Abstract
Gastric cancer is one of the most common cancers. Molecular events in the carcinogenesis of gastric cancer remain, however, largely undefined. We investigated changes in DNA copy number in 102 gastric cancers by CGH. We found changes in DNA copy number in all cases, with frequent (≧30% of patients) gains at 20q, 8q, 20p, 7q, 17q, 5p, and 13q. Frequent (≧20%) losses were found at 19p, 18q, 5q, 21q, 4p, 4q, 15q, and 17p. The mean number of total alterations was significantly lower in grade 3 and scirrhous-type carcinomas (10.81 in grade 3 vs 13.98 in grade 1 and grade 2, 9.31 in scirrhous-type vs 13.18 in medullary- and intermediate-type). The mean number of losses and total alterations were higher in tumors at pT2, pT3 and pT4 (4.68 and 12.77 in pT2, pT3, and pT4 vs 2.55 and 9.22 in pT1). The mean number of losses was higher in carcinomas with lymph node metastasis (4.83). The mean number of gains and total alterations were higher in carcinomas with venous invasion (8.44 and 13.28). Several chromosomal alterations were linked in a statistically significant manner to specific clinicopathological parameters. Gain of 17q, 20p, and 20q and loss of 4p were associated with the pattern of the cancer–stroma relationship; loss of 18q was associated with pT category; gain of 5p was associated with pN category; loss of 4q and loss of 21q were associated with lymphatic invasion; gain of 7p and loss of 4q and 18q were associated with venous invasion; and loss of 18q was associated with pathological stage. These data suggest that gain of 20q and loss of 18q might play an important role in the development and progression of gastric cancer. Moreover, some genes on 20q and 18q might be target genes of gastric cancer.
Similar content being viewed by others
Main
Gastric cancer is one of the most common cancers worldwide and is the second most common cause of cancer-related death.1 Gastric cancer has generally been resistant to chemotherapy, but effective antineoplastic drugs are now being developed in parallel with advances in cytogenetics. However, treatment of gastric cancer at advanced stages remains difficult and the prognosis is still poor, as a consequence of local recurrence and/or metastasis. The overall relative 5-year survival rate is currently less than 20%.
Molecular events in the carcinogenesis of gastric cancer remain largely unknown. Carcinogenesis, including the development of gastric cancer, is widely regarded as a multistep process involving, for example, the accumulation of genetic alterations in cellular oncogenes, tumor-suppressor genes, regulators of the cell cycle and DNA-repair genes.
Recently available genomic technologies and approaches enable us to accumulate genetic information at a rapid pace. The chromosome localization of about 26 000 genes in the human genome have already been mapped accurately.2 The expression or amplification of various genes, for example, genes for p53,3 p27,4 smad4,5 c-met,6, 7 c-erbB2,8, 9 c-myc, l-myc,10, 11 K-sam,12 E-cadherin,13β-catenin,13 VEGF,14 and FHIT15 and mutations in the genes for TP53,16 APC,17 K-ras18, 19 and E-cadherin20 have been associated with gastric carcinogenesis. There also have been reports of loss of heterozygosity (LOH) on chromosomes 1p, 2q, 3p, 4p, 5q, 7q, 8p, 9p, 11q, 13q, 14q, 17p, and 18q, suggesting a relationship between carcinogenesis and potential tumor suppressor genes.21, 22, 23 There is also a report of alterations in microsatellite DNA and other mutations in target genes in gastric cancers.24
Comparative genomic hybridization (CGH), described first by Kallioniemi et al,25 allows the detection of genetic amplifications and deletions in each chromosome in tumor cells. In a previous report, we described chromosomal alterations in squamous cell carcinomas of the esophagus26 and lung.27 We concluded that gain of a region of 3q might play an important role in the development and progression in human squamous cell carcinoma. Similarly, many chromosomal alterations have been identified by CGH in gastric cancer, but there are significant discrepancies among the reported results. In several studies, frequent chromosomal alterations, namely, gains on 1p, 6p, 7q, 8q, 11q, 16p, 17q, 20q, and 22q, and deletions on 3p, 4q, 5q, 9p, 16q, 17p, 18q, and 19p were observed in gastric cancer.28, 29, 30, 31, 32, 33, 34, 35 Recently, microarray technologies have emerged as key tools for the expression analysis of gene and genes expressed in gastric cancers have been examined using cDNA microarray technique.36
In the present study, to identify regions of the genome that might be involved in the oncogenesis of gastric cancers, we made an extensive study of chromosomal alterations in such cancers by CGH. In addition, we examined the association between chromosomal alterations and clinicopathologic parameters in patients with gastric cancer. The number of patients that we analyzed in this study, 102, is the largest in all studies reported to date.
