obtain a sufficient amount of PCR product.
References
Crisan D, Cadoff EM, Mattson JC, Hartle KA . Polymerase chain reaction: amplification of DNA from fixed tissue. Clin Biochem 1990; 23: 489–495.
Ben-Ezra J, Johnson DA, Rossi J, Cook N, Wu A . Effect of fixation on the amplification of nucleic acids from paraffin-embedded material by the polymerase chain reaction. J Histochem Cytochem 1991; 39: 351–354.
Sukpanichnant S, Vnencak-Jones CL, McCurley TL . Detection of clonal immunoglobulin heavy chain gene rearrangements by polymerase chain reaction in scrapings from archival hematoxylin and eosin-stained histologic sections: implications for molecular genetic studies of focal pathologic lesions. Diagn Mol Pathol 1993; 2: 168–176.
Trümper LH, Brady G, Bagg A, Gray D, Loke SL, Griersser H, et al. Single-cell analysis of Hodgkin and Reed-Sternberg cells: molecular heterogeneity of gene expression and p53 mutations. Blood 1993; 81: 3097–3115.
Delabie J, Tierens A, Wu G, Weisenburger DD, Chan WC . Lymphocyte predominance Hodgkin’s disease: lineage and clonality determination using a single-cell assay. Blood 1994; 84: 3291–3298.
Roehrl MH, Becker KF, Becker I, Hšfler H . Efficiency of single-cell polymerase chain reaction from stained histologic slides and integrity of DNA in archival tissue. Diagn Mol Pathol 1997; 6: 292–297.
d'Amore F, Stribley JA, Ohno T, Wu G, Wickert RS, Delabie J, et al. Molecular studies on single cells harvested by micromanipulation from archival tissue sections previously stained by immunohistochemistry or nonisotopic in situ hybridization. Lab Invest 1997; 76: 219–224.
Burton MP, Schneider BG, Brown R, Escamilla-Ponce N, Gulley ML . Comparison of histologic stains for use in PCR analysis of microdissected, paraffin-embedded tissues. Biotechniques 1998; 24: 86–92.
Medintz I, Chiriboga L, Hotaling H, Kobilinsky L . DNA identity testing in stained tissue samples. Anal Lett 1997; 30: 909–916.
Frank TS, Svoboda-Newman SM, Hsi ED . Comparison of methods for extracting DNA from formalin-fixed paraffin sections for nonisotopic PCR. Diagn Mol Pathol 1996; 5: 220–224.
Diaz-Cano SJ, Brady SP . DNA extraction from formalin-fixed, paraffin-embedded tissues: protein digestion as a limiting step for retrieval of high-quality DNA. Diagn Mol Pathol 1997; 6: 342–346.
An SF, Fleming KA . Removal of inhibitor(s) of the polymerase chain reaction from formalin fixed, paraffin wax embedded tissues. J Clin Pathol 1991; 44: 924–927.
Pelliniemi LJ, Dym M, Karnovsky MJ . Peroxidase histochemistry using diaminobenzidine tetrahydrochloride stored as a frozen solution. J Histochem Cytochem 1980; 28: 191–192.
Hsu SM, Soban E . Color modification of diaminobenzidine (DAB) precipitation by metallic ions and its application for double immunohistochemistry. J Histochem Cytochem 1982; 30: 1079–1082.
Cordell JL, Falini B, Erber WN, Ghosh AK, Abdulaziz Z, MacDonald S, et al. Immunoenzymatic labeling of monoclonal antibodies using immune complexes of alkaline phosphatase and monoclonal anti-alkaline phosphatase (APAAP complexes). J Histochem Cytochem 1984; 32: 219–219.
Wright DK, Manos MM . Sample preparation from paraffin-embedded tissues. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols: a guide to methods and applications. New York: Academic Press; 1990. pp. 153–158.
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 1988; 239: 487–491.
Chen JT, Lane MA, Clark DP . Inhibitors of the polymerase chain reaction in Papanicolaou stain: removal with a simple destaining procedure. Acta Cytol 1996; 40: 873–877.
Bettinger C, Zimmermann HW . New investigations on hematoxylin, hematein, and hematein-aluminium complexes: II. Hematein-aluminium complexes and hemalum staining. Histochemistry 1991; 96: 215–228.
Gelfand DH . Taq DNA polymerase. In: Erlich HA, editor. PCR technology: principles and applications for DNA amplification. New York: Oxford University Press; 1992. pp. 17–22.
Tuite E, Sehlstedt U, Hagmar P, Norden B, Takahashi M . Effects of minor and major groove-binding drugs and intercalators on the DNA association of minor groove-binding proteins RecA and deoxyribonuclease I detected by flow linear dichroism. Eur J Biochem 1997; 243: 482–492.
Kim SK, Norden B . Methyl green: a DNA major-groove binding drug. FEBS Lett 1993; 315: 61–64.
Bailly C, Henichart JP, Colson P, Houssier C . Drug-DNA sequence-dependent interactions analysed by electric linear dichroism. J Mol Recogn 1992; 5: 155–171.
Fox KR, Higson SL, Scott JE . Methyl green and its analogues bind selectively to AT-rich regions of native DNA. Eur J Histochem 1992; 36: 263–270.
Melnick J, Pickering M . The mechanism of the drug induced partial displacement of methyl green from DNA. Biochem Int 1988; 16: 69–75.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Murase, T., Inagaki, H. & Eimoto, T. Influence of Histochemical and Immunohistochemical Stains on Polymerase Chain Reaction. Mod Pathol 13, 147–151 (2000). https://doi.org/10.1038/modpathol.3880028
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3880028
Keywords
This article is cited by
-
Molecular analysis of H&E- and Papanicolau-stained samples—systematic review
Histochemistry and Cell Biology (2020)
-
3,3′-Diaminobenzidine staining interferes with PCR-based DNA analysis
Scientific Reports (2018)
-
Comparison of Different Drying Methods for Recovery of Mushroom DNA
Scientific Reports (2017)
-
A simple and cost-effective method of DNA extraction from small formalin-fixed paraffin-embedded tissue for molecular oncologic testing
BMC Clinical Pathology (2014)
-
Laser capture microdissection in forensic research: a review
International Journal of Legal Medicine (2010)