Materials and methods
Tissues Specimens and DNA Extraction
Samples of tumor tissues were obtained, with informed consent, from 102 patients with gastric cancer who underwent surgical resection of their tumors at the 2nd Department of Surgery 2 of Oita University Hospital, Oita, Japan, between 1998 and 2003. The patients, 74 men and 28 women, age ranged from 37 to 89 years (mean age, 68.4 years). None of the patients had received chemotherapy or radiotherapy before surgery. Samples of tumors were collected immediately after surgical resection, frozen in liquid nitrogen and stored at −80°C prior to extraction of DNA. DNA was isolated by digestion with proteinase K (2 μg/ml) followed by phenol–chloroform extraction according to standard protocols using a DNA extraction kit (Qiagen). The histopathological grade and stage of each tumor were classified according to the TNM classification of the International Union against Cancer.37 Other histological features (cancer–stroma relationship, lymphatic invasion, and venous invasion) were classified according to the Japanese Classification of Gastric Carcinoma.38 The results for the cancer–stroma relationship were as follows: med (medullary-type), 24 cases; int (intermediate-type), 59 cases; and sci (scirrhous-type), 19 cases. The histopathological grades were as follows: grade 1(well-differentiated adenocarcinoma), 29 cases; grade 2 (moderately differentiated adenocarcinoma), 24 cases; and grade 3 (poorly differentiated adenocarcinoma), 49 cases. In this study, eight cases of signet-ring cell carcinoma were classified as grade 3. The histopathological primary tumors (pT) were as follows: pT1, nine cases; pT2a, 14 cases; pT2b, 22 cases; pT3, 47 cases; and pT4, 10 cases. Lymph node metastases were detected in 73 of the 102 patients, with distant metastases in 10 of the 102 patients. The postsurgical pathological stages were as follows: stage Ia, eight cases; stage Ib, 18 cases; stage II, 15 cases; stage IIIa, 15 cases; stage IIIb, 13 cases; and stage, 33 cases.
Comparative Genomic Hybridization
We performed CGH using DNA that was labeled with fluorescent dUTP, as described previously with minor modifications. In brief, DNA isolated from each tumor was labeled with Spectrum Green dUTP (Vysis, Inc), and reference DNA from blood leukocytes of healthy donors was labeled with Spectrum Red dUTP (Vysis, Inc) by nick translation. The nick-translation reaction was stopped by heating at 70°C for 15 min. The lengths of fragments used as probes ranged from approximately 300 to 3000 base pairs (bp). Lengths were confirmed by electrophoresis on a nondenaturing agarose gel. The hybridization mixture was prepared by mixing 200–400 ng of Spectrum Green-labeled tumor DNA, 200–400 ng of Spectrum Red-labeled reference DNA, 20 μg of human Cot-1 DNA (Vysis, Inc.), 0.1 vol of 3 M sodium acetate and 2.5 vol of 100% ethanol. DNA from tumors from males were always mixed with male reference DNA, and that of tumors from females was always mixed with female reference DNA. Samples were mixed briefly on a vortex mixer and then placed at −80°C for 15 min. The DNA in the mixture was then pelleted by centrifugation at 14 000 rpm for 30 min at 4°C. The supernatant was decanted and the pellet was air-dried. The DNA in the pellet was then dissolved in 10 μl of hybridization buffer (50% formamide, 10% dextran sulfate, 2 × SSC, pH 7.0), incubated at 37°C for 30 min, and denatured by incubation of 75°C for 5 min. Reference slides of normal metaphase spreads (Vysis, Inc) were incubated, to denature the DNA, at 73°C in 70% deionized formamide and 2 × SSC (pH 7.0) for 2.5 min and then dehydrated through a graded ethanol series (70, 90, and 100% ethanol). The hybridization mixture, including the probe, was immediately applied to one of the above-mentioned normal metaphase spreads. A coverslip was placed over the spread and sealed with rubber cement. Then the slide was placed in a sealed moist hybridization chamber in an incubator and incubated at 37°C for 3–5 days. After hybridization, each slide was subjected to four 12-min washes in 50% formamide in 2 × SSC (pH 7.0) at 45°C, which were followed by two 10-min washes in 2 × SSC at 45°C, one 10-min wash in 2 × SSC at room temperature, and two 5-min washes in distilled water at room temperature. Samples were counterstained with 4′,6-diamino-2-phenylindole (DAPI; Vysis, Inc.) in antifade solution.
Image Acquisition and Analysis
Three single-color images (due to the fluorescence of DAPI, Spectrum Green and Spectrum Red, respectively) were collected from each metaphase spread under an epifluorescence microscope (Olympus, Tokyo), with a 12-bit cooled charge-coupled device (CCD) camera; and a power Gene/G3-SS 1400 system (Perceptive Scientific International, PSI), and analyzed using a digital image analysis system, The Power Gene, Mac Probe (PSI). The DAPI image was used for identification of chromosomes. The fluorescence from Spectrum Green and Spectrum Red, which was specific for the tumor and the reference genome, respectively, was used to compute fluorescence ratio images and ratio profiles. For each tumor, we analyzed an average of 13 metaphases for each chromosome, including only those metaphase spreads with high-intensity hybridization and low granularity. We evaluated the corresponding ratio profiles provided that the 95% confidence limits did not exceed 0.15. To define the chromosomal regions with losses and gains of DNA, we used a 50% threshold (upper, 1.25; lower, 0.75).
Statistical Analysis
We examined associations between aberrations revealed by CGH and clinicopathologic factors by Student's t-test, the χ2-test and Fisher's probability test. Statistical significance was recognized when values of P were less than 0.05.
Results
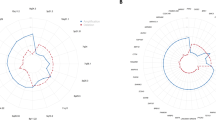
We analyzed the DNA from 102 primary gastric cancers by CGH and we detected changes in the copy number of DNA sequences in all of them. The mean number of total changes in copy number was 12.5 (range, 3–27). The mean number of gains in DNA copy number was 8.0 (range, 1–18) and the mean number of losses was 4.5 (range, 1–13). An overview of the genetic changes in the 102 gastric cancers is shown in Figure 1. Frequent (≧30% of patients) gains were found at 20q (70%), 8q (56%), 20p (47%), 7q (33%), 17q (33%), 5p (32%), and 13q (30%). Frequent (≧20% of patients) losses were found at 19p (40%), 18q (39%), 5q (27%), 21q (26%), 4p (23%), 4q (23%), 15q (21%), and 17p (21%) in more than 20% of tumors, as shown schematically in Figure 2. In particular, gain of 21q (0%) and loss of 8q (0%), 11p (0%), and 20p (0%) were not detected in any of the tumors examined. However, there was no chromosomal region in which either a gain or a loss was not detected.
Associations between the number of chromosomal alterations and clinicopathologic factors are shown in Table 1. The mean number of total (gain and loss) chromosomal alterations was significantly lower in grade 3 carcinomas and scirrhous-type carcinomas (10.81±5.12 and 9.31±5.10) than in grade 1 and grade 2 carcinomas and in medullary- and intermediate-type carcinomas (13.98±5.22 and 13.18±5.22; P=0.002 and 0.004). The mean numbers of losses and total chromosomal alterations were higher in pT2, pT3, and pT4 tumors (4.68±3.03 and 12.77±5.41; P=0.04 and 0.05). The mean number of losses was higher in carcinomas with lymph node metastasis (4.83±3.02). The mean numbers of gains and total chromosomal alterations were higher in carcinomas with venous invasion (8.44±3.42 and 13.28±5.18). Associations between the total number of chromosomal alterations and clinicopathologic factors (grade, pT, and stage) are shown in Table 2. The mean number of total chromosomal alterations decreased with histological severity, from grade 1 to grade 2 and grade 3. The mean number of total chromosomal alterations was lowest in pT1 tumors and in tumors at stage I(9.22±4.11 and 8.75±4.13).
We analyzed possible associations between clinicopathologic parameters and each chromosomal change (gain and loss). Several chromosomal alterations were associated in a statistically significant manner with specific clinicopathological parameters (P<0.05), as shown in Table 3. Gain of 17q, of 20p, and of 20q and loss of 4p were associated with the pattern of the cancer–stroma relationship; loss of 18q was associated with pT category; gain of 5p was associated with pN category, loss of 4q and of 21q was associated with lymphatic invasion; gain of 7p and loss of 4q and of 18q were associated with venous invasion; and loss of 18q was correlated with pathological stage.
Discussion
We analyzed the chromosomal alterations in 102 primary gastric cancers from patients who had not received any preoperative treatment, such as chemotherapy or irradiation.
In the present study, frequent gains (≧30% of patients) were detected at 20q, 8q, 20p, 7q, 17q, 5p, 13q by CGH. Previous CGH studies of gastric cancers yielded similar results. Moreover, several CGH studies identified the 20q region as the most frequent site of gain of DNA in gastric cancer.29, 30, 31, 32, 34, 35, 39 Amplification at 20q has been reported in several cancers, such as colon cancer,40 pancreatic cancer,41, 42 lung adenocarcinoma,43 ovarian carcinoma,44 and osteosarcoma.45 In our study, gain of 20q was detected in 71 cases (70%) and was associated with the pattern of the cancer–stroma relationship. Candidate genes at 20q are BTAK (20q13),46, 47 AIB1 (20q12),48, 39, 49, 50 TOP1 (20q12–13.1),50 TFAP2C (20q13),50 ZNF217 (20q13.2),51, 52 NABC1 (20q13.2),51 and CYP24 (20q13.2).52 Amplification and overexpression of the BTAK gene, which encodes breast tumor-amplified kinase (identical to aurora2, ARK1 and STK15), have been reported in primary gastric adenocarcinomas that were associated with aneuploidy and poor prognosis.47 Moreover, amplification and overexpression of the AIB1 (amplified in breast cancer 1) gene, a member of the steroid receptor coactivator family, appears to be useful as a marker of poor prognosis in gastric cancer.49 Recently, it was reported that TOP1, TFAP2C and NCOA3 (AIB1) might be prognostic indicators in breast cancer.50 The NCOA3 gene encodes a coactivator of steroid receptor that interacts with estrogen receptors to enhance ligand-dependent transcription. The TOP1 gene encodes topoisomerase 1, a nuclear enzyme that catalyzes single-strand breakage and rejoining of DNA, allowing the relaxation of supercoiled DNA during transcription and DNA replication. The TFAP2C gene encodes a member of the activating enhancer-binding protein-2 (AP-2) family of transcription factors. It was also been reported that the ZNF217 gene, which encodes a transcription factor, and the CYP24 gene, for a suppressor of the active form of vitamin D that inhibits cell growth, at 20q13.2 might be relevant to gastric carcinogenesis.52 Thus, expressions of these genes at 20q12–13 are likely regulated not only by amplifications of these genes but also by other mechanisms such as transcriptional activation, and expressions of these genes might play an important role in the development and progression of gastric cancers.
A gain in DNA copy number at 8q was the second most frequent gain, but it was not associated with clinicopathologic factors in our series. Amplification at 8q has been reported in several cancers, such as pancreatic cancer,42 osteosarcoma,45 and colorectal cancer.40 The genes at 8q (8q-23–24) include the c-myc, EIF3S3 and PRL-3 genes. Amplification and polymorphism of the c-myc gene has been reported in gastric cancer.10, 11 Amplification of the EIF3S3 gene has been reported as a marker of tumor progression and poor prognosis in prostate cancer.53 The EIF3S3 gene encodes the p40 subunit of eukaryotic translation initiation factor 3.54 In Addition, the PRL-3 gene has recently been demonstrated to be associated with metastasis of colorectal cancer.55 Unfortunately, we were unable to detect any correlation between these changes and clinicopathologic factors. However, the c-myc, EIF3S3, and PRL-3 genes might be relevant to the development and progression of gastric cancer.
A gain in DNA copy number at 20p was the third most frequent gain (47%) and was associated with the pattern of tumor expansion. Several CGH studies have identified 20p as a region with frequent gains in copy number in gastric cancer.28, 33 The PCNA genes at 20p (20p12) encodes proliferating cell nuclear antigen, which is expressed in advanced gastric cancers with poor prognosis.56 We considered that some strange genes exist on 20p, further research on 20p may prove fruitful.
In our study, we detected gains in DNA copy number at 7q and 17q in 34 cases (33%). Gain at 7q was not associated with clinicopathologic factors in our series. Expression of the c-met gene, which is located at 7q31, is associated with clinical stage and/or prognosis in gastric cancer.6, 7, 57 Furthermore, amplification of 7q21 is associated with expression of the HGF gene, which encodes hepatocyte growth factor. Serum levels of HGF were reported to correlate with the aggressiveness of gastric carcinomas.58 Gain at 17q was associated with clinicopathologic factor (the pattern of the cancer–stroma relationship), as was the case for gain at 20q. Strong amplification of the region 17q12–21 was reported in the intestinal type of gastric cancer.59 Amplification and/or overexpression of some genes on chromosome 17q21.1, such as ERBB-2 (HER2/neu)8, 9, 60, 61 and TOP2A (for topoisomerase IIα),62 have been reported in gastric cancer and breast cancer, and might represent prognostic factors. Recently, an antineoplastic drug that targets the HER2 gene has been used to treat breast cancer and there are reports of overexpression of the HER2/neu gene in gastric cancer.61 Therefore, the same antineoplastic drug might be useful for the treatment of gastric cancer. It has also been reported that gain at 17q is a powerful prognostic factor and a candidate gene on 17q23 is PPM1D, whose expression is associated with prognosis in neuroblastoma.63 Strong amplification of the 17q22–23 regions has been detected in breast cancer,64 and the amplified genes include RCH1, PAT1, PS6K, APPBP2, and MUL genes.64, 65 In our study, gains at 17q were associated with the patterns of tumor infiltration and expansion. As noted above, genes on 17q might be associated with tumor progression.
Overexpressions and/or amplifications of genes on 20q, 8q, 20p, 7q, and 17q were associated with clinicopathological factors, and this result indicates that 20q, 8q, 20p, 7q, and 17q may harbor putative oncogenes that play an important role in gastric cancer pathogenesis.
There also have been reports of loss of heterozygosity (LOH) on 1p, 2q, 3p, 4p, 5q, 7q, 8p, 9p, 11q, 13q, 14q, 17p, 18q, and 21q, suggesting an association between potential tumor suppressor genes and the development and progression of gastric cancers.22, 23, 24 In our study, the most common losses were found at 19p and 18q, and the mean numbers of losses and of total chromosomal alterations were higher in pT2, pT3, and pT4 tumors (4.68 and 12.77; P=0.04 and 0.05) and the mean number of losses of DNA was higher in carcinomas with lymph node metastasis (4.83), as shown in Table 1. Deletion of 19p has been reported in gastric cancer, and chromosome 19p might include tumor suppressor genes that play an important role in the development and progression in gastric cancers.39 Recently, the gene for antizyme (OAZ1), a negative regulator of cellular polymines, is mapped to 19p13.3, where frequent allelic imbalance is observed in ovarian cancer. But it was reported that one or more tumor suppressor genes other than OAZ1 gene exist on 19p13.3.66 It has also been reported that inactivation of LKB1/STK11 gene on 19p is a very common event and might be important in the development of sporadic lung adenocarcinoma.67 The LKB1/STK11 protein is a serine–threonine kinase whose biological function has not been fully elucidated. Although it was considered that some strange tumor suppressor genes exist on 19p, researches of 19p are rare so far and further researches are needed.
With respect to associations between clinicopathologic parameters and individual chromosomal alterations (gain and loss), losses at 18q were associated with pT category and pathological stage in our series. Several candidate genes on 18q, including DCC (detected in colorectal cancer) (18q21.3), Smad2 (18q21), Smad4 (18q21.1), and bcl2 (18q21.3) have been mapped to this chromosome. LOH at the DCC locus (18q21) is frequently found in gastric cancer.68, 69 A decreased level of DCC mRNA in gastric cancer is associated not only with the expanding phenotype but also with metastasis to the liver.70 The expression of Smad4 is a favorable prognostic factor in gastric cancer.5 Smad4 appears to be the key regulator of the transforming growth factor β signaling pathway and of control transcription driven by this superfamily. Smad2 and DCC have been found to be inactivated in subgroups of colon cancers,71, 72, 73 and the expression of bcl-2 is associated with a better prognosis in gastric cancer.74, 75 In this tudy, losses at 18q were associated with pT category and pathological stage in our series. DCC, Smad2, and Smad4 are known as tumor suppressor genes, and loss of these tumor suppressor genes might be associated with the tumor progression, and might play a significant role in carcinogenesis of gastric cancer.
The mean numbers of gains and of total chromosomal alterations were higher in carcinomas with venous invasion (8.44 and 13.28). Gains at 7p and losses at 4q and 18q were associated with venous invasion. Gains at 7p and losses at 4q were detected by previous studies of gastric cancers by CGH.76, 77, 78 Candidate genes in these regions might include the gene for EGFR (epidermal growth factor receptor) at 7p12. A study by differential PCR demonstrated amplification of the gene for EGFR in 30% of esophageal adenocarcinomas.79 And genetic deletions at 4q have been detected in esophageal adenocarcinomas.80, 81 However, no genes have been implicated in the deletions in 4q region, undiscovered tumor suppressor genes might reside here.
In the association between the number of chromosomal alterations and clinicopathologic factors, the mean number of total chromosomal alterations was significantly lower in grade 3 carcinomas. Although genetic alterations tend to accumulate as the histopathological grade goes up, we found the opposite to be true in our series. There is a greater amount of stroma in grade 3 carcinomas than in grade 1 and grade 2 carcinomas, so it is possible that insufficient quantities of cancer cells were extracted for analysis by CGH from grade 3 carcinomas. The mean number of total chromosomal alterations in our study was significantly lower in scirrhous-type carcinoma than in medullary- and intermediate-type carcinoma.
We analyzed the chromosomal aberrations in 102 primary gastric cancers. The results of the present and previous studies suggest that gains at 20q, 17q, and 8q and losses at 19p and 18q might play an important role in the development and progression of gastric cancer. In particular, gains at 20q, which were the most frequent gains, and losses at 18q, which were associated with clinicopathologic factors (pT category and pathological stage) might be markers of the progression of human adenocarcinomas that include gastric cancer, lung adenocarcinoma, colorectal adenocarcinoma, and others. The relevant genes within these regions remain to be identified and further studies are required to identify these genes and to determine whether genes in these regions of the human genome behave as oncogenes that are important in gastric cancer, as might now be expected.
References
Whelan S, Parkin D, Masuyer E . Trends in Cancer Incidence and Mortality. IARC Scientific Publ, no. 102 IARC: Lyon, France, 1993.
Venter JC, Adams MD, Myers EW, et al. The sequence of the human genome. Science 2001;291:1304–1351.
Lee WJ, Shun CT, Hong RL, et al. Overexpression of p53 predicts shorter survival in diffuse-type gastric cancer. Br J Surgery 1998;85:1138–1142.
Nitti D, Belluco C, Mammano E, et al. Low level of p27 (Kip1) protein expression in gastric adenocarcinoma is associated with disease progression and poor outcome. J Surg Oncol 2002;81:167–176.
Xiangming C, Natsugoe S, Takeo S, et al. Preserved Smad4 expression in the transforming growth factor β signaling pathway is a favorable prognostic factor in patients with advanced gastric cancer. Clin Cancer Res 2001;7:277–282.
Kuniyasu H, Yasui W, Yokozaki H, et al. Aberrant expression of c-met mRNA in human gastric carcinomas. Int J Cancer 1993;55:72–75.
Heideman DAM, Snijders PJF, Bloemena E, et al. Absence of tpr-met and expression of c-met in human gastric mucosa and carcinoma. J Pathol 2001;194:428–435.
Yokota J, Yamamoto T, Miyajima N, et al. Genetic alterations of the c-erbB-2 oncogene occur frequently in tubular adenocarcinoma of the stomach and are often accompanied by amplification of v-erbA homologue. Oncogene 1988;2:283–288.
Vidgren V, Varis A, Kokkola A, et al. Concomitant gastrin and ERBB2 gene amplifications at 17q12–q21 in the intestinal type of gastric cancer. Genes Chromosomes Cancer 1999;24:24–29.
Suzuki S, Tenjin T, Watanabe H, et al. Low level c-myc gene amplification in gastric cancer detected by dual-color fluorescence in situ hybridization analysis. J Surg Oncol 1997;66:173–178.
Shibuta K, Mori M, Haraguchi M, et al. Association between restriction fragment length polymorphism of the L-myc gene and susceptibility to gastric cancer. Br J Surg 1998;85:681–684.
Hattori Y, Odagiri H, Nakatani H, et al. K-sam, an amplified gene in stomach cancer, is a member of the heparin-binding growth factor receptor genes. Proc Natl Acad Sci USA 1990;87:5983–5987.
Tanaka M, Kitajima Y, Edakuni G, et al. Abnormal expression of E-cadherin and β-catenin may be a molecular marker of submucosal invasion and lymph node metastasis in early gastric cancer. Br J Surg 2002;89:236–244.
Kimura H, Konishi K, Nukui T, et al. Prognostic significance of expression of thymidine phosphorylase and vascular endothelial growth factor in human gastric carcinoma. J Surg Oncol 2001;76:31–36.
Huiping C, Kristjansdottir S, Bergthorsson JT, et al. High frequency of LOH, MSI and abnormal expression of FHIT in gastric cancer. Eur J Cancer 2002;38:728–735.
Tamura G, Kihana T, Nomura K, et al. Detection of frequent p53 gene mutations in primary gastric cancer by cell sorting and polymerase chain reaction single-strand conformation polymorphism analysis. Cancer Res 1991;51:3056–3058.
Horii A, Nakasturu S, Miyoshi Y, et al. The APC gene, responsible for familial adenomatous polyposis, is mutated in human gastric cancer. Cancer Res 1992;52:3231–3233.
Hiyama T, Haruma K, Kitadai Y, et al. K-ras mutation in Helicobacter pylori-associated chronic gastritis in patients with and without gastric cancer. Int J Cancer 2002;97:562–566.
Ranzani GN, Renault B, Pellegra NS, et al. Loss of heterozygosity and K-ras gene mutations in gastric cancer. Hum Genet 1993;92:244–249.
Becker KF, Atkinson MJ, Reich U, et al. E-cadherin gene mutations provide clues to diffuse-type gastric carcinomas. Cancer Res 1994;54:3845–3852.
Yustein AS, Harper JC, Petroni GR, et al. Allotype of gastric adenocarcinoma. Cancer Res 1999;59:1437–1441.
Tamura G, Sakata K, Nishizuka S, et al. Allelotype of adenoma and differentiated adenocarcinoma of the stomach. J Pathol 1996;180:371–377.
Nishizuka S, Tamura G, Terashima M, et al. Loss of heterozygosity during the development and progression of differentiated adenocarcinoma of the stomach. J Pathol 1998;185:38–43.
Ogata S, Tamura G, Endoh Y, et al. Microsatellite alterations and target gene mutations in the early stage of multiple gastric cancer. J Pathol 2001;194:334–340.
Kallionemi OP, Kallionemi A, Piper J, et al. Optimizing comparative genomic hybridization for analysis of DNA sequence copy number changes in solid tumors. Genes Chromosomes Cancer 1994;10:231–243.
Noguchi T, Kimura Y, Takeno S, et al. Chromosomal imbalance in esophageal squamous cell carcinoma: 3q gain correlates with tumor progression but not prognostic significance. Oncol Rep 2003;10:1393–1400.
Chujo M, Noguchi T, Miura T, et al. Comparative genomic hybridization analysis detected frequent overrepresentation of chromosome 3q in squamous cell carcinoma of the lung. Lung Cancer 2002;38:23–29.
Sakakura C, Mori T, Sakabe T, et al. Gains, losses, and amplifications of genomic materials in primary gastric cancers analyzed by comparative genomic hybridization. Genes Chromosomes Cancer 1999;24:299–305.
Varis A, Puolakkainen P, Savolainen H, et al. DNA copy number profiling in esophageal Barrett adenocarcinoma: comparison with gastric adenocarcinoma and esophageal squamous cell carcinoma. Cancer Genet Cytogenet 2001;127:53–58.
Okada K, Sugihara H, Bamba M, et al. Sequential numerical changes of chromosomes 7 and 18 in diffuse-type stomach cancer cell lines: combined comparative genomic hybridization, fluorescence in situ hybridization, and ploidy analyses. Genes Chromosomes Cancer 2000;118:99–107.
Wu MS, Chang MC, Huang AP, et al. Correlation of histologic subtype and replication error phenotype with comparative genomic hybridization in gastric cancer. Genes Chromosomes Cancer 2001;30:80–86.
Dekken H, Alers JC, Riegman PHJ, et al. Molecular cytogenetic evaluation of gastric cardia adenocarcinoma and precursor lesions. Am J Pathol 2001;158:1961–1967.
Kim YH, Kim NG, Lim JG, et al. Chromosomal alterations in paired gastric adenoma and carcinoma. Am J Pathol 2001;158:655–662.
Varis A, van Rees B, Weterman M, et al. DNA copy number changes in young gastric cancer patients with special reference to chromosome 19. Br J Cancer 2003;16:1914–1919.
Noguchi T, Wirtz HC, Michaelis S, et al. Chromosomal imbalance in gastric cancer. Correlation with histologic subtype and tumor progression. Am J Clin Pathol 2001;115:828–834.
Lee S, Back M, Yang H, et al. Identification of genes differentially expressed between gastric cancers and normal gastric mucosa with cDNA microarrays. Cancer Letters 2002;184:197–206.
Sobin LH, Wittekind CH . TNM Classification of Malignant Tumors. UICC (International Union against Cancer). 6th edn. Wiley: New York, 2002, pp 65–68.
Japanese Gastric Cancer Association. Japanese Classification of Gastric Carcinoma 2nd English edn. Gastric Cancer 1998;1:10–24.
Guan XY, Fu SB, Xia JC, et al. Recurrent chromosome changes in 62 primary gastric carcinomas detected by comparative genomic hybridization. Cancer Genet Cytogenet 2000;123:27–34.
Ghadimi BM, Grade M, Liersch, et al. Gain of chromosome 8q23–24 is a predictive marker for lymph node positivity in colorectal cancer. Clin Cancer Res 2003;9:1808–1814.
Toldo SS, Wallrapp C, Pillasch FM, et al. Mapping of chromosomal imbalances in pancreatic carcinoma by comparative genomic hybridization. Cancer Res 1996;56:3803–3807.
Mahlamaki EH, Barlund M, Tanner M, et al. Frequent amplification of 8q24, 11q, 17q, and 20q-specific genes in pancreatic cancer. Genes Chromosomes Cancer 2002;35:353–358.
Wong MP, Fung LF, Wang E, et al. Chromosomal aberrations of primary lung adenocarcinoma in nonsmokers. Cancer 2003;97:1263–1270.
Iwabuchi H, Sakamoto M, Sakunaga H, et al. Genetic analysis of benign, low-grade, and high-grade ovarian tumors. Cancer Res 1995;55:6172–6180.
Bayani J, Zielebska M, Pandita A, et al. Spectral karyotyping identifies recurrent complex rearrangements of chromosomes 8, 17, and 20 in osteosarcoma. Genes Chromosomes Cancer 2003;36:7–16.
Collins C, Rommens JM, Kowbel D, et al. Positional cloning of ZNF217 and NABC1: genes amplified at 20q13.2 and overexpressed in breast carcinoma. Proc Natl Acad Sci USA 1998;8703–8708.
Sakakura C, Hagiwara A, Yasuoka R, et al. Tumor-amplified kinase BTAK is amplified and overexpressed in gastric cancers with possible involvemant in aneuploid formation. Br J Cancer 2001;84:824–831.
Anzick SL, Kononen J, Walker RL, et al. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science 1997;277:965–968.
Sakakura C, Hagiwara A, Yasuoka R, et al. Amplification and overexpression of the AIB1 nuclear receptor co-activator gene in primary gastric cancer. Int J Cancer 2000;89:217–223.
Zhao C, Yasui K, Lee CJ, et al. Elevated expression levels of NCOA3, TOP1, and TFAP2C in breast tumors as predictors of poor prognosis. Cancer 2003;98:18–23.
Sen S, Zhou H, White RA . A putative serine/threonine kinase-encoding gene BTAK on chromosome 20q13 is amplified and overexpressed in human breast cancer cell lines. Oncogene 1997;14:2195–2200.
Weiss MM, Snijders AM, Kuipers EJ, et al. Determination of amplicon boundaries at 20q13.2 in tissue samples of human gastric adenocarcinomas by high-resolution microarray comparative genomic hybridization. J Pathol 2003;200:320–326.
Saramaki O, Willi N, Bratt O, et al. Amplification of EIF3S3 gene is associated with advanced stage in prostate cancer. Am J Pathol 2001;159:2089–2094.
Hershey JW, Asano K, Naranda T, et al. Conservation and diversity in the structure of translation initiation factor EIF3 from humans and yeast. Biochimie 1996;78:903–907.
Saha S, Bardelli A, Buckhaults P, et al. Phosphatase associated with metastasis of colorectal cancer. Science 2001;294:1343–1346.
Yonemura Y, Kimura H, Fushida S, et al. Analysis of proliferative activity using anti-proliferating cell nuclear antigen antibody in gastric cancer tissue specimens obtained by endoscopic biopsy. Cancer 1993;71:2448–2453.
Kuniyasy H, Yasui W, Kitadai Y, et al. Frequent amplification of c-met genes in scirrhous-type stomach cancer. Biochem Biophys Res Commum 1992;189:227–232.
Han SU, Lee JH, Kim WH, et al. Significant correlation between serum level of hepatocyte growth factor and progression of gastric carcinoma. World J Surg 1999;23:1176–1180.
Kokkola A, Monni O, Puolakkainen P, et al. 17q12–21 amplicon, a novel recurrent genetic change in intestinal type of gastric carcinoma: a comparative genomic hybridization study. Genes Chromosomes Cancer 1997;20:38–43.
Zhao J, Wu R, Au A, et al. Determination of HER2 gene amplification by chromogenic in situ hybridization (CISH) in archival breast carcinoma. Mod Pathol 2002;15:657–665.
Kono K, Takahashi A, Amemiya H, et al. Frequencies of HER2/NEU overexpression relating to HLA haplotype in patients with gastric cancer. Int J Cancer 2002;98:216–220.
Jarvinen TA, Holli K, Kuukasjarvi T, et al. Predictive value of topoisomerase II alpha and other prognostic factors for epirubicin chemotherapy in advanced breast cancer. Br J Cancer 2001;61:903–907.
Saito F, Imoto I, Inoue J, et al. PPM1D is potential target for 17q gain in neuroblastoma. Cancer Res 2003;63:1876–1883.
Forozan F, Mahlamaki EH, Monni O, et al. Comparative genomic hybridization analysis of 38 breast cancer cell lines: a basis for interpreting complementary DNA microarray data. Cancer Res 2000;60:4519–4525.
Couch FJ, Wang XY, Wu GJ, et al. Localization of PS6 K to chromosomal region 17q23 and determination of its amplification in breast cancer. Cancer Res 1999;59:1408–1411.
Yanahara N, Okamoto A, Matsufuji S . A commonly deleted region in ovarian cancer on chromosome 19p13.3, not including the OAZ1 gene. Int J Oncol 2003;23:567–575.
Sanchez-Cespedes M, Parrella P, Estella M, et al. Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res 2002;62:3659–3662.
Sato K, Tamura G, Tsuchiya T, et al. Frequent loss of expression without sequence mutations of the DCC gene in primary gastric cancer. Br J Cancer 2001;85:199–203.
Uchino S, Tsuda H, Nogichi M, et al. Frequent loss of heterozygosity at the DCC locus in gastric cancer. Cancer Res 1992;52:3099–3102.
Yoshida Y, Itoh F, Endo T, et al. Decreased DCC mRNA expression in human gastric cancers is clinicopathologically significant. Int J Cancer 1998;79:634–639.
Thiagalingam S, Lengauer C, Leach FS, et al. Evaluation of candidate tumor suppressor genes on chromosome 18 in colorectal cancers. Nature Genet 1996;13:343–346.
Riggins GJ, Kinzler KW, Vogelstein B, et al. Frequency of Smad gene mutations in human cancers. Cancer Res 1997;57:2578–2580.
Fearon E, Cho K, Nigro J, et al. Identification of a chromosome 18q gene that is altered in colorectal cancers. Science 1989;247:49–56.
Aizawa K, Ueki K, Suzuki S, et al. Apoptosis and Bcl-2 expression in gastric carcinoma: correlation with clinicopathological variables, p53 expression, cell proliferation and prognosis. Int J Oncol 1999;14:85–91.
Inada T, Kikuyama S, Ichikawa A, et al. Bcl-2 expression as a prognostic factor of survival of gastric carcinoma. Anticancer Res 1998;18:2003–2010.
Moskaluk CA, Hu J, Perlman EJ . Comparative genomic hybridization of esophageal and gastroesophageal adenocarcinomas shows consensus areas of DNA gain and loss. Genes Chromosomes Cancer 1998;22:305–311.
Rifai WE, Harper JC, Cummings OW, et al. Consistent genetic alterations in xenografts of proximal stomach and gastro-esophageal junction adenocarcinoma. Cancer Res 1998;58:34–37.
Dekken H, Geelen E, Dinjens WNM, et al. Comparative genomic hybridization of cancer of the gastroesophageal junction: deletion of 14q31–32.1 discriminates between esophageal (Barrett's) and gastric cardia adenocarcinomas. Cancer Res 1999;59:748–752.
Kasspooles M, Moore JH, Orringer MB, et al. Amplification and over-expression of the EGFR and erbB-2 genes in human esophageal adenocarcinomas. Int J Cancer 1993;54:213–219.
Hommoud ZT, Kaleem Z, Cooper JD, et al. Allelotype analysis of esophageal adenocarcinomas: evidence for the involvement of sequences on the long arm of chromosome 4. Cancer Res 1996;56:4499–4502.
Rumpel CA, Powell SM, Moskaluk CA . Mapping of genetic deletions on the long arm of chromosome 4 in human esophageal adenocarcinomas. Am J Pathol 1999;154:1329–1334.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kimura, Y., Noguchi, T., Kawahara, K. et al. Genetic alterations in 102 primary gastric cancers by comparative genomic hybridization: gain of 20q and loss of 18q are associated with tumor progression. Mod Pathol 17, 1328–1337 (2004). https://doi.org/10.1038/modpathol.3800180
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800180
Keywords
This article is cited by
-
TTPAL promotes gastric tumorigenesis by directly targeting NNMT to activate PI3K/AKT signaling
Oncogene (2021)
-
Association between Genetic Polymorphisms in Superoxide Dismutase Gene Family and Risk of Gastric Cancer
Pathology & Oncology Research (2020)
-
Genomic landscape of platinum resistant and sensitive testicular cancers
Nature Communications (2020)
-
Molecular Profiles and Metastasis Markers in Chinese Patients with Gastric Carcinoma
Scientific Reports (2019)
-
Genomic copy number analysis of a spectrum of blue nevi identifies recurrent aberrations of entire chromosomal arms in melanoma ex blue nevus
Modern Pathology (2016